Abstract
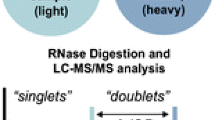
Mass spectrometry (MS) has become an enabling technology for the characterization of post-transcriptionally modified nucleosides within ribonucleic acids (RNAs). These modified RNAs tend to be more challenging to completely characterize using conventional genomic-based sequencing technologies. As with many biological molecules, information relating to the presence or absence of a particular compound (i.e., qualitative measurement) is only one step in sample characterization. Additional useful information is found by performing quantitative measurements on the levels of the compound of interest in the sample. Phosphate labeling of modified RNAs has been developed by our laboratory to enhance conventional mass spectrometry techniques. By taking advantage of the mechanism of action of many ribonucleases (RNases), digesting RNA samples in the presence of 18O-labeled water generates an 18O-labeled 3′-phosphate in each digestion product. We describe the historical development of this approach, contrast this stable isotope labeling strategy with others used in RNA mass spectrometry, and provide examples of new analytical mass spectrometry methods that are enabled by phosphate labeling in this fashion.


Reproduced with permission from Berhane et al. [28] Copyright 2003

Reproduced with permission from Meng et al. [29] Copyright 2005

Reproduced with permission from Li et al. [33] Copyright 2012

Reproduced with permission from Li et al. [34] Copyright 2013

Reproduced with permission from Wetzel et al. [35] Copyright 2014
Similar content being viewed by others
References
Nachtergaele S, He C (2016) The emerging biology of RNA post-transcriptional modifications. RNA Biol 14(2):156–163
Song J, Yi C (2017) Chemical modifications to RNA: a new layer of gene expression regulation. ACS Chem Biol 12(2):316–325
Zhang X et al (2016) Small RNA modifications: integral to function and disease. Trends Mol Med 22(12):1025–1034
Kowalak JA et al (1993) A novel method for the determination of post-transcriptional modification in RNA by mass spectrometry. Nucleic Acids Res 21(19):4577–4585
Kowalak JA, Bruenger E, McCloskey JA (1995) Posttranscriptional modification of the central loop of domain V in Escherichia coli 23 S ribosomal RNA. J Biol Chem 270(30):17758–17764
Ni J et al (1996) Interpretation of oligonucleotide mass spectra for determination of sequence using electrospray ionization and tandem mass spectrometry. Anal Chem 68(13):1989–1999
McLuckey SA, Van Berkel GJ, Glish GL (1992) Tandem mass spectrometry of small, multiply charged oligonucleotides. J Am Soc Mass Spectrom 3:60–70
Kirpekar F, Douthwaite S, Roepstorff P (2000) Mapping posttranscriptional modifications in 5S ribosomal RNA by MALDI mass spectrometry. RNA 6:296–306
Meng Z, Limbach PA (2006) Mass spectrometry of RNA: linking the genome to the proteome†. Brief Funct Genom 5(1):87–95
Ross R et al (2016) Sequence mapping of transfer RNA chemical modifications by liquid chromatography tandem mass spectrometry. Methods 107:73–78
Limbach PA, Paulines MJ (2017) Going global: the new era of mapping modifications in RNA. Wiley Interdiscip Rev: RNA 8(1):e1367
Li X, Xiong X, Yi C (2016) Epitranscriptome sequencing technologies: decoding RNA modifications. Nat Methods 14(1):23–31
Ong SE et al (2002) Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Mol Cell Proteom 1(5):376–386
Ong SE, Mann M (2006) A practical recipe for stable isotope labeling by amino acids in cell culture (SILAC). Nat Protoc 1(6):2650–2660
Brückl T et al (2009) Parallel isotope-based quantification of modified tRNA nucleosides. Angew Chem Int Ed Engl 48(42):7932–7934
Kellner S et al (2014) Absolute and relative quantification of RNA modifications via biosynthetic isotopomers. Nucleic Acids Res 42(18):e142
Waghmare SP, Dickman MJ (2011) Characterization and quantification of RNA post-transcriptional modifications using stable isotope labeling of RNA in conjunction with mass spectrometry analysis. Anal Chem 83(12):4894–4901
Popova AM, Williamson JR (2014) Quantitative analysis of rRNA modifications using stable isotope labeling and mass spectrometry. J Am Chem Soc 136(5):2058–2069
Taoka M et al (2015) A mass spectrometry-based method for comprehensive quantitative determination of post-transcriptional RNA modifications: the complete chemical structure of Schizosaccharomyces pombe ribosomal RNAs. Nucleic Acids Res 43(18):e115
Paulines MJ, Limbach PA (2017) Stable isotope labeling for improved comparative analysis of RNA digests by mass spectrometry. J Am Soc Mass Spectrom. doi:10.1007/s13361-017-1593-3
Sprinson DB, Rittenberg D (1951) Nature of the activation process in enzymatic reactions. Nature 167(4247):484
Desiderio DM, Kai M (1983) Preparation of stable isotope-incorporated peptide internal standards for field desorption mass spectrometry quantification of peptides in biologic tissue. Biomed Mass Spectrom 10(8):471–479
Mirgorodskaya OA et al (2000) Quantitation of peptides and proteins by matrix-assisted laser desorption/ionization mass spectrometry using (18)O-labeled internal standards. Rapid Commun Mass Spectrom 14(14):1226–1232
Yao X et al (2001) Proteolytic 18O labeling for comparative proteomics: model studies with two serotypes of adenovirus. Anal Chem 73(13):2836–2842
Qian WJ et al (2009) Large-scale multiplexed quantitative discovery proteomics enabled by the use of an (18)O-labeled “universal” reference sample. J Proteome Res 8(1):290–299
Lange S et al (2010) Identification of phosphorylation-dependent interaction partners of the adapter protein ADAP using quantitative mass spectrometry: SILAC vs (18)O-labeling. J Proteome Res 9(8):4113–4122
Hamasaki T et al (2013) Synthesis of (1)(8)O-labeled RNA for application to kinetic studies and imaging. Nucleic Acids Res 41(12):e126
Berhane BT, Limbach PA (2003) Stable isotope labeling for matrix-assisted laser desorption/ionization mass spectrometry and post-source decay analysis of ribonucleic acids. J Mass Spectrom 38:872–878
Meng Z, Limbach PA (2005) Quantitation of ribonucleic acids using 18-O labeling and mass spectrometry. Anal Chem 77:1891–1895
Hartmer R et al (2003) RNase T1 mediated base-specific cleavage and MALDI-TOF MS for high-throughput comparative sequence analysis. Nucleic Acids Res 31(9):e47
Castleberry CM, Limbach PA (2010) Relative quantitation of transfer RNAs using liquid chromatography-mass spectrometry (LC–MS) and signature digestion products. Nucleic Acids Res 38:e162
Hossain M, Limbach PA (2007) Mass spectrometry-based detection of transfer RNAs by their signature endonuclease digestion products. RNA 13(2):295–303
Li S, Limbach PA (2012) Method for comparative analysis of ribonucleic acids using isotope labeling and mass spectrometry. Anal Chem 84(20):8607–8613
Li S, Limbach PA (2013) Mass spectrometry sequencing of transfer ribonucleic acids by the comparative analysis of RNA digests (CARD) approach. Analyst 138(5):1386–1394
Wetzel C, Li S, Limbach PA (2014) Metabolic de-isotoping for improved LC–MS characterization of modified RNAs. J Am Soc Mass Spectrom 25(7):1114–1123
Li S, Limbach PA (2014) Identification of RNA sequence isomers by isotope labeling and LC–MS/MS. J Mass Spectrom 49(11):1191–1198
Bruce AG, Uhlenbeck OC (1978) Reactions at the termini of tRNA with T4 RNA ligase. Nucleic Acids Res 5(10):3665–3677
England TE, Bruce AG, Uhlenbeck OC (1980) Specific labeling of 3′ termini of RNA with T4 RNA ligase. Meth Enzymol 65(1):65–74
Viollet S et al (2011) T4 RNA ligase 2 truncated active site mutants: improved tools for RNA analysis. BMC Biotechnol 11:72
Abad MG, Rao BS, Jackman JE (2010) Template-dependent 3′-5′ nucleotide addition is a shared feature of tRNAHis guanylyltransferase enzymes from multiple domains of life. Proc Natl Acad Sci USA 107(2):674–679
Jackman JE, Gott JM, Gray MW (2012) Doing it in reverse: 3′-to-5′ polymerization by the Thg1 superfamily. RNA 18(5):886–899
Timms JF, Cutillas PR (2010) Overview of quantitative LC–MS techniques for proteomics and activitomics. Methods Mol Biol 658:19–45
Acknowledgements
Financial support for our lab’s decade-long work in the area of stable isotope labeling of RNA has been generously supported by the National Science Foundation, including our current NSF support (CHE1507357). The generous support of the University of Cincinnati and the Rieveschl Endowment for these studies are also appreciated.
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is part of the Topical Collection “Phosphate Labeling and Sensing in Chemical Biology”; edited by Henning Jessen.
Rights and permissions
About this article
Cite this article
Borland, K., Limbach, P.A. Applications and Advantages of Stable Isotope Phosphate Labeling of RNA in Mass Spectrometry. Top Curr Chem (Z) 375, 33 (2017). https://doi.org/10.1007/s41061-017-0121-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s41061-017-0121-z