Abstract
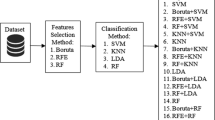
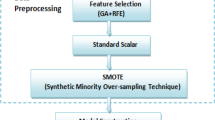
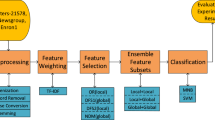
Classification of high dimensional data is a very crucial task in bioinformatics. Cancer classification of the microarray is a typical application of machine learning due to the large numbers of genes. Feature (genes) selection and classification with computational intelligent techniques play an important role in diagnosis and prediction of disease in the microarray. Artificial neural networks (ANN) is an artificial intelligence technique for classifying, image processing and predicting the data. This paper evaluates the performance of ANN classifier using six different hybrid feature selection techniques, for gene selection of microarray data. These hybrid techniques use Independent component analysis (ICA), as an extraction technique, popular filter techniques and bio-inspired algorithm for optimization of the ICA feature vector. Five binary gene expression microarray datasets are used to compare the performance of these techniques and determine how these techniques improve the performance of ANN classifier. These techniques can be extremely useful in feature selection because they achieve the highest classification accuracy along with the lowest average number of selected genes. Furthermore, to check the significant difference between these different algorithms a statistical hypothesis test was employed with a certain level of confidence. The experimental result shows that a combination of ICA with genetic bee colony algorithm shows superior performance as it heuristically removes non-contributing features to improve the performance of classifiers.





Similar content being viewed by others
References
Xu Y, Selaru FM, Yin J, Zou TT, Shustova V, Mori Y, Sato F, Liu TC, Olaru A, Wang S (2002) Artificial neural networks and gene filtering distinguish between global gene expression profiles of Barrett’s esophagus and esophageal cancer. Cancer Res 62(12):3493–3497
Ng RT, Pei J (2007) Introduction to the special issue on data mining for health informatics. ACM SIGKDD Explor Newsl 9(1):1–2
Aziz R, Verma C, Srivastava N (2017) Dimension reduction methods for microarray data: a review. AIMS Bioeng 4:179–197
Shang C, Shen Q (2005) Aiding classification of gene expression data with feature selection: a comparative study. Int J Comput Intell Res 1(1):68–76
Aziz R, Verma C, Srivastava N (2015) A weighted-SNR feature selection from independent component subspace for NB classification of microarray data. Int J Adv Biotechnol Res 6(2):245–255
Peng Y (2006) A novel ensemble machine learning for robust microarray data classification. Comput Biol Med 36(6):553–573
Mohan A, Rao MD, Sunderrajan S, Pennathur G (2014) Automatic classification of protein structures using physicochemical parameters. Interdiscip Sci Comput Life Sci 6(3):176–186
Dash R (2017) A two stage grading approach for feature selection and classification of microarray data using Pareto based feature ranking techniques: a case study. J King Saud Univ Comput Inf Sci. https://doi.org/10.1016/j.jksuci.2017.08.005
Jain AK, Mao J, Mohiuddin K (1996) Artificial neural networks: a tutorial. Computer 3:31–44
Tong DL, Schierz AC (2011) Hybrid genetic algorithm-neural network: feature extraction for unpreprocessed microarray data. Artif Intell Med 53(1):47–56
Peterson LE, Ozen M, Erdem H, Amini A, Gomez L, Nelson CC, Ittmann M (2005) Artificial neural network analysis of DNA microarray-based prostate cancer recurrence. In: Proceedings of the 2005 IEEE symposium on computational intelligence in bioinformatics and computational biology, CIBCB’05. IEEE, pp 1–8
Khan J, Wei JS, Ringner M, Saal LH, Ladanyi M, Westermann F, Berthold F, Schwab M, Antonescu CR, Peterson C (2001) Classification and diagnostic prediction of cancers using gene expression profiling and artificial neural networks. Nat Med 7(6):673–679
Lancashire LJ, Lemetre C, Ball GR (2009) An introduction to artificial neural networks in bioinformatics—application to complex microarray and mass spectrometry datasets in cancer studies. Brief Bioinform 10:315–329
Sarhan AM (2009) Cancer classification based on microarray gene expression data using DCT and ANN. J Theor Appl Inf Technol 6(2):208–216
Huynh H, Kim J-J, Won Y (2009) Classification study on DNA micro array with feed forward neural network trained by singular value decomposition. Int J Bio-Sci Bio-Technol 1(1):17–24
Catto JW, Abbod MF, Wild PJ, Linkens DA, Pilarsky C, Rehman I, Rosario DJ, Denzinger S, Burger M, Stoehr R (2010) The application of artificial intelligence to microarray data: identification of a novel gene signature to identify bladder cancer progression. Eur Urol 57(3):398–406
Fernández-Navarro F, Hervás-Martínez C, Ruiz R, Riquelme JC (2012) Evolutionary generalized radial basis function neural networks for improving prediction accuracy in gene classification using feature selection. Appl Soft Comput 12(6):1787–1800
Yu H, Hong S, Yang X, Ni J, Dan Y, Qin B (2013) Recognition of multiple imbalanced cancer types based on DNA microarray data using ensemble classifiers. BioMed Res Int. https://doi.org/10.1155/2013/239628
Dela Rosa JL, Magpantay AE, Gonzaga AC, Solano G (2014) Cluster center genes as candidate biomarkers for the classification of Leukemia. In: The 5th international conference on information, intelligence, systems and applications, IISA 2014. IEEE, pp 124–129
Akadi E, Ouardighi E (2009) A new gene selection approach based on minimum redundancy-maximum relevance (MRMR) and genetic algorithm (GA). In: 2009 IEEE/ACS international conference on computer systems and applications, pp 69–75
Alshamlan H, Badr G, Alohali Y (2015) mRMR-ABC: a hybrid gene selection algorithm for cancer classification using microarray gene expression profiling. BioMed Res Int. https://doi.org/10.1155/2015/604910
Alshamlan HM, Badr GH, Alohali YA (2014) The performance of bio-inspired evolutionary gene selection methods for cancer classification using microarray dataset. Int J Biosci Biochem Bioinform 4(3):166–170
Yu H, Ni J, Zhao J (2013) ACOSampling: an ant colony optimization-based undersampling method for classifying imbalanced DNA microarray data. Neurocomputing 101:309–318
Ghorai S, Mukherjee A, Sengupta S, Dutta PK (2011) Cancer classification from gene expression data by NPPC ensemble. IEEE/ACM Trans Comput Biol Bioinform (TCBB) 8(3):659–671
Sahu B, Mishra D (2012) A novel feature selection algorithm using particle swarm optimization for cancer microarray data. Procedia Eng 38:27–31
Danaee P, Ghaeini R, Hendrix DA (2017) A deep learning approach for cancer detection and relevant gene identification. In: Pacific symposium on biocomputing 2017. World Scientific, pp 219–229
Aziz R, Verma C, Jha M, Srivastava N (2017) Artificial neural network classification of microarray data using new hybrid gene selection method. Int J Data Min Bioinform 17(1):42–65
Hyvarinen A, Karhunen J, Oja E (2001) Independent component analysis. Wiley, London
Aziz R, Verma C, Srivastava N (2016) A fuzzy based feature selection from independent component subspace for machine learning classification of microarray data. Genom Data 8:4–15
Hsu C-C, Chen M-C, Chen L-S (2010) Integrating independent component analysis and support vector machine for multivariate process monitoring. Comput Ind Eng 59(1):145–156
Rabia A, Namita S, Chandan KV (2015) t-Independent component analysis for SVM classification of DNA- microarray data. Int J Bioinform Res 6(1):305–312
Hengpraprohm S (2013) GA-based classifier with SNR weighted features for cancer microarray data classification. Int J Signal Process Syst 1:29–33
Cesar I (2012) Feature selection using fuzzy entropy measures with Yu’s Similarity measure. Dissertation, Lappeenranta University of Technology
Lee H-M, Chen C-M, Chen J-M, Jou Y-L (2001) An efficient fuzzy classifier with feature selection based on fuzzy entropy. IEEE Trans Syst Man Cybern Part B Cybern 31(3):426–432
Huerta EB, Duval B, Hao J-K (2006) A hybrid GA/SVM approach for gene selection and classification of microarray data. In: Workshops on applications of evolutionary computation. Springer, pp 34–44
Karaboga D (2005) An idea based on honey bee swarm for numerical optimization. Technical report-tr06, Erciyes university, engineering faculty, computer engineering department
Garro BA, Rodríguez K, Vázquez RA (2015) Classification of DNA microarrays using artificial neural networks and ABC algorithm. Appl Soft Comput 38:548–560
Aziz R, Verma C, Srivastava N (2017) A novel approach for dimension reduction of microarray. Comput Biol Chem 71:161–169
Kıran MS, Özceylan E, Gündüz M, Paksoy T (2012) A novel hybrid approach based on particle swarm optimization and ant colony algorithm to forecast energy demand of Turkey. Energy Convers Manag 53(1):75–83
Jatoth RK, Rajasekhar A (2010) Speed control of pmsm by hybrid genetic artificial bee colony algorithm. In: 2010 IEEE international conference on communication control and computing technologies (ICCCCT). IEEE, pp 241–246
Alshamlan HM, Badr GH, Alohali YA (2015) Genetic bee colony (GBC) algorithm: a new gene selection method for microarray cancer classification. Comput Biol Chem 56:49–60
McCulloch WS, Pitts W (1943) A logical calculus of the ideas immanent in nervous activity. Bull Math Biophys 5(4):115–133
Cybenko G (1989) Approximation by superpositions of a sigmoidal function. Math Control Signals Syst 2(4):303–314
Rosenblatt F (1961) Principles of neurodynamics. Perceptrons and the theory of brain mechanisms. DTIC Document
Rumelhart DE, Hinton GE, Williams RJ (1985) Learning internal representations by error propagation. DTIC Document
Alon U, Barkai N, Notterman DA, Gish K, Ybarra S, Mack D, Levine AJ (1999) Broad patterns of gene expression revealed by clustering analysis of tumor and normal colon tissues probed by oligonucleotide arrays. Proc Natl Acad Sci 96(12):6745–6750
Golub TR, Slonim DK, Tamayo P, Huard C, Gaasenbeek M, Mesirov JP, Coller H, Loh ML, Downing JR, Caligiuri MA (1999) Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science 286(5439):531–537
Singh D, Febbo PG, Ross K, Jackson DG, Manola J, Ladd C, Tamayo P, Renshaw AA, D’Amico AV, Richie JP (2002) Gene expression correlates of clinical prostate cancer behavior. Cancer Cell 1(2):203–209
Gordon GJ, Jensen RV, Hsiao L-L, Gullans SR, Blumenstock JE, Ramaswamy S, Richards WG, Sugarbaker DJ, Bueno R (2002) Translation of microarray data into clinically relevant cancer diagnostic tests using gene expression ratios in lung cancer and mesothelioma. Cancer Res 62(17):4963–4967
Nutt CL, Mani D, Betensky RA, Tamayo P, Cairncross JG, Ladd C, Pohl U, Hartmann C, McLaughlin ME, Batchelor TT (2003) Gene expression-based classification of malignant gliomas correlates better with survival than histological classification. Cancer Res 63(7):1602–1607
Song B, Zhang G, Zhu W, Liang Z (2014) ROC operating point selection for classification of imbalanced data with application to computer-aided polyp detection in CT colonography. Int J Comput Assist Radiol Surg 9(1):79–89
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
All authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Aziz, R., Verma, C.K. & Srivastava, N. Artificial Neural Network Classification of High Dimensional Data with Novel Optimization Approach of Dimension Reduction. Ann. Data. Sci. 5, 615–635 (2018). https://doi.org/10.1007/s40745-018-0155-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40745-018-0155-2