Abstract
Background and Objective
Long non-coding RNAs (lncRNAs) may serve as biomarkers for complex disease states, such as intracranial aneurysms. In this study, we investigated lncRNA expression differences in the whole blood of patients with unruptured aneurysms.
Methods
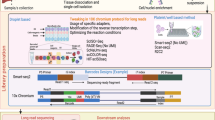
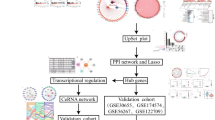
Whole blood RNA from 67 subjects (34 with aneurysm, 33 without) was used for next-generation RNA sequencing. Differential expression analysis was used to define a signature of intracranial aneurysm-associated lncRNAs. To estimate the signature’s ability to classify aneurysms and to identify the most predictive lncRNAs, we implemented a nested cross-validation pipeline to train classifiers using linear discriminant analysis. Ingenuity pathway analysis was used to study potential biological roles of differentially expressed lncRNAs, and lncRNA ontology was used to investigate ontologies enriched in signature lncRNAs. Co-expression correlation analysis was performed to investigate associated differential protein-coding messenger RNA expression.
Results
Of 4639 detected lncRNAs, 263 were significantly different (p < 0.05) between the two groups, and 84 of those had an absolute fold-change ≥ 1.5. An eight-lncRNA signature (q < 0.35, fold-change ≥ 1.5) was able to separate patients with and without aneurysms on principal component analysis, and had an estimated accuracy of 70.9% in nested cross-validation. Bioinformatics analyses showed that networks of differentially expressed lncRNAs (p < 0.05) were enriched for cell death and survival, connective tissue disorders, carbohydrate metabolism, and cardiovascular disease. Signature lncRNAs shared ontologies that reflected regulation of gene expression, signaling, ubiquitin processing, and p53 signaling. Co-expression analysis showed correlations with messenger RNAs related to inflammatory responses.
Conclusions
Differential expression in whole blood lncRNAs is detectable in patients harboring aneurysms, and reflects expression/signaling regulation, and ubiquitin and p53 pathways. Following validation in larger cohorts, these lncRNAs may be potential diagnostic targets for aneurysm detection by blood testing.





Similar content being viewed by others
References
Wiebers DO, Whisnant JP, Huston J III, International Study of Unruptured Intracranial Aneurysms Investigators, et al. Unruptured intracranial aneurysms: natural history, clinical outcome, and risks of surgical and endovascular treatment. Lancet. 2003;362(9378):103–10.
Go AS, Mozaffarian D, Roger VL, Benjamin EJ, Berry JD, Blaha MJ, et al. Executive summary: heart disease and stroke statistics: 2014 update: a report from the American Heart Association. Circulation. 2014;129(3):399–410.
Rinkel GJ, Djibuti M, Algra A, Van Gijn J. Prevalence and risk of rupture of intracranial aneurysms: a systematic review. Stroke. 1998;29(1):25–6.
Thompson BG, Brown RD Jr, Amin-Hanjani S, Broderick JP, Cockroft KM, Connolly ES Jr, et al. Guidelines for the management of patients with unruptured intracranial aneurysms: a guideline for healthcare professionals from the American Heart Association/American Stroke Association. Stroke. 2015;46(8):2368–400.
Power SP, Moloney F, Twomey M, James K, O’Connor OJ, Maher MM. Computed tomography and patient risk: facts, perceptions and uncertainties. World J Radiol. 2016;8(12):902–15.
Doyle J, Abraham S, Feeney L, Reimer S, Finkelstein A. Clinical decision support for high-cost imaging: a randomized clinical trial. PLoS ONE. 2019;14(3):e0213373.
Derrien T, Johnson R, Bussotti G, Tanzer A, Djebali S, Tilgner H, et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res. 2012;22(9):1775–899.
Fernandes JCR, Acuña SM, Aoki JI, Floeter-Winter LM, Muxel SM. Long non-coding RNAs in the regulation of gene expression: physiology and disease. Noncoding RNA. 2019;5(1):17.
Deniz E, Erman B. Long noncoding RNA (lincRNA), a new paradigm in gene expression control. Funct Integr Genom. 2017;17(2):135–43.
Jaé N, Dimmeler S. Noncoding RNAs in vascular diseases. Circ Res. 2020;126(9):1127–45.
Hung J, Miscianinov V, Sluimer JC, Newby DE, Baker AH. Targeting non-coding RNA in vascular biology and disease. Front Physiol. 2018;9:1655.
Huang F, Yi J, Zhou T, Gong X, Jiang H, Yao X. Toward understanding non-coding RNA roles in intracranial aneurysms and subarachnoid hemorrhage. Transl Neurosci. 2017;8:54–64.
Li H, Wang W, Zhang L, Lan Q, Wang J, Cao Y, et al. Identification of a long noncoding RNA-associated competing endogenous RNA network in intracranial aneurysm. World Neurosurg. 2017;97:684–92 (e4).
Li H, Yue H, Hao Y, Li H, Wang S, Yu L, et al. Expression profile of long noncoding RNAs in human cerebral aneurysms: a microarray analysis. J Neurosurg. 2017;127(5):1055–62.
Wang W, Li H, Yu L, Zhao Z, Wang H, Zhang D, et al. Aberrant expression of lncRNAs and mRNAs in patients with intracranial aneurysm. Oncotarget. 2017;8(2):2477–84.
Wu C, Song H, Wang Y, Gao L, Cai Y, Cheng Q, et al. Long non-coding RNA TCONS_00000200 as a non-invasive biomarker in patients with intracranial aneurysm. Biosci Rep. 2019;39(11):BSR20182224.
Robinson MD, McCarthy DJ, Smyth GK. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2009;26(1):139–40.
McCarthy DJ, Chen Y, Smyth GK. Differential expression analysis of multifactor RNA-seq experiments with respect to biological variation. Nucleic Acids Res. 2012;40(10):4288–97.
Hart SN, Therneau TM, Zhang Y, Poland GA, Kocher J-P. Calculating sample size estimates for RNA sequencing data. J Comput Biol. 2013;20(12):970–8.
Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B Stat Methodol. 1995;57(1):289–300.
Damiano RJ. Predicting treatment outcomes of coiled intracranial aneurysms using finite element modeling and machine learning. Buffalo: State University of New York at Buffalo; 2020.
Stone M. Cross-validatory choice and assessment of statistical predictions. J R Stat Soc Ser B Stat Methodol. 1974;36(2):111–33.
Krämer A, Green J, Pollard J Jr, Tugendreich S. Causal analysis approaches in ingenuity pathway analysis. Bioinformatics. 2014;30(4):523–30.
Li J-P, Liu L-H, Li J, Chen Y, Jiang X-W, Ouyang Y-R, et al. Microarray expression profile of long noncoding RNAs in human osteosarcoma. Biochem Biophys Res Commun. 2013;433(2):200–6.
Niebler M, Qian X, Höfler D, Kogosov V, Kaewprag J, Kaufmann AM, et al. Post-translational control of IL-1β via the human papillomavirus type 16 E6 oncoprotein: a novel mechanism of innate immune escape mediated by the E3-ubiquitin ligase E6-AP and p53. PLoS Pathog. 2013;9(8):e1003536.
Tutino VM, Poppenberg KE, Jiang K, Jarvis JN, Sun Y, Sonig A, et al. Circulating neutrophil transcriptome may reveal intracranial aneurysm signature. PLoS ONE. 2018;13(1):e0191407.
Tutino VM, Poppenberg KE, Li L, Shallwani H, Jiang K, Jarvis JN, et al. Biomarkers from circulating neutrophil transcriptomes have potential to detect unruptured intracranial aneurysms. J Transl Med. 2018;16(1):373.
Wang KC, Chang HY. Molecular mechanisms of long noncoding RNAs. Mol Cell. 2011;43(6):904–14.
Simion V, Haemmig S, Feinberg MW. LncRNAs in vascular biology and disease. Vasc Pharmacol. 2019;114:145–56.
Schlosser K, Hanson J, Villeneuve PJ, Dimitroulakos J, McIntyre L, Pilote L, et al. Assessment of circulating lncRNAs under physiologic and pathologic conditions in humans reveals potential limitations as biomarkers. Sci Rep. 2016;6(1):36596.
Tsai M-C, Spitale RC, Chang HY. Long intergenic noncoding RNAs: new links in cancer progression. Cancer Res. 2011;71(1):3–7.
Ransohoff JD, Wei Y, Khavari PA. The functions and unique features of long intergenic non-coding RNA. Nat Rev Mol Cell Biol. 2018;19(3):143–57.
Moriwaki T, Takagi Y, Sadamasa N, Aoki T, Nozaki K, Hashimoto N. Impaired progression of cerebral aneurysms in interleukin-1beta-deficient mice. Stroke. 2006;37(3):900–5.
Zhang HF, Zhao MG, Liang GB, Song ZQ, Li ZQ. Expression of pro-inflammatory cytokines and the risk of intracranial aneurysm. Inflammation. 2013;36(6):1195–200.
Leeper NJ, Raiesdana A, Kojima Y, Kundu Ramendra K, Cheng H, Maegdefessel L, et al. Loss of CDKN2B promotes p53-dependent smooth muscle cell apoptosis and aneurysm formation. Arterioscler Thromb Vasc Biol. 2013;33(1):e1–.
Meng H, Tutino VM, Xiang J, Siddiqui A. High WSS or low WSS? Complex interactions of hemodynamics with intracranial aneurysm initiation, growth, and rupture: toward a unifying hypothesis. Am J Neuroradiol. 2014;35(7):1254–62.
Ouimet M, Drouin S, Lajoie M, Caron M, St-Onge P, Gioia R, et al. A childhood acute lymphoblastic leukemia-specific lncRNA implicated in prednisolone resistance, cell proliferation, and migration. Oncotarget. 2017;8(5):7477–88.
Yang X, Yang J, Wang J, Wen Q, Wang H, He J, et al. Microarray analysis of long noncoding RNA and mRNA expression profiles in human macrophages infected with Mycobacterium tuberculosis. Sci Rep. 2016;6(1):38963.
Riege K, Hölzer M, Klassert TE, Barth E, Bräuer J, Collatz M, et al. Massive effect on lncRNAs in human monocytes during fungal and bacterial infections and in response to vitamins A and D. Sci Rep. 2017;7:40598.
Valadkhan S, Gunawardane LS. lncRNA-mediated regulation of the interferon response. Virus Res. 2016;2(212):127–36.
Mariotti B, Servaas NH, Rossato M, Tamassia N, Cassatella MA, Cossu M, et al. The long non-coding RNA NRIR drives IFN-response in monocytes: implication for systemic sclerosis. Front Immunol. 2019;10:100.
Hussain S, Barbarite E, Chaudhry NS, Gupta K, Dellarole A, Peterson EC, et al. Search for biomarkers of intracranial aneurysms: a systematic review. World Neurosurg. 2015;84(5):1473–83.
Sandalcioglu IE, Wende D, Eggert A, Regel JP, Stolke D, Wiedemayer H. VEGF plasma levels in non-ruptured intracranial aneurysms. Neurosurg Rev. 2006;29(1):26–9.
Phillips J, Roberts G, Bolger C, el Baghdady A, Bouchier-Hayes D, Farrell M, et al. Lipoprotein (a): a potential biological marker for unruptured intracranial aneurysms. Neurosurgery. 1997;40(5):1112–5 (discussion 5–7).
Baker CJ, Fiore A, Connolly ES Jr, Baker KZ, Solomon RA. Serum elastase and alpha-1-antitrypsin levels in patients with ruptured and unruptured cerebral aneurysms. Neurosurgery. 1995;37(1):56–61 (discussion 61–2).
Chalouhi N, Theofanis T, Starke RM, Zanaty M, Jabbour P, Dooley SA, et al. Potential role of granulocyte-monocyte colony-stimulating factor in the progression of intracranial aneurysms. DNA Cell Biol. 2015;34(1):78–81.
Chen J, Han L, Xu X, Tang H, Wang H, Wei B. Serum biomarkers VEGF-C and IL-6 are associated with severe human peripheral artery stenosis. J Inflamm. 2015;12:50.
Kimura H, Okada O, Tanabe N, Tanaka Y, Terai M, Takiguchi Y, et al. Plasma monocyte chemoattractant protein-1 and pulmonary vascular resistance in chronic thromboembolic pulmonary hypertension. Am J Respir Crit Care Med. 2001;164(2):319–24.
Ray L, Khemka VK, Behera P, Bandyopadhyay K, Pal S, Pal K, et al. Serum homocysteine, dehydroepiandrosterone sulphate and lipoprotein (a) in Alzheimer's disease and vascular dementia. Aging Dis. 2013;4(2):57–64.
Sabatino G, Rigante L, Minella D, Novelli G, Della Pepa GM, Esposito G, et al. Transcriptional profile characterization for the identification of peripheral blood biomarkers in patients with cerebral aneurysms. J Biol Regul Homeost Agents. 2013;27(3):729–38.
Li P, Zhang Q, Wu X, Yang X, Zhang Y, Li Y, et al. Circulating microRNAs serve as novel biological markers for intracranial aneurysms. J Am Heart Assoc. 2014;3(5):e000972.
Jin H, Li C, Ge H, Jiang Y, Li Y. Circulating microRNA: a novel potential biomarker for early diagnosis of intracranial aneurysm rupture a case control study. J Transl Med. 2013;11:296.
Meeuwsen JAL, van’t Hof FNG, van Rheenen W, Rinkel GJE, Veldink JH, Ruigrok YM. Circulating microRNAs in patients with intracranial aneurysms. PLoS ONE. 2017;12(5):e0176558.
Zhao M, Xu L, Qian H. Bioinformatics analysis of microRNA profiles and identification of microRNA-mRNA network and biological markers in intracranial aneurysm. Medicine (Baltimore). 2020;99(31):e21186.
Acknowledgements
We thank the patients who participated in this study. We acknowledge Jonathan Bard, MA and Brandon Marzullo, MS for RNA sequencing data analysis assistance, and Jennifer L. Gay, CCRP for study protocol management. This work was performed in part at the New York State Center of Excellence in Bioinformatics and Life Sciences’ Genomics and Bioinformatics Core.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This work was funded by the Brain Aneurysm Foundation (VMT), the New York State Center for Advanced Technology in Big Data and Health Sciences (VMT), and the Cummings Foundation (VMT).
Conflict of interest
Vincent M. Tutino is the principal investigator for National Science Foundation Award No. 1746694 and NIH NINDS award R43 NS115314-0, awardee of the abovementioned Brain Aneurysm Foundation grant, Center for Advanced Technology grant, and Cummings Foundation grant, and a co-founder of Neurovascular Diagnostics, Inc. Kerry E. Poppenberg, Robert J. Damiano, Tatsat R. Patel, Muhammad Waqas, and Adam A. Dmytriw have no conflicts of interest that are directly relevant to the content of this article. Kenneth V. Snyder is a consultant and teacher for Canon Medical Systems Corporation, Penumbra Inc., Medtronic, and Jacobs Institute and a co-founder of Neurovascular Diagnostics, Inc. Adnan H. Siddiqui has a financial interest/stock options/ownership/investment in Adona Medical, Inc., Amnis Therapeutics, BlinkTBI, Inc, Boston Scientific Corp (for purchase of Claret Medical), Buffalo Technology Partners, Inc., Cardinal Consultants, LLC, Cerebrotech Medical Systems, Inc, Cognition Medical, Endostream Medical, Ltd, Imperative Care, Inc., International Medical Distribution Partners, Neurovascular Diagnostics, Inc., Q’Apel Medical, Inc., Radical Catheter Technologies, Inc., Rebound Therapeutics Corp. (purchased 2019 by Integra Lifesciences, Corp), Rist Neurovascular, Inc., Sense Diagnostics, Inc., Serenity Medical, Inc., Silk Road Medical, Spinnaker Medical, Inc., StimMed, Synchron, Three Rivers Medical, Inc., Vastrax, LLC, VICIS, Inc., and Viseon, Inc.; is a consultant/advisory board member for Amnis Therapeutics, Boston Scientific, Canon Medical Systems USA, Inc., Cerebrotech Medical Systems, Inc., Cerenovus, Corindus, Inc., Endostream Medical, Ltd, Imperative Care, Inc., Integra LifeSciences Corp., Medtronic, MicroVention, Minnetronix Neuro, Inc., Northwest University – DSMB Chair for HEAT Trial, Penumbra, Q’Apel Medical, Inc., Rapid Medical, Rebound Therapeutics Corp., Serenity Medical, Inc., Silk Road Medical, StimMed, Stryker, Three Rivers Medical, Inc., VasSol, and W.L. Gore & Associates; is a national principal investigator/steering committee member for Cerenovus LARGE Trial and ARISE II Trial, Medtronic SWIFT PRIME and SWIFT DIRECT Trials, MicroVention FRED Trial & CONFIDENCE Study, MUSC POSITIVE Trial, Penumbra 3D Separator Trial, COMPASS Trial, and INVEST Trial; has received research grants/co-investigator for NIH/NINDS 1R01NS091075 Virtual Intervention of Intracranial Aneurysms; and is a co-principal investigator for NIH-NINDS R21 NS109575-01 Optimizing Approaches to Endovascular Therapy of Acute Ischemic Stroke. James N. Jarvis is the principal investigator for NIH Grant R01-AR-060604.
Ethics approval
This study was approved by the University at Buffalo Institutional Review Board (no. 030-474433).
Consent to participate
All subjects provided written informed consent before participating in this study, as described in the approved protocol.
Consent for publication
Not applicable.
Availability of data and material
Raw data are available upon reasonable request to the corresponding author.
Code availability
Not applicable.
Author contributions
VMT and KEP conceived and designed the research. VMT, KEP, MW, KVS, and AHS collected and reviewed the data. VMT, KEP, RJD, and TRP analyzed the data and performed the statistical analysis. VMT handled the funding and supervision of the research. VMT, KEP, RJD, and JNJ drafted the manuscript. VMT, KEP, RJD, TRP, MW, AAD, KVS, AHS, and JNJ revised the manuscript and reviewed the final version.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Tutino, V.M., Poppenberg, K.E., Damiano, R.J. et al. Characterization of Long Non-coding RNA Signatures of Intracranial Aneurysm in Circulating Whole Blood. Mol Diagn Ther 24, 723–736 (2020). https://doi.org/10.1007/s40291-020-00494-3
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40291-020-00494-3