Abstract
Purpose
Tumor microenvironment (TME) affects the progression of rectal cancer (RC), and the clinical relevance of its immune elements was widely reported. Here we aim to delineate the complete TME landscape, including non-immune features, to improve our understanding of RC heterogeneity and provide a better strategy for precision medicine.
Methods
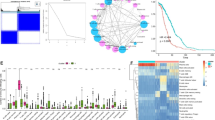
Single-cell analysis of GSE161277 using Seurat and Cellcall was performed to identify cell-cell interactions. The ssGSEA was employed to quantify the TME elements in TCGA patients, which were further clustered into subtypes by hclust. WGCNA and LASSO were combined to construct a degenerated signature for prognosis, and its performance was validated in two GEO datasets.
Results
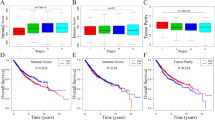
We proposed a subtyping strategy based on the abundance of both immune and non-immune components, which divided all RC patients into 4 subtypes (Immune-, Canonical-, Dormant- and Stem-like). Different subtypes exhibited distinct mutation landscapes, biological features, immune characteristics, immunotherapy responses and prognoses. Next, WGCNA and LASSO regression were combined to construct a 10-gene signature based on differentially expressed genes among different subtypes. Subgroups divided by this signature also exhibited different clinical parameters and responses to immune checkpoint blockades. Diverse machine learning algorithms were applied to achieve higher accuracy for survival prediction and a nomogram was further established in combination with M stage and age to provide an accurate and visual prediction of prognosis.
Conclusions
We identified four TME-based RC subtypes with distinct biological and clinical features. Based on those subtypes, we also proposed a degenerated 10-gene signature to predict the prognosis and immunotherapy response.








Similar content being viewed by others
Data availability
All sequencing data were public available datasets (TCGA https://portal.gdc.cancer.gov/cart and GEO https://www.ncbi.nlm.nih.gov/geo/). All other data supporting the conclusions of this article are presented within the article and its supplementary files.
References
H. Sung, J. Ferlay, R.L. Siegel, M. Laversanne, I. Soerjomataram, A. Jemal, F. Bray, Global Cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. (2021). https://doi.org/10.3322/caac.21660
D.S. Keller, M. Berho, R.O. Perez, S.D. Wexner, M. Chand, The multidisciplinary management of rectal cancer. Nat. Rev. Gastroenterol. Hepatol. (2020). https://doi.org/10.1038/s41575-020-0275-y
W.J. Tan, H.J. Tan, S.R. Dorajoo, F.J. Foo, C.L. Tang, M.H. Chew, Rectal Cancer surveillance-recurrence patterns and survival outcomes from a cohort followed up beyond 10 years. J. Gastrointest. Cancer (2018). https://doi.org/10.1007/s12029-017-9984-z
R. Braun, L. Anthuber, D. Hirsch, D. Wangsa, J. Lack, N.E. McNeil, K. Heselmeyer-Haddad, I. Torres, D. Wangsa, M.A. Brown, A. Tubbs, N. Auslander, E.M. Gertz, P.R. Brauer, M.C. Cam, D.L. Sackett, J.K. Habermann, A. Nussenzweig, E. Ruppin, et al., Single-cell-derived primary rectal carcinoma cell lines reflect Intratumor heterogeneity associated with treatment response. Clin. Cancer Res. (2020). https://doi.org/10.1158/1078-0432.CCR-19-1984
Z. Lakkis, G. Manceau, V. Bridoux, A. Brouquet, S. Kirzin, L. Maggiori, C. de Chaisemartin, J.H. Lefevre, Y. Panis, Management of rectal cancer: The 2016 French guidelines. Color. Dis. (2017). https://doi.org/10.1111/codi.13550
N.M. Anderson, M.C. Simon, The tumor microenvironment. Curr. Biol. (2020). https://doi.org/10.1016/j.cub.2020.06.081
I. Vitale, G. Manic, L.M. Coussens, G. Kroemer, L. Galluzzi, Macrophages and metabolism in the tumor microenvironment. Cell Metab. (2019). https://doi.org/10.1016/j.cmet.2019.06.001
M. Saxena, N. Bhardwaj, Re-emergence of dendritic cell vaccines for Cancer treatment. Trends Cancer (2018). https://doi.org/10.1016/j.trecan.2017.12.007
J.J. Loh, S. Ma, The role of Cancer-associated fibroblast as a dynamic player in mediating Cancer Stemness in the tumor microenvironment. Front. Cell Dev. Biol. (2021). https://doi.org/10.3389/fcell.2021.727640
S. Sui, X. An, C. Xu, Z. Li, Y. Hua, G. Huang, S. Sui, Q. Long, Y. Sui, Y. Xiong, M. Ntim, W. Guo, M. Chen, W. Deng, X. Xiao, M. Li, An immune cell infiltration-based immune score model predicts prognosis and chemotherapy effects in breast cancer. Theranostics (2020). https://doi.org/10.7150/thno.49451
W. Jiang, D. Zhu, C. Wang, Y. Zhu, An immune relevant signature for predicting prognoses and immunotherapeutic responses in patients with muscle-invasive bladder cancer (MIBC). Cancer Med. (2020). https://doi.org/10.1002/cam4.2942
R. Zhou, J. Zhang, D. Zeng, H. Sun, X. Rong, M. Shi, J. Bin, Y. Liao, W. Liao, Immune cell infiltration as a biomarker for the diagnosis and prognosis of stage I-III colon cancer. Cancer Immunol. Immunother. (2019). https://doi.org/10.1007/s00262-018-2289-7
P. Ge, W. Wang, L. Li, G. Zhang, Z. Gao, Z. Tang, X. Dang, Y. Wu, Profiles of immune cell infiltration and immune-related genes in the tumor microenvironment of colorectal cancer. Biomed. Pharmacother. (2019). https://doi.org/10.1016/j.biopha.2019.109228
M. Binnewies, E.W. Roberts, K. Kersten, V. Chan, D.F. Fearon, M. Merad, L.M. Coussens, D.I. Gabrilovich, S. Ostrand-Rosenberg, C.C. Hedrick, R.H. Vonderheide, M.J. Pittet, R.K. Jain, W. Zou, T.K. Howcroft, E.C. Woodhouse, R.A. Weinberg, M.F. Krummel, Understanding the tumor immune microenvironment (TIME) for effective therapy. Nat. Med. (2018). https://doi.org/10.1038/s41591-018-0014-x
Z. Wu, K. Zhu, Q. Liu, Y. Liu, L. Chen, J. Cui, H. Guo, N. Zhou, Y. Zhu, Y. Li, B. Shi, Profiles of immune infiltration in bladder Cancer and its clinical significance: An integrative genomic analysis. Int. J. Med. Sci. (2020). https://doi.org/10.7150/ijms.42151
J. Liu, Z. Tan, J. He, T. Jin, Y. Han, L. Hu, J. Song, S. Huang, Identification of three molecular subtypes based on immune infiltration in ovarian cancer and its prognostic value. Biosci. Rep. (2020). https://doi.org/10.1042/BSR20201431
P. Hu, Y. Gao, Y. Huang, Y. Zhao, H. Yan, J. Zhang, L. Zhao, Gene expression-based immune cell infiltration analyses of prostate Cancer and their associations with survival outcome. DNA Cell Biol. (2020). https://doi.org/10.1089/dna.2020.5371
S. Yang, T. Liu, Y. Cheng, Y. Bai, G. Liang, Immune cell infiltration as a biomarker for the diagnosis and prognosis of digestive system cancer. Cancer Sci. (2019). https://doi.org/10.1111/cas.14216
X. Liu, S. Wu, Y. Yang, M. Zhao, G. Zhu, Z. Hou, The prognostic landscape of tumor-infiltrating immune cell and immunomodulators in lung cancer. Biomed. Pharmacother. (2017). https://doi.org/10.1016/j.biopha.2017.08.003
N. Kim, H.K. Kim, K. Lee, Y. Hong, J.H. Cho, J.W. Choi, J.I. Lee, Y.L. Suh, B.M. Ku, H.H. Eum, S. Choi, Y.L. Choi, J.G. Joung, W.Y. Park, H.A. Jung, J.M. Sun, S.H. Lee, J.S. Ahn, K. Park, et al., Single-cell RNA sequencing demonstrates the molecular and cellular reprogramming of metastatic lung adenocarcinoma. Nat. Commun. (2020). https://doi.org/10.1038/s41467-020-16164-1
L.M. Becker, J.T. O'Connell, A.P. Vo, M.P. Cain, D. Tampe, L. Bizarro, H. Sugimoto, A.K. McGow, J.M. Asara, S. Lovisa, K.M. McAndrews, R. Zielinski, P.L. Lorenzi, M. Zeisberg, S. Raza, V.S. LeBleu, R. Kalluri, Epigenetic reprogramming of Cancer-associated fibroblasts deregulates glucose metabolism and facilitates progression of breast Cancer. Cell Rep. (2020). https://doi.org/10.1016/j.celrep.2020.107701
A. Butler, P. Hoffman, P. Smibert, E. Papalexi, R. Satija, Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. (2018). https://doi.org/10.1038/nbt.4096
M.C. Cieslak, A.M. Castelfranco, V. Roncalli, P.H. Lenz, D.K. Hartline, T-distributed stochastic neighbor embedding (t-SNE): A tool for eco-physiological transcriptomic analysis. Mar. Genomics (2020). https://doi.org/10.1016/j.margen.2019.100723
Q. Huang, Y. Liu, Y. Du, L.X. Garmire, Evaluation of cell type annotation R packages on single-cell RNA-seq data. Genomics Proteomics Bioinformatics (2021). https://doi.org/10.1016/j.gpb.2020.07.004
Y. Zhang, T. Liu, X. Hu, M. Wang, J. Wang, B. Zou, P. Tan, T. Cui, Y. Dou, L. Ning, Y. Huang, S. Rao, D. Wang, X. Zhao, CellCall: Integrating paired ligand-receptor and transcription factor activities for cell-cell communication. Nucleic Acids Res. (2021). https://doi.org/10.1093/nar/gkab638
S. Mariathasan, S.J. Turley, D. Nickles, A. Castiglioni, K. Yuen, Y. Wang, E.I. Kadel, H. Koeppen, J.L. Astarita, R. Cubas, S. Jhunjhunwala, R. Banchereau, Y. Yang, Y. Guan, C. Chalouni, J. Ziai, Y. Senbabaoglu, S. Santoro, D. Sheinson, et al., TGFbeta attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature (2018). https://doi.org/10.1038/nature25501
P. Charoentong, F. Finotello, M. Angelova, C. Mayer, M. Efremova, D. Rieder, H. Hackl, Z. Trajanoski, Pan-cancer Immunogenomic analyses reveal genotype-Immunophenotype relationships and predictors of response to checkpoint blockade. Cell Rep. (2017). https://doi.org/10.1016/j.celrep.2016.12.019
G. Bindea, B. Mlecnik, M. Tosolini, A. Kirilovsky, M. Waldner, A.C. Obenauf, H. Angell, T. Fredriksen, L. Lafontaine, A. Berger, P. Bruneval, W.H. Fridman, C. Becker, F. Pages, M.R. Speicher, Z. Trajanoski, J. Galon, Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer. Immunity (2013). https://doi.org/10.1016/j.immuni.2013.10.003
X. Zhang, Y. Lan, J. Xu, F. Quan, E. Zhao, C. Deng, T. Luo, L. Xu, G. Liao, M. Yan, Y. Ping, F. Li, A. Shi, J. Bai, T. Zhao, X. Li, Y. Xiao, CellMarker: A manually curated resource of cell markers in human and mouse. Nucleic Acids Res. (2019). https://doi.org/10.1093/nar/gky900
S. Hanzelmann, R. Castelo, J. Guinney, GSVA: Gene set variation analysis for microarray and RNA-seq data. Bmc Bioinformatics (2013). https://doi.org/10.1186/1471-2105-14-7
J. Wu, L. Li, H. Zhang, Y. Zhao, H. Zhang, S. Wu, B. Xu, A risk model developed based on tumor microenvironment predicts overall survival and associates with tumor immunity of patients with lung adenocarcinoma. Oncogene (2021). https://doi.org/10.1038/s41388-021-01853-y
R. Cao, B. Ma, G. Wang, Y. Xiong, Y. Tian, L. Yuan, Characterization of hypoxia response patterns identified prognosis and immunotherapy response in bladder cancer. Mol. Ther. Oncolytics (2021). https://doi.org/10.1016/j.omto.2021.06.011
P. Langfelder, S. Horvath, WGCNA: An R package for weighted correlation network analysis. Bmc Bioinformatics (2008). https://doi.org/10.1186/1471-2105-9-559
T. Wu, E. Hu, S. Xu, M. Chen, P. Guo, Z. Dai, T. Feng, L. Zhou, W. Tang, L. Zhan, X. Fu, S. Liu, X. Bo, G. Yu, clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation (Camb.) (2021). https://doi.org/10.1016/j.xinn.2021.100141
P. Geeleher, N. Cox, R.S. Huang, pRRophetic: An R package for prediction of clinical chemotherapeutic response from tumor gene expression levels. PLoS One (2014). https://doi.org/10.1371/journal.pone.0107468
D. Sun, Y. Zhu, H. Zhao, T. Bian, T. Li, K. Liu, L. Feng, H. Li, H. Hou, Loss of ARID1A expression promotes lung adenocarcinoma metastasis and predicts a poor prognosis. Cell Oncol. (Dordr.) (2021). https://doi.org/10.1007/s13402-021-00616-x
Y. Chen, J. Zhao, Identification of an immune gene signature based on tumor microenvironment characteristics in Colon adenocarcinoma. Cell Transplant. (2021). https://doi.org/10.1177/09636897211001314
Y. Zhao, C. Chen, X. Xu, X. Ge, K. Ding, S. Zheng, J. Wang, L. Sun, An efficient prognostic immune scoring system for colorectal Cancer patients with peritoneal metastasis. Oncoimmunology (2021). https://doi.org/10.1080/2162402X.2021.1901464
W. Chong, L. Shang, J. Liu, Z. Fang, F. Du, H. Wu, Y. Liu, Z. Wang, Y. Chen, S. Jia, L. Chen, L. Li, H. Chen, M(6)a regulator-based methylation modification patterns characterized by distinct tumor microenvironment immune profiles in colon cancer. Theranostics (2021). https://doi.org/10.7150/thno.52717
J. Rao, W. Li, C. Chen, Pyroptosis-mediated molecular subtypes and tumor microenvironment infiltration characterization in Colon Cancer. Front. Cell Dev. Biol. (2021). https://doi.org/10.3389/fcell.2021.766503
W. Song, J. Ren, R. Xiang, C. Kong, T. Fu, Identification of pyroptosis-related subtypes, the development of a prognosis model, and characterization of tumor microenvironment infiltration in colorectal cancer. Oncoimmunology (2021). https://doi.org/10.1080/2162402X.2021.1987636
X. Zhu, X. Tian, L. Ji, X. Zhang, Y. Cao, C. Shen, Y. Hu, J. Wong, J.Y. Fang, J. Hong, H. Chen, A tumor microenvironment-specific gene expression signature predicts chemotherapy resistance in colorectal cancer patients. NPJ Precis. Oncol. (2021). https://doi.org/10.1038/s41698-021-00142-x
C.A. Doubeni, D.A. Corley, V.P. Quinn, C.D. Jensen, A.G. Zauber, M. Goodman, J.R. Johnson, S.J. Mehta, T.A. Becerra, W.K. Zhao, J. Schottinger, V.P. Doria-Rose, T.R. Levin, N.S. Weiss, R.H. Fletcher, Effectiveness of screening colonoscopy in reducing the risk of death from right and left colon cancer: A large community-based study. Gut (2018). https://doi.org/10.1136/gutjnl-2016-312712
R. Yaeger, W.K. Chatila, M.D. Lipsyc, J.F. Hechtman, A. Cercek, F. Sanchez-Vega, G. Jayakumaran, S. Middha, A. Zehir, M. Donoghue, D. You, A. Viale, N. Kemeny, N.H. Segal, Z.K. Stadler, A.M. Varghese, R. Kundra, J. Gao, A. Syed, et al., Clinical sequencing defines the genomic landscape of metastatic colorectal Cancer. Cancer Cell (2018). https://doi.org/10.1016/j.ccell.2017.12.004
S. Talukdar, P. Bhoopathi, L. Emdad, S. Das, D. Sarkar, P.B. Fisher, Dormancy and cancer stem cells: An enigma for cancer therapeutic targeting. Adv. Cancer Res. (2019). https://doi.org/10.1016/bs.acr.2018.12.002
F. Liu, W. Hou, J. Liang, L. Zhu, C. Luo, LRP1B mutation: A novel independent prognostic factor and a predictive tumor mutation burden in hepatocellular carcinoma. J. Cancer (2021). https://doi.org/10.7150/jca.53124
E. Tabouret, M. Labussiere, A. Alentorn, Y. Schmitt, Y. Marie, M. Sanson, LRP1B deletion is associated with poor outcome for glioblastoma patients. J. Neurol. Sci. (2015). https://doi.org/10.1016/j.jns.2015.09.345
P. Li, J. Xiao, B. Zhou, J. Wei, J. Luo, W. Chen, SYNE1 mutation may enhance the response to immune checkpoint blockade therapy in clear cell renal cell carcinoma patients. Aging (Albany NY) (2020). https://doi.org/10.18632/aging.103781
Q. Luo, D. Chen, X. Fan, X. Fu, T. Ma, D. Chen, KRAS and PIK3CA bi-mutations predict a poor prognosis in colorectal cancer patients: A single-site report. Transl. Oncol. (2020). https://doi.org/10.1016/j.tranon.2020.100874
S. Jang, M. Hong, M.K. Shin, B.C. Kim, H.S. Shin, E. Yu, S.M. Hong, J. Kim, S.M. Chun, T.I. Kim, K.C. Choi, Y.W. Ko, J.W. Kim, KRAS and PIK3CA mutations in colorectal adenocarcinomas correlate with aggressive histological features and behavior. Hum. Pathol. (2017). https://doi.org/10.1016/j.humpath.2017.01.010
D.S. Chen, I. Mellman, Elements of cancer immunity and the cancer-immune set point. Nature (2017). https://doi.org/10.1038/nature21349
Y. Chen, Z. Sun, W. Chen, C. Liu, R. Chai, J. Ding, W. Liu, X. Feng, J. Zhou, X. Shen, S. Huang, Z. Xu, The immune subtypes and landscape of gastric Cancer and to predict based on the whole-slide images using deep learning. Front. Immunol. (2021). https://doi.org/10.3389/fimmu.2021.685992
C. Tang, J. Ma, X. Liu, Z. Liu, Identification of four immune subtypes in bladder Cancer based on immune gene sets. Front. Oncol. (2020). https://doi.org/10.3389/fonc.2020.544610
M. Sasaki, N. Miyoshi, S. Fujino, K. Saso, T. Ogino, H. Takahashi, M. Uemura, H. Yamamoto, C. Matsuda, M. Yasui, M. Ohue, T. Mizushima, Y. Doki, H. Eguchi, The meiosis-specific cohesin component stromal antigen 3 promotes cell migration and chemotherapeutic resistance in colorectal cancer. Cancer Lett. (2021). https://doi.org/10.1016/j.canlet.2020.10.006
J.M. Kraveka, L. Li, Z.M. Szulc, J. Bielawski, B. Ogretmen, Y.A. Hannun, L.M. Obeid, A. Bielawska, Involvement of dihydroceramide desaturase in cell cycle progression in human neuroblastoma cells. J. Biol. Chem. (2007). https://doi.org/10.1074/jbc.M700647200
P. Munoz-Guardiola, J. Casas, E. Megias-Roda, S. Sole, H. Perez-Montoyo, M. Yeste-Velasco, T. Erazo, N. Dieguez-Martinez, S. Espinosa-Gil, C. Munoz-Pinedo, G. Yoldi, J.L. Abad, M.F. Segura, T. Moran, M. Romeo, J. Bosch-Barrera, A. Oaknin, J. Alfon, C. Domenech, et al., The anti-cancer drug ABTL0812 induces ER stress-mediated cytotoxic autophagy by increasing dihydroceramide levels in cancer cells. Autophagy (2021). https://doi.org/10.1080/15548627.2020.1761651
S.Z. Xie, L. Garcia-Prat, V. Voisin, R. Ferrari, O.I. Gan, E. Wagenblast, K.B. Kaufmann, A. Zeng, S.I. Takayanagi, I. Patel, E.K. Lee, J. Jargstorf, G. Holmes, G. Romm, K. Pan, M. Shoong, A. Vedi, C. Luberto, M.D. Minden, et al., Sphingolipid modulation activates Proteostasis programs to govern human hematopoietic stem cell self-renewal. Cell Stem Cell (2019). https://doi.org/10.1016/j.stem.2019.09.008
C. Di Malta, D. Siciliano, A. Calcagni, J. Monfregola, S. Punzi, N. Pastore, A.N. Eastes, O. Davis, R. De Cegli, A. Zampelli, L.G. Di Giovannantonio, E. Nusco, N. Platt, A. Guida, M.H. Ogmundsdottir, L. Lanfrancone, R.M. Perera, R. Zoncu, P.G. Pelicci, et al., Transcriptional activation of RagD GTPase controls mTORC1 and promotes cancer growth. Science (2017). https://doi.org/10.1126/science.aag2553
L. Ding, X. Liang, Ras related GTP binding D promotes aerobic glycolysis of hepatocellular carcinoma. Ann. Hepatol. (2021). https://doi.org/10.1016/j.aohep.2021.100307
Acknowledgements
We acknowledge the owners of TCGA, GEO and all other databases for providing valuable platforms and for making all those meaningful data available. All the contributions to those public datasets are deeply appreciated.
Funding
This work was supported by grants from the Beijing Nova Program of Science and Technology (Z191100001119128), the National Natural Science Foundation of China (82073390, 81702314), the Beijing Municipal Science and Technology Project (Z191100006619081), the Beijing Municipal Administration of Hospitals’ Youth Programme (QML20180108), the Funding Program for Excellent Talents of Beijing (2017000021469G212) and the Digestive Medical Coordinated Development Center of Beijing Municipal Administration of Hospitals (XXZ02, XXZ01). The study sponsors had no role in the design of the study and the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
Study concept and design: FB and LM. Data acquisition and data cleaning: YZ and FB. Data analysis and interpretation: FB, YSZ, YC and LS. Drafting of the manuscript: FB, XY, YZ and YSZ. Supervision: LM and SZ. Proofreading and revision: LM, SZ, LS, and YC. Critical revision of the manuscript for important intellectual content: All authors.
Corresponding authors
Ethics declarations
Ethics approval and inform consent
This study was conducted based on the open-source data from the TCGA and GEO databases. The patients involved in those databases have already provided ethical approval. All users were permitted to download relevant data for research and to publish relevant articles. Therefore, there are no ethical issues and other conflicts of interest regarding this article.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors have no competing interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bu, F., Zhao, Y., Zhao, Y. et al. Distinct tumor microenvironment landscapes of rectal cancer for prognosis and prediction of immunotherapy response. Cell Oncol. 45, 1363–1381 (2022). https://doi.org/10.1007/s13402-022-00725-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-022-00725-1