Abstract
Background and Objectives
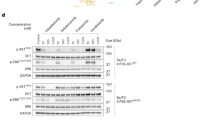
Rearranged during transfection (RET) is a transmembrane receptor tyrosine kinase that plays a crucial role in tumorigenesis. FHND5071, a potent and selective RET kinase inhibitor, could exert antitumor effects by inhibiting RET autophosphorylation. The present work aims to profile the pharmacokinetics of FHND5071 in in vivo and in vitro experiments as a ground work for further clinical research.
Methods
The absorption, distribution, metabolism, and excretion properties of FHND5071 were examined, along with metabolite production and cytochrome P450 (CYP) phenotyping assay. Additionally, plasma protein binding and pharmacokinetics in mice were investigated.
Results
Microsomal stability assay corroborated moderate to high clearance of FHND5071, and the use of UPLC-Q-TOF-MS identified a total of six metabolites and suggested a possible metabolic pathway involving oxidation, demethylation, and N-dealkylation. Primary contributors to the CYP-mediated metabolism of FHND5071 were found to be CYP2C8 and CYP3A4, and FHND5071 displayed low permeability and acted as a substrate for the P-glycoprotein (P-gp). FHND5071 had a moderate to high binding in plasma and exhibited a moderate absorption degree (absolute bioavailability > 60%) The distribution of FHND5071 in mouse tissues was rapid (mostly peaking at 1–4 h) and wide (detectable in almost all tissues and organs), with the highest exposure in the spleen. A small fraction of FHND5071 was excreted via the urine and feces, and a presumed metabolic pathway involving 20 metabolites in mice is proposed.
Conclusion
Pharmacokinetic characteristics of FHND5071 were systemically profiled, which may lay the foundation for further clinical development as a drug candidate.
Graphical Abstract











Similar content being viewed by others
References
Takahashi M, Ritz J, Cooper GM. Activation of a novel human transforming gene, ret, by DNA rearrangement. Cell. 1985;42(2):581–8. https://doi.org/10.1016/0092-8674(85)90115-1.
Trupp M, Scott R, Whittemore SR, Ibanez CF. Ret-dependent and -independent mechanisms of glial cell line-derived neurotrophic factor signaling in neuronal cells. J Biol Chem. 1999;274(30):20885–94. https://doi.org/10.1074/jbc.274.30.20885.
Andreozzi F, Melillo R, Carlomagno F, Oriente F, Miele C, Fiory F, et al. Protein kinase Calpha activation by RET: evidence for a negative feedback mechanism controlling RET tyrosine kinase. Oncogene. 2003;22(19):2942–9. https://doi.org/10.1038/sj.onc.1206475.
Bhattarai C, Poudel P, Ghosh A, Kalthur SJAN. RETThe gene encodes RET protein, which triggers intracellular signaling pathways for enteric neurogenesis, and mutation results in Hirschsprung’s disease. AIMS Neurosci. 2022;9(1):128–49. https://doi.org/10.3934/Neuroscience.2022008.
Fukuda T, Kiuchi K, Takahashi M. Novel mechanism of regulation of Rac activity and lamellipodia formation by RET tyrosine kinase. J Biol Chem. 2002;277(21):19114–21. https://doi.org/10.1074/jbc.M200643200.
Chi X, Michos O, Shakya R, Riccio P, Enomoto H, Licht JD, et al. Ret-dependent cell rearrangements in the Wolffian duct epithelium initiate ureteric bud morphogenesis. Develop Cell. 2009;17(2):199–209. https://doi.org/10.1016/j.devcel.2009.07.013.
de Graaff E, Srinivas S, Kilkenny C, D’Agati V, Mankoo BS, Costantini F, et al. Differential activities of the RET tyrosine kinase receptor isoforms during mammalian embryogenesis. Genes Develop. 2001;15(18):2433–44. https://doi.org/10.1101/gad.205001.
Kohno T, Tabata J, Nakaoku T. REToma: a cancer subtype with a shared driver oncogene. Carcinogenesis. 2020;41(2):123–9. https://doi.org/10.1093/carcin/bgz184.
Liu X, Hu X, Shen T, Li Q, Mooers BHM, Wu J. RET kinase alterations in targeted cancer therapy. Cancer Drug Resist. 2020;3(3):472-81.https://doi.org/10.20517/cdr.2020.15
Tsuzuki T, Takahashi M, Asai N, Iwashita T, Matsuyama M, Asai JP. Spatial and temporal expression of the ret proto-oncogene product in embryonic, infant and adult rat tissues. Oncogene. 1995;10(1):191–8. https://doi.org/10.1016/0014-5793(94)01388-H.
Kohno T, Ichikawa H, Totoki Y, Yasuda K, Hiramoto M, Nammo T, et al. KIF5B-RET fusions in lung adenocarcinoma. Nat Med. 2012;18(3):375–7. https://doi.org/10.1038/nm.2644.
Lipson D, Capelletti M, Yelensky R, Otto G, Parker A, Jarosz M, et al. Identification of new ALK and RET gene fusions from colorectal and lung cancer biopsies. Nat Med. 2012;18(3):382–4. https://doi.org/10.1038/nm.2673.
Mulligan LM. RET revisited: expanding the oncogenic portfolio. Nat Rev Cancer. 2014;14(3):173–86. https://doi.org/10.1038/nrc3680.
Romei C, Ciampi R, Elisei R. A comprehensive overview of the role of the RET proto-oncogene in thyroid carcinoma. Nat Rev Endocrinol. 2016;12(4):192–202. https://doi.org/10.1038/nrendo.2016.11.
Stransky N, Cerami E, Schalm S, Kim JL, Lengauer C. The landscape of kinase fusions in cancer. Nat Commun. 2014;5:4846. https://doi.org/10.1038/ncomms5846.
Drilon A, Hu ZI, Lai GGY, Tan DSW. Targeting RET-driven cancers: lessons from evolving preclinical and clinical landscapes. Nat Rev Clin Oncol. 2018;15(3):151–67. https://doi.org/10.1038/nrclinonc.2017.175.
Pall G, Gautschi O. Advances in the treatment of RET-fusion-positive lung cancer. Lung Cancer. 2021;156:136–9. https://doi.org/10.1016/j.lungcan.2021.04.017.
Subbiah V, Cote GJ. Advances in targeting RET-dependent cancers. Cancer Discov. 2020;10(4):498–505. https://doi.org/10.1158/2159-8290.CD-19-1116.
Subbiah V, Velcheti V, Tuch BB, Ebata K, Busaidy NL, Cabanillas ME, et al. Selective RET kinase inhibition for patients with RET-altered cancers. Ann Oncol. 2018;29(8):1869–76. https://doi.org/10.1093/annonc/mdy137.
Thein KZ, Velcheti V, Mooers BHM, Wu J, Subbiah V. Precision therapy for RET-altered cancers with RET inhibitors. Trends Cancer. 2021;7(12):1074–88. https://doi.org/10.1016/j.trecan.2021.07.003.
Ekpenyong O, Gao X, Ma J, Cooper C, Nguyen L, Olaleye OA, et al. Pre-clinical pharmacokinetics, tissue distribution and physicochemical studies of CLBQ14, a novel methionine aminopeptidase inhibitor for the treatment of infectious diseases. Drug Des Devel Ther. 2020;14:1263–77. https://doi.org/10.2147/DDDT.S238148.
Mosure KW, Knipe JO, Browning M, Arora V, Sinz MJJOPS. Preclinical Pharmacokinetics and In Vitro Metabolism of Asunaprevir (BMS-650032), a Potent Hepatitis C Virus NS3 Protease Inhibitor. J Pharm Sci. 2015;104(9)https://doi.org/10.1002/jps.24356
He P, Niu S, Wang S, Shi X, Feng S, Du L, et al. Discovery of WS-157 as a highly potent, selective and orally active EGFR inhibitor. Acta Pharm Sin B. 2019;9(6):1193–203. https://doi.org/10.1016/j.apsb.2019.06.010.
Zhang S, Zhao Y, Wang S, Li M, Xu Y, Ran J, et al. Discovery of novel diarylamides as orally active diuretics targeting urea transporters. Acta Pharm Sin B. 2021;11(1):181–202. https://doi.org/10.1016/j.apsb.2020.06.001.
Zhang X, Cheng X, Wu Y, Feng D, Qian Y, Chen L, et al. In Vitro and In Situ characterization of the intestinal absorption of Capilliposide B and Capilliposide C from Lysimachia capillipes Hemsl. Molecules. 2019. https://doi.org/10.3390/molecules24071227.
Yu L, Chen X, Zhang WS, Zheng L, Xu WW, Xu MY, et al. Metabolite identification, tissue distribution, excretion and preclinical pharmacokinetic studies of ET-26-HCl, a new analogue of etomidate. R Soc Open Sci. 2020;7(2):191666. https://doi.org/10.1098/rsos.191666.
Houston JB. Utility of in vitro drug metabolism data in predicting in vivo metabolic clearance. Biochem Pharmacol. 1994;47(9):1469–79. https://doi.org/10.1016/0006-2952(94)90520-7.
Chen J, Liu D, Zheng X, Zhao Q, Jiang J, Hu PJEoodm, et al. Relative contributions of the major human CYP450 to the metabolism of icotinib and its implication in prediction of drug-drug interaction between icotinib and CYP3A4 inhibitors/inducers using physiologically based pharmacokinetic modeling. Expert Opin Drug Metab Toxicol.2015;11(6):857-68.https://doi.org/10.1517/17425255.2015.1034688
Tang L, Ye L, Lv C, Zheng Z, Gong Y, Liu Z. Involvement of CYP3A4/5 and CYP2D6 in the metabolism of aconitine using human liver microsomes and recombinant CYP450 enzymes. Toxicol Lett. 2011;202(1):47–54. https://doi.org/10.1016/j.toxlet.2011.01.019.
Ohmori S, Horie T, Guengerich F, Kiuchi M, Kitada MJAob, biophysics. Purification and characterization of two forms of hepatic microsomal cytochrome P450 from untreated cynomolgus monkeys. Arch Biochem Biophys. 1993;305(2):405-13.https://doi.org/10.1006/abbi.1993.1439
Komori M, Kikuchi O, Sakuma T, Funaki J, Kitada M, Kamataki T. Molecular cloning of monkey liver cytochrome P-450 cDNAs: similarity of the primary sequences to human cytochromes P-450. Biochim Biophys Acta. 1992;1171(2):141–6. https://doi.org/10.1016/0167-4781(92)90113-e.
Peter F, Interactions GJC-B. Comparisons of catalytic selectivity of cytochrome P450 subfamily enzymes from different species. Chem Biol Interact.1997;106(3):161-82. https://doi.org/10.1016/s0009-2797(97)00068-9
Li M, de Graaf I, van de Steeg E, de Jager M, Groothuis GJ, Tivaijpiaw B. The consequence of regional gradients of P-gp and CYP3A4 for drug-drug interactions by P-gp inhibitors and the P-gp/CYP3A4 interplay in the human intestine ex vivo. Toxicol In Vitro. 2017;40:26–33. https://doi.org/10.1016/j.tiv.2016.12.002.
Ahmed EY, Abdelhafez OM, Zaafar D, Serry AM, Ahmed YH, El-Telbany RFA, et al. Antitumor and multikinase inhibition activities of some synthesized coumarin and benzofuran derivatives. Arch Pharm (Weinheim). 2022:e2100327.https://doi.org/10.1002/ardp.202100327
Li L, Chen X, Zhou J, Zhong DJDm, chemicals dtbfo. In vitro studies on the oxidative metabolism of 20(s)-ginsenoside Rh2 in human, monkey, dog, rat, and mouse liver microsomes, and human liver s9. Drug Metab Dispos. 2012;40(10):2041-53.https://doi.org/10.1124/dmd.112.046995
Jin Z, Qiu W, Liu H, Jiang X, Wang LJCjonm. Enhancement of oral bioavailability and immune response of Ginsenoside Rh2 by co-administration with piperine. Chin J Nat Med. 2018;16(2):143-9.https://doi.org/10.1016/s1875-5364(18)30041-4
Pang KJDm, chemicals dtbfo. Modeling of intestinal drug absorption: roles of transporters and metabolic enzymes (for the Gillette Review Series). Drug Metab Dispos. 2003;31(12):1507-19.https://doi.org/10.1124/dmd.31.12.1507
Acknowledgements
We thank the Jiangsu Chia Tai Fenghai Pharmaceutical Co. Ltd. for providing funds and instruments. We are grateful to all members of the Zhu laboratory for valuable discussion.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This work was supported by the National Natural Science Foundation of China (121877061 to YQ.Z.) and Jiangsu Chia Tai Fenghai Pharmaceutical Co. Ltd.
Conflict of Interest
Jia Wang and Jinmiao Shi are employees of Jiangsu Chia Tai Fenghai Pharmaceutical Co. Ltd. All other authors have no potential conflicts of interest, financial or otherwise, to declare.
Ethics Approval
All experiments involving animals were performed in accordance with protocols approved by Institutional Animal Care and Use Committee of Nanjing Normal University under approval number IACUC-20200506 on June 17, 2021 and the guidelines of the Animal Welfare Council of China.
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Availability of Data and Materials
The data used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Code Availability
Not applicable.
Author Contributions
Yiran Han (First Author): Conceptualization, Methodology, Software, Investigation, Formal Analysis, Writing - Original Draft; Tiantian Wen: Methodology, Formal Analysis. Jingmiao Shi: Methodology, Software, Formal Analysis. Jia Wang: Visualization, Writing - Review & Editing. Yongqiang Zhu (Corresponding Author). Conceptualization, Funding Acquisition, Resources, Supervision, Writing - Review & Editing.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Han, Y., Wen, T., Wang, J. et al. Preclinical Pharmacokinetics and in vitro Metabolism of FHND5071, a Novel Selective RET Kinase Inhibitor. Eur J Drug Metab Pharmacokinet 48, 595–614 (2023). https://doi.org/10.1007/s13318-023-00844-6
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13318-023-00844-6