Abstract
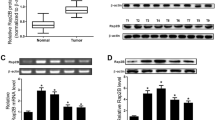
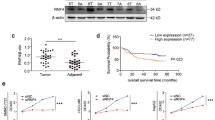
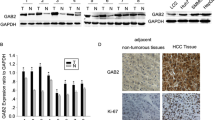
IQ motif-containing GTPase-activating proteins (IQGAPs) belong to a conserved family, and they are involved in various intracellular processes. IQGAP1 is expressed in all cells, while IQGAP2 and IQGAP3 are mainly expressed in hepatic cells. IQGAP1 has been suggested to be an oncogene, while IQGAP2 is considered a tumor-suppressor gene. However, the relationship between RAS family genes and IQGAP genes remains unclear. We recently demonstrated this interaction in a chemically induced mouse liver cancer. In this study, IQGAP1 expression was partially silenced in human hepatocellular carcinoma (HepG2) cells. We investigated the impact of IQGAP1 silencing on the interactions of IQGAP and RAS with several apoptotic proteins, including caspase-3 (CASP3), BCL2-associated X protein (BAX), and B-cell leukemia/lymphoma 2 (BCL2). Additionally, we investigated the effects of the interactions of these genes on cell viability, proliferation, apoptosis, and invasive capacity. IQGAP1 siRNA-treated HepG2 cells showed lower invasive capacity than the control cells, and this reduction was time- and vector concentration-dependent. In addition, IQGAP1 silencing resulted in significantly lower IQGAP1 level and subsequently higher IQGAP2 and IQGAP3 expression in HepG2 cells than in the control. Flow cytometry analyses indicated that the silencing of IQGAP1 can induce early and late apoptosis in HepG2 cells. Additionally, IQGAP2, IQGAP3, CASP3, and BAX were upregulated whereas IQGAP1 and BCL2 were downregulated in the siRNA-treated cells. Furthermore, we observed that the mRNA levels of HRAS, KRAS, NRAS, and MRAS decreased upon IQGAP1 silencing. These findings indicate that IQGAP1 potentially regulates the expression of IQGAP and RAS gene families and demonstrate its regulatory role in the apoptotic network. Taken together, our findings suggest that IQGAP1 silencing plays crucial roles in the apoptosis of HepG2 cells and lowers their proliferative and invasive capacities.






Similar content being viewed by others
References
Bernards A. GAPs galore! A survey of putative Ras superfamily GTPase activating proteins in man and drosophila. Biochim Biophys Acta. 2003;1603:47–82.
Chen F, Zhu H, Zhou L, Wu S, Wang J, Chen Z. IQGAP1 is overexpressed in hepatocellular carcinoma and promotes cell proliferation by Akt activation. EXPERIMENTAL and MOLECULAR MEDICINE. July 2010;42(7):477–83.
Fukata M, Nakagawa M, Kaibuchi K. Roles of rho-family GTPases in cell polarisation and directional migration. Curr Opin Cell Biol. 2003;15(5):590–7.
Machesky LM. Curr Biol. 1998;8:R202.
Noritake J, Watanabe T, Sato K, Wang S, Kaibuchi K. J Cell Sci. 2005;118:2085.
Brown MD, Sacks DB. IQGAP1 in cellular signaling: bridging the gap. Trends Cell Biol. 2006;16(5):242–9.
Schmidt VA, Chiariello CS, Capilla E, Miller F, Bahou WF. Development of hepatocellular carcinoma in Iqgap2 deficient mice is IQGAP1 dependent. Mol Cell Biol. 2008;28(5):1489–502.
Schmidt VA, Scudder L, Devoe CE, Bernards A, Cupit LD, Bahou WF. IQGAP2 functions as a GTP-dependent effector protein in thrombin-induced platelet cytoskeletal reorganization. Blood. 2003;101(8):3021–8.
Wang S, Watanabe T, Noritake J, Fukata M, Yoshimura T, Itoh N, Harada T, Nakagawa M, Matsuura Y, Arimura N, et al. IQGAP3, a novel effector of Rac1 and Cdc42, regulates neurite outgrowth. J Cell Sci. 2007;120(4):567–77.
Briggs MW, Sacks DB. IQGAP proteins are integral components of cytoskeletal regulation EMBO rep. 2003;4:571–4.
White CD, Brown MD, Sacks DB IQGAPs in cancer: a family of scaffold proteins underlying tumorigenesis. FEBS Lett 2009;583:1817–1824
Zoheir KMA, Abd-Rabou AA, Harisa GI, Ashour AE, Ahmad SF, Attia SM, et al. Gene expression of IQGAPs and Ras families in an experimental mouse model for hepatocellular carcinoma: a mechanistic study of cancer progression. International journal of clinical and experimental pathology. 2015;8(8):8821.
Johnson M, Sharma M, Henderson BR. IQGAP1 regulation and roles in cancer cellular signalling. 2009;21:1471–8.
Jadeski L, Mataraza JM, Jeong HW, Li Z, Sacks DB. IQGAP1 stimulates proliferation and enhances tumorigenesis of human breast epithelial cells. J Biol Chem 2008;283: 1008–1017
Galuppo R, Maynard E, Shah M, Daily MF, Chen C, Spear BT, Gedaly R. Synergistic inhibition of HCC and liver cancer stem cell proliferation by targeting RAS/RAF/MAPK and WNT/β-catenin pathways. Anticancer Res. 2014;34:1709–14.
Kim Y, Sills RC, Houle CD. Overview of the molecular biology of hepatocellular neoplasms and hepatoblastomas of the mouse liver. Toxicol Pathol. 2005;33:175–80.
Thorgeirsson SS, Grisham JW. Molecular pathogenesis of human hepatocellular carcinoma. Nat Genet. 2002;31:339–46.
Fire A et al. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature. 1998;391:806–11.
Elbashir SM et al. Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells. Nature. 2001;411:494–8.
McCaffrey AP et al. Gene expression: RNA interference in adult mice. Nature. 2002;418:38–9.
Morrissey DV et al. Potent and persistent in vivo anti-HBV activity of chemically modified siRNAs. Nature Biotech. 2005;23:1002–7.
Okumura A, Pitha PM, Harty RN. ISG15 inhibits Ebola VP40 VLP budding in an L-domain-dependent manner by blocking Nedd4 ligase activity. Proc Natl Acad Sci U S A. 2008;105:3974–9.
Ptasznik A, Nakata Y, Kalota A, Emerson SG, Gewirtz AM. Short interfering RNA (siRNA) targeting the Lyn kinase induces apoptosis in primary, and drug-resistant, BCR-ABL1+ leukemia cells. Nature Med. 2004;10:1187–9.
Kim SH, Jeong JH, Lee SH, Kim SW, Park TG. Local and systemic delivery of VEGF siRNA using polyelectrolyte complex micelles for effective treatment of cancer. J Control Release. 2008;129:107–16.
Xia C-F, Zhang Y, Zhang Y, Boado R, Pardridge W. Intravenous siRNA of brain cancer with receptor targeting and avidin–biotin technology. Pharm Res. 2007;24:2309–16.
Dong P.X., Jia N., Xu Z.J, Liu Y.T., Li D.J, Feng Y.J., J. Exp. Clin. Cancer Res. 27 (2008)
Clark EA, Golub TR, Lander ES, Hynes RO. Nature. 2000;406:532.
Yamaoka-Tojo M et al. IQGAP1 a novel vascular endothelial growth factor receptor binding protein, is involved in reactive oxygen species-dependent endothelial migration and proliferation. Circ Res. 2004;95:276–83.
Meyer RD, Sacks DB, Rahimi N. IQGAP1-dependent signaling pathway regulates endothelial cell proliferation and angiogenesis. PLoS One. 2008;3:e3848–58.
Liang C, Park AY, Guan J. In vitro scratch assay: a convenient and inexpensive method for analysis of cell migration in vitro. Nat Protoc. 2007;2:329–33.
Dong P, Nabeshima K, Nishimura N, Kawakami T, Hachisuga T, Kawarabayashi T, Iwasaki H. Overexpression and diffuse expression pattern of IQGAP1 at invasion fronts are independent prognostic parameters in ovarian carcinomas. Cancer Lett. 2006;243:120–7.
Nabeshima K, Shimao Y, Inoue T, Koono M. Immunohistochemical analysis of IQGAP1 expression in human colorectal carcinomas: its overexpression in carcinomas and association with invasion fronts. Cancer Lett. 2002;176:101–9.
Walch A, Seidl S, Hermannstadter C, et al. Combined analysis of Rac1, IQGAP1, Tiam1 and E-cadherin expression in gastric cancer. Mod Pathol. 2008;21:544–52.
Jin SH, Akiyama Y, Fukamachi H, et al. IQGAP2 inactivation through aberrant promoter methylation and promotion of invasion in gastric cancer cells. Int J Cancer. 2008;122:1040–6.
Xie Y, Yan J, Cutz JC, et al. IQGAP2, a candidate tumour suppressor of prostate tumorigenesis. Biochim Biophys Acta. 2012;1822:875–84.
Schmidt VA. Watch the GAP: emerging roles for IQ motif-containing GTPase-activating proteins IQGAPs in hepatocellular carcinoma. Int J Hepatol. 2012;2012:958673.
Zhou J, et al. Quantitative proteomic analysis of HepG2 cells treated with quercetin suggests IQGAP1 is involved in quercetin-induced regulation of cell proliferation and migration. Omics. 2009.
Roy M, Li Z, Sacks DB. IQGAP1 is a scaffold for mitogen-activated protein kinase signaling. Mol Cell Biol. 2005;25:7940–52.
Gnatenko DV, Xu X, Zhu W, Schmidt VA. Transcript profiling identifies iqgap2(−/−) mouse as a model for advanced human hepatocellular carcinoma. PLoS One. 2013;8:e71826.
Brill S, Li S, Lyman CW, et al. The Ras GTPase-activating-protein-related human protein IQGAP2 harbors a potential actin binding domain and interacts with calmodulin and Rho family GTPases. Mol Cell Biol. 1996;16:4869–78.
Hart MJ, Callow MG, Souza B, Polakis P. IQGAP1, a calmodulin-binding protein with a RasGAP-related domain, is a potential effector for cdc42Hs. EMBO J. 1996;15:2997–3005.
McCallum SJ, WJ W, Cerione RA. Identification of a putative effector for Cdc42Hs with high sequence similarity to the RasGAP-related protein IQGAP1 and a Cdc42Hs binding partner with similarity to IQGAP2. J Biol Chem. 1996;271:21732–7.
Joyal JL, Annan RS, Ho YD, et al. Calmodulin modulates the interaction between IQGAP1 and Cdc42. Identification of IQGAP1 by nanoelectrospray tandem mass spectrometry. J Biol Chem. 1997;272:15419–25.
Van Hengel J, D’Hooge P, Hooghe B, et al. Continuous cell injury promotes hepatic tumorigenesis in cdc42-deficient mouse liver. Gastroenterology. 2008;134:781–92.
Acknowledgments
The project was funded by the National Plan for Science, Technology and Innovation (MAARIFAH) King Abdulaziz City for Science and Technology, Kingdom of Saudi Arabia, award number 12-BIO2926-02.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
None
Rights and permissions
About this article
Cite this article
Zoheir, K.M., Abd-Rabou, A.A., Harisa, G.I. et al. IQGAP1 gene silencing induces apoptosis and decreases the invasive capacity of human hepatocellular carcinoma cells. Tumor Biol. 37, 13927–13939 (2016). https://doi.org/10.1007/s13277-016-5283-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-016-5283-8