Abstract
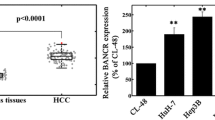
Long noncoding RNAs (lncRNAs) have been found dysregulated in human disease, especially in cancer. Small nucleolar RNA host gene 3 (SNHG3) is an lncRNA whose potential function and mechanism in hepatocellular carcinoma (HCC) remain largely unknown. In the present study, we aimed to determine SNHG3 expression and its clinical significance in HCC. Our results showed that the expression level of SNHG3 was significantly upregulated in HCC tissues compared with paired noncancerous tissues from 51 HCC patients, as determined by quantitative real-time polymerase chain reaction (qRT-PCR; P < 0.001), which was consistent with the results of two independent HCC cohorts from The Cancer Genome Atlas (TCGA) and Oncomine databases (P < 0.0001 and P = 0.0325, respectively). These results were further confirmed in 144 paired paraffin-embedded HCC specimens by in situ hybridization assay (ISH). Furthermore, SNHG3 expression was significantly correlated with tumor size (P = 0.003), portal vein tumor thrombus (PVTT; P = 0.014), and relapse (P = 0.038). The high expression level of SNHG3 was markedly correlated with overall survival (OS; P < 0.0001), recurrence-free survival (RFS; P = 0.006), and disease-free survival (DFS; P < 0.0001). More importantly, multivariate analysis indicated that SNHG3 expression was an independent prognostic factor for HCC patients (P < 0.001). In conclusion, increased SNHG3 expression is associated with malignant status and poor prognosis in HCC patients.



Similar content being viewed by others
References
Meng X, Franklin DA, Dong J, Zhang Y. MDM2-p53 pathway in hepatocellular carcinoma. Cancer Res. 2014;74:716–7167.
Maluccio M, Covey A. Recent progress in understanding, diagnosing, and treating hepatocellular carcinoma. CA Cancer J Clin. 2012;62:394–9.
He G, Dhar D, Nakagawa H, Font-Burgada J, Ogata H, Jiang Y, et al. Identification of liver cancer progenitors whose malignant progression depends on autocrine IL-6 signaling. Cell. 2013;155:384–96.
Villanueva A, Minguez B, Forner A, Reig M, Llovet JM. Hepatocellular carcinoma: novel molecular approaches for diagnosis, prognosis, and therapy. Annu Rev Med. 2010;61:317–28.
Batista PJ, Chang HY. Long noncoding RNAs: cellular address codes in development and disease. Cell. 2013;152:1298–307.
Karreth FA, Pandolfi PP. CeRNA cross-talk in cancer: when ce-bling rivalries Go awry. Cancer Discov. 2013;3:1113–21.
Ulitsky I, Bartel DP. LincRNAs: genomics, evolution, and mechanisms. Cell. 2013;154:26–46.
Shi X, Sun M, Liu H, Yao Y, Song Y. Long non-coding RNAs: a new frontier in the study of human diseases. Cancer Lett. 2013;339:159–66.
Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP. A ceRNA hypothesis: the Rosetta stone of a hidden RNA language? Cell. 2011;146:353–8.
Shabalina SA1, Spiridonov NA. The mammalian transcriptome and the function of non-coding DNA sequences. Genome Biol [NLM - MEDLINE]. 2004;5:105.
Guttman M, Amit I, Garber M, French C, Lin MF, Feldser D, et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature. 2009;458:223–7.
Quinodoz S, Guttman M. Long noncoding RNAs: an emerging link between gene regulation and nuclear organization. Trends Cell Biol. 2014;24:651–63.
Brown CJ et al. A gene from the region of the human X inactivation centre is expressed exclusively from the inactive X chromosome. Nature. 1991;349:38–44.
Rinn JL1, Kertesz M, Wang JK, Squazzo SL, Xu X, Brugmann SA, et al. Functional demarcation of active and silent chromatin domains in human HOX loci by Non-coding RNAs. Cell. 2007;129:1311–23.
Gupta RA, Shah N, Wang KC, Kim J, Horlings HM, Wong DJ, et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature. 2010;464:1071–6.
Tripathi V, Ellis JD, Shen Z, Song DY, Pan Q, Watt AT, et al. The Nuclear-Retained noncoding RNA MALAT1 regulates alternative splicing by modulating SR splicing factor phosphorylation. Mol Cell. 2010;39:925–38.
Xia T, Liao Q, Jiang X, Shao Y, Xiao B, Xi Y, et al. Long noncoding RNA associated-competing endogenous RNAs in gastric cancer. Sci Rep-UK. 2014; 4.
Su S, Liu Q, Chen J, Chen J, Chen F, He C, et al. A positive feedback loop between mesenchymal-like cancer cells and macrophages is essential to breast cancer metastasis. Cancer Cell. 2014;25:605–20.
Trimarchi T, Bilal E, Ntziachristos P, Fabbri G, Dalla-Favera R, Tsirigos A, et al. Genome-wide mapping and characterization of notch-regulated long noncoding RNAs in acute leukemia. Cell. 2014;158:593–606.
Yuan JH, Yang F, Wang F, Ma JZ, Guo YJ, Tao QF, et al. A long noncoding RNA activated by TGF-beta promotes the invasion-metastasis cascade in hepatocellular carcinoma. Cancer Cell. 2014;25:666–81.
Cao C, Sun J, Zhang D, Guo X, Xie L, Li X, et al. The long intergenic noncoding RNA UFC1, a target of MicroRNA 34a, interacts with the mRNA stabilizing protein HuR to increase levels of beta-Catenin in HCC cells. Gastroenterology. 2015;148:415–26.
Huang JL, Zheng L, Hu YW, et al. Characteristics of long non-coding RNA and its relation to hepatocellular carcinoma. Carcinogenesis. 2014;35:507–14.
Sun J, Bie B, Zhang S, et al. Long non-coding RNAs: critical players in hepatocellular carcinoma. Int J Mol Sci. 2014;15:20434–48.
Xing Z, Lin A, Li C, Liang K, Wang S, Liu Y, et al. LncRNA directs cooperative epigenetic regulation downstream of chemokine signals. Cell. 2014;159:1110–25.
Sun J, Bie B, Zhang S, Yang J, Li Z. Long non-coding RNAs: critical players in hepatocellular carcinoma. Int J Mol Sci. 2014;15:20434–48.
El Serag HB, Rudolph KL. Hepatocellular carcinoma: epidemiology and molecular carcinogenesis. Gastroenterology. 2007;132:2557–76.
He Y, Meng X, Huang C, Wu B, Zhang L, Lv X, et al. Long noncoding RNAs: novel insights into hepatocelluar carcinoma. Cancer Lett. 2014;344:20–7.
Zhu J, Liu S, Ye F, et al. The long noncoding RNA expression profile of hepatocellular carcinoma identified by microarray analysis. PLoS One. 2014;9:e101707.
Panzitt K, Tschernatsch MM, Guelly C, Moustafa T, Stradner M, Strohmaier HM, et al. Characterization of HULC, a novel gene with striking up-regulation in hepatocellular carcinoma, as noncoding RNA. Gastroenterology. 2007;132(1):330–42.
Geng YJ, Xie SL, Li Q, Ma J, Wang GY. Large intervening non-coding RNA HOTAIR is associated with hepatocellular carcinoma progression. J Int Med Res. 2011;39(6):2119–28.
Yang Z, Zhou L, Wu LM, Lai MC, Xie HY, Zhang F, et al. Overexpression of long non-coding RNA HOTAIR predicts tumor recurrence in hepatocellular carcinoma patients following liver transplantation. Ann Surg Oncol. 2011;18(5):1243–50.
Pelczar P et al. The host gene for intronic U17 small nucleolar RNAs in mammals has no protein-coding potential and is a member of the 5′-terminal oligopyrimidine gene family. Mol Cell Biol. 1998;18(8):4509–18.
Compliance with ethical standards
Funding
This study was funded by National Nature Science Foundation of China (81401180 and 81372283).
Conflicts of interest
None
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. This article does not contain any studies with animals performed by any of the authors.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Zhang, T., Cao, C., Wu, D. et al. SNHG3 correlates with malignant status and poor prognosis in hepatocellular carcinoma. Tumor Biol. 37, 2379–2385 (2016). https://doi.org/10.1007/s13277-015-4052-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-015-4052-4