Abstract
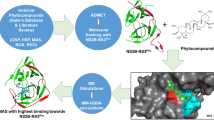
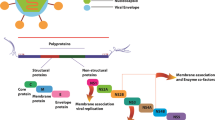
The current study aimed to assess the binding potential of herbal lead molecules against the prioritized molecular targets of chikungunya virus (CHIKV) and dengue virus (DENV) by computational virtual screening and suggests a novel therapeutic intervention. Based on the metabolic pathway analysis and virulent functions, the non-structural and envelop proteins present in CHIKV and DENV were identified as putative drug targets. The structures of the protein not available in their native forms were computationally predicted by homology modeling. The lead compounds from 43 herbal sources were screened and their drug likeliness and pharmacokinetics properties were computationally predicted. The binding potential of selected phytoligands against the prioritized drug targets were analyzed by molecular docking studies. This study revealed that Kaempferol (3,5,7-trihydroxy-2-(4-hydroxyphenyl)chromen-4-one) and Chymopain (disodium;4,5-dihydroxybenzene-1,3-disulfonate), natural flavonols present in Carica papaya and Gossypetin (3, 5, 7, 8, 3′, 4′-hexahydroxyflavone), a natural flavonoid available in Hibiscus sabdariffa were demonstrated promising good binding potential with minimum binding energy (kcal/mol) and maximum stabilizing interactions to the putative drug targets of CHIKV and DENV. The selected lead molecules demonstrated ideal drug likeliness, ADMET (adsorption, distribution, excretion, metabolism and toxicity) features required for the drug development. The molecular docking studies suggested that the presence of these compounds probably responsible for the antiviral properties of Carica papaya, which was traditionally known as therapeutic remedy for dengue viral infections. This study provides profound insight for the experimental validation of the applied approach and industrial scale-up of the suggested herbal lead molecules as promising lead candidates against CHIKV and DENV infections.

(Source: KEGG pathway database)




Similar content being viewed by others
References
Abu Bakar F, Ng LFP (2018) Nonstructural proteins of alphavirus-potential targets for drug development. Viruses. https://doi.org/10.3390/v10020071
Ajay A, Walters P, Murcko A (1998) Can we learn to distinguish between “drug-like” and “nondrug-like” molecules? J Med Chem 41:3314–3324
Ajay A, Bemis W, Murcko A (1999) Blood brain barrier: designing libraries with CNS activity. J Med Chem 42:4942–4951
Al-Tawfiq JA, Memish ZA (2018) Dengue Hemorrhagic Fever Virus in Saudi Arabia: A Review. Vector Borne Zoonotic Dis 18:75–81
Ames BN, Gold SL (2000) Paracelsus to parascience: The environmental cancer distraction. Mutat Res 447:3–33
Ashour J, Laurent-Rolle M, Shi PY, García-Sastre A (2009) NS5 of dengue virus mediates STAT2 binding and degradation. J Virol 83:5408–5418
Bowie A (2010) TRAF3: Uncovering the Real but Restricted Role in Human. Immunity 33:293–295
Bruno C (2011) Antiviral research and development against dengue Virus, WHO Rep, pp 1–101
Byler K, Collins J, Ogungbe I, Setzer W (2016) Alphavirus protease inhibitors from natural sources: a homology modeling and molecular docking investigation. Comput Biol Chem 64:163–184
Carrillo-Hernández MY, Ruiz-Saenz J, Villamizar LJ, Gómez-Rangel SY, Martínez-Gutierrez M (2018) Co-circulation and simultaneous co-infection of dengue, chikungunya, and zika viruses in patients with febrile syndrome at the Colombian-Venezuelan border. BMC Infect Dis. 18(1): 61. https://doi.org/10.1186/s12879-018-2976-1
Colovos C, Yeates T (1993) Verification of protein structures: Patterns of nonbonded atomic interactions. Protein Sci 2:1511–1519
Dayaraj C (2014) Current status of dengue and chikungunya in India. Seajph Jour 3:22–27
Dhama K, Karthik K, Khandia R, Munjal A, Tiwari R, Rana R, Khurana SK, Khan RU, Alagawany M, Farag MR, Dadar M, Joshi SK (2018) Medicinal and therapeutic potential of herbs and plant metabolites / extracts countering viral pathogens—current knowledge and future prospects. Curr Drug Metab. https://doi.org/10.2174/1389200219666180129145252
Fiser A, Sali A (2003) Modeller: Generation and refinement of homology-based protein structure models. Methods Enzymol 374:461–491
Frimurer T, Bywater R, Nærum L, Lauritsen L, Brunak S (2000) Improving the odds in discriminating “drug-like” from “non drug-like” compounds. J Chem Inform Comput Sci 40:1315–1324
Gómez-Calderón C, Mesa-Castro C, Robledo S, Gómez S, Bolivar-Avila S, Diaz-Castillo F, Martínez-Gutierrez M (2017) Antiviral effect of compounds derived from the seeds of Mammea americana and Tabernaemontana cymosa on dengue and chikungunya virus infections. BMC Complement Altern Med 17:57. https://doi.org/10.1186/s12906-017-1562-1
Irvine D, Takahashi L, Lockhart K, Cheong J, Tolan W, Selick E, Grove R (1999) MDCK (Madin-Darby canine kidney) cells: a tool for membrane permiability screening. J Pharm Sci 88:28–33
Kawai T, Akira S (2008) Toll-like receptor and RIG-1-like receptor signaling. Ann N Y Acad Sci 1143:1–20
Khan AH, Morita K, Parquet Md Mdel C, Hasebe F, Mathenge EG, Igarashi A (2002).Complete nucleotide sequence of chikungunya virus and evidence for an internal polyadenylation site. J Gen Virol 83(Pt 12):3075–3084
Laskowski A, MacArthur W, Moss D, Thornton M (1993) PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Cryst 26:283–291
Laurie AT, Jackson RM (2005) Q-SiteFinder: an energy-based method for the prediction of protein-ligand binding sites. Bioinformatics 21:1908–1916
Li Y, Siripanyaphinyo U, Tumkosit U, Noranate N, Atchareeya A, Pan Y, Kameoka M, Kurosu T, Ikuta K, Takeda N, Ananrapreecha S (2012) Poly (I:C), an agonist of toll-like receptor-3, inhibits replication of the Chikungunya virus in BEAS-2B cells. Virol J 9:114
Lipinski C, Lombardo F, Dominy B, Feeney P (2001) Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv 46:3–26
Luis K, Shaohong L, Gabriel M, Ricardo J, Soares M, Archie C, Wenbiao H, Patricia B, Francesca F, Rebecca D, Laith Y (2016) Co-distribution and co-infection of chikungunya and dengue viruses, Furuya-Kanamori. BMC Infect Dis 16–84
Ma L, Jones CT, Groesch TD, Kuhn RJ, Post CB (2004) Solution structure of dengue virus capsid protein reveals another fold. Proc Natl Acad Sci USA 101:3414–3419
Malet H, Coutard B, Jamal S, Dutartre H, Papageorgiou N, Neuvonen M, Ahola T, Forrester N, Gould EA, Lafitte D, Ferron F, Lescar J, Gorbalenya AE, de Lamballerie X, Canard B (2009) The crystal structures of Chikungunya and Venezuelan equine encephalitis virus nsP3 macro domains define a conserved adenosine binding pocket. J Virol 83:6534–6545
Meyer E (1991) The first years of the protein data bank. Protein Sci 6:1591–1597
Moore C, Bergstralh D, Duncan A, Lei Y, Morrison E, Zimmermann G, Accavitti-Loper A, Madden J, Sun L, Ye Z, Lich D, Heise T, Chen Z, Ting P (2008) Nlrx1 is a regulator of mitochondrial antiviral immunity. Nature 451:573–577
Mortelmans K, Zeiger E (2000) The Ames Salmonella/microsome mutagenicity assay. Mutat Res 2:29–60
O’Boyle NM, Banck M, James CA, Morley C, Vandermeersch T, Hutchison GR (2011) Open babel: an open chemical toolbox. J Cheminform 3:33. https://doi.org/10.1186/1758-2946-3-33
Paymen N, Pierre G, Ahmet C, John H (2009) RIG-I-like receptors: Sensing and responding to RNA virus infection. Semin Immunol 21: 215–222
Perera-Lecoin M, Luplertlop N, Surasombatpattana P, Liegeois F, Hamel R, Thongrungkiat S, Vargas M, Yssel H, Misse D (2016) Dengue and chikungunya coinfection—the emergence of an underestimated threat. In: Rodriguez-Morale s AJ (ed) Current topics in chikungunya. Chap. 5, InTech, Rijeka, Croatia
Powers A (2015) Chikungunya virus outbreak expansion and micro evolutionary events affecting epidemiology and epidemic potential. Res Rep Trop Med 6:11–19
Powers CN, Setzer WN (2016). In in-silico investigation of phytochemicals as antiviral agents against dengue fever. Comb Chem High Throughput Screen 19: 516–36
Puerta-Guardo H, Glasner DR, Harris E (2016) Dengue Virus NS1 disrupts the endothelial glycocalyx, leading to hyperpermeability. PLoS Pathog 12:e1005738. https://doi.org/10.1371/journal.ppat.1005738
Ramakrishnan C, Kutumbarao N, Suhitha S, Velmurugan D (2017) Structure-function relationship of Chikungunya nsP2 protease: a comparative study with papain. Chem Biol Drug Des 89:772–782
Rashad A, Mahalingam S, Keller A (2014) Chikungunya virus: emerging targets and new opportunities for medicinal chemistry. J Med Chem 57:1147–1166
Ruwali P, Rai N, Kumar N, Gautam P (2013) Antiviral potential of medicinal plants: an overview. Int Res J Pharm 4:8–16
Santhanam V, Waheeta H (2017) Towards the identification of novel phytochemical leads as macrodomain inhibitors of chikungunya virus using molecular docking approach. J Appl Pharm Sci 7:074–082
Seeliger D, de Groot B (2010) Ligand docking and binding site analysis with PyMOL and Autodock/Vina. J Comput Aided Mol Des 24:417–422
Seyedi S, Shukri M, Hassandarvish P, Oo A, Shankar E, Abubakar S, Zandi K (2016) Computational approach towards exploring potential anti-chikungunya activity of selected flavonoids. Sci Rep 6:1–7
Soe HJ, Yong YK, Al-Obaidi MMJ, Raju CS, Gudimella R, Manikam R, Sekaran SD (2018) Identifying protein biomarkers in predicting disease severity of dengue virus infection using immune-related protein microarray. Medicine (Baltimore) 97(5):e9713. https://doi.org/10.1097/MD.0000000000009713
Tan WL, Lee YK, Ho YF, Yusof R, Abdul Rahman N, Karsani SA (2018) Comparative proteomics reveals that YK51, a 4-Hydroxypandurantin-A analogue, down regulates the expression of proteins associated with dengue virus infection. PeerJ 5:e3939. https://doi.org/10.7717/peerj.3939
Thompson J, Gibson T, Hiqquins D (2002) Multiple sequence alignment using ClustalW and ClustalX. Curr Protoc Bioinform Chap 2:Unit 2.3. https://doi.org/10.1002/0471250953.bi0203s00
Torsten S, Jurgen K, Nicolas G, Manuel P (2003) SWISS-MODEL: an automated protein homology-modeling server. Nucleic Acids Res 31:3381–3385
Trott O, Olson AJ (2010) AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization and multithreading. J Comput Chem 31:455–461
Vázquez-Calvo Á, Jiménez de Oya N, Martín-Acebes MA, Garcia-Moruno E, Saiz JC (2017) Antiviral properties of the natural polyphenols delphinidin and epigallocatechin gallate against the flaviviruses West Nile virus, Zika virus, and Dengue virus. Front Microbiol 8:1314. https://doi.org/10.3389/fmicb.2017.01314
Veber F, Johnson R, Chen HY, Smith BR, Ward KW, Kopple KD (2002) Molecular properties that influence the oral bioavailability of drug candidates. J Med Chem 45:2615–2623
Wiederstein J, Sippl M (2007) ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res 35:W407–W410
Xie X, Zou J, Puttikhunt C, Yuan Z, Shi PY (2015) Two distinct sets of NS2A molecules are responsible for dengue virus RNA synthesis and virion assembly. J Virol 89:1298–1313
Xu D, Zhang Y (2011) Improving the physical realism and structural accuracy of protein models by a two-step atomic-level energy minimization. Biophys J 101:2525–2534
Yazdanian M, Glynn L, Wright L, Hawi (1998) Correlating partitioning and Caco-2 cell permeability of structurally diverse small molecular weight compounds”. Pharm Res 15:1490–1494
Yoneyama M, Fujita T (2009) RNA recognition and signal transduction by RIG-I-like receptors. Immunol Rev 227:54–65
Zhong B, Yang Y, Li S, Wang Y, Li Y, Diao F, Lei C, He X, Zhang L, Tien P, Shu H (2008) The adaptor protein MITA links virus-sensing receptors to IRF3 transcription factor activation. Immunity 29:538–550
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no potential conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Keramagi, A.R., Skariyachan, S. Prediction of binding potential of natural leads against the prioritized drug targets of chikungunya and dengue viruses by computational screening. 3 Biotech 8, 274 (2018). https://doi.org/10.1007/s13205-018-1303-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-018-1303-2