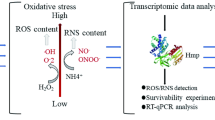
Abstract
To date, the mechanisms underlying the flavoprotein l-amino acid oxidase (LAAO) accumulation in cells remain unclear. In this study, using LAAO-producer Pseudoalteromonas spp. as model organisms, we found that the cell biomass is negatively associated with LAAO accumulation, whereas the LAAO accumulation is positively associated with the reactive oxygen species (ROS)-scavenging capability. The expression levels of ROS-scavenging-associated genes were up-regulated with LAAO accumulation in Pseudoalteromonas cells, which is presumably due to the requirement for the removal of LAAO-induced ROS. Exogenous H2O2 exposure experiment supported that the ROS-scavenging system is associated with LAAO accumulation in Pseudoalteromonas. All these observations indicate that ROS-scavenging capacity determines LAAO accumulation in bacterial cells. Our results shed a light on understanding the mechanism underlying controlling and adapting to LAAO accumulation in Pseudoalteromonas. Besides, our findings are critical to the improvement of heterologous expression of active LAAO in the future.





Similar content being viewed by others
References
Boehm DE, Vincent K, Brown OR (1976) Oxygen and toxicity inhibition of amino acid biosynthesis. Nature 262(5567):418–420
Chacón-Verdú MD, Gómez D, Solano F, Lucas-Elío P, Sánchez-Amat A (2013) LodB is required for the recombinant synthesis of the quinoprotein l-lysine-ε-oxidase from Marinomonas mediterranea. Appl Microbiol Biotechnol 98(7):2981–2989
Chen WM, Lin CY, Chen CA (2010) Involvement of an l-amino acid oxidase in the activity of the marine bacterium Pseudoalteromonas flavipulchra against methicillin-resistant staphylococcus aureus. Enzyme Microb Technol 47:52–58
Christman MF, Morgan RW, Jacobson FS, Ames N (1985) Positive control of a regulon for defenses against oxidative stress and some heat-shock proteins in Salmonella typhimurium. Cell 41(3):753–762
Gómez D, Espinosa E, Bertazzo M, Lucas-Elío P, Solano F, Sanchez-Amat A (2008) The macromolecule with antimicrobial activity synthesized by Pseudoalteromonas luteoviolacea strains is an l-amino acid oxidase. Appl Microbiol Biotechnol 79:925–930
Hassett DJ, Ma JF, Elkins JG, McDermott TR, Ochsner UA, West SE, Huang CT, Fredericks J, Burnett S, Stewart PS, McPheters G, Passador L, Iglewski BH (1999) Quorum sensing in Pseudomonas aeruginosa controls expression of catalase and superoxide dismutase genes and mediates biofilm susceptibility to hydrogen peroxide. Mol Microbiol 34(5):1082–1093
Imlay JA (2013) The molecular mechanisms and physiological consequences of oxidative stress: lessons from a model bacterium. Nat Rev Microbiol 11(7):443–454
Isnansety A, Kamei Y (2003) Pseudoalteromonas phenolica sp. nov. a novel marine bacterium that produces phenolic anti-methicillin-resistant Staphylococcus aureus substances. Int J Syst Evol Microbiol 53:583–588
James SG, Holmstrom C, Kjelleberg S (1996) Purification and characterization of a novel antibacterial protein from the marine bacterium D2. Appl Environ Microbiol 1(62):2783–2788
Kitani Y, Mori T, Nagai H, Toyooka K, Ishizaki S, Shimakura K, Shiomi K, Nagashima Y (2007) Gene expression and distribution of antibacterial l-amino acid oxidase in the rockfish Sebastes schlegeli. Fish Shellfish Immunol 23:1178–1186
Li JJ, Tang Q, Li Y, Hu BR, Ming ZY, Fu Q, Qian JQ, Xiang JZ (2006) Role of oxidative stress in the apoptosis of hepatocellular carcinoma induced by combination of arsenic trioxide and ascorbic acid. Acta Pharmacol Sin 27(8):1078–1084
Lourenco P, Batista P, Oliveira P, Silvestre C, Chen P (2008) Hydrogen peroxide linked to lysine oxidase activity facilitates biofilm differentiation and dispersal in several Gram-negative bacteria. J Bacteriol 190(15):5493–5501
Lucas EP, Gomez D, Solano F, Sanchez-Amat A (2006) The antimicrobial activity of marinocine, synthesized by Marinomonas mediterranea, is due to hydrogen peroxide generated by its lysine oxidase activity. J Bacteriol 188(7):2493–2501
Mishra S, Imlay JA (2012) Why do bacteria use so many enzymes to scavenge hydrogen peroxide? Arch Biochem Biophys 525(2):145–160
Nakano M, Danowski TS (1966) Crystalline mammalian l-amino acid oxidase from rat kidney mitochondria. J Biol Chem 241(9):2075–2083
Ochsner UA, Vasil ML, Alsabbagh E, Parvatiyar K, Hassett DJ (2000) Role of the Pseudomonas aeruginosa oxyR-recG operon in oxidative stress defense and DNA repair: oxyR-dependent regulation of katB, ahpB and ahpCF. J Bacteriol 182:4533–4544
Ohkawa H, Ohishi N, Yagi K (1979) Assay for lipid peroxides in animal tissues by thiobarbituric acid reaction. Anal Biochem 95(2):351–358
Pfaffl MW (2001) A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res 29(9):e45
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative C T method. Nat Protoc 3(3):1101–1108
Storz G, Imlay JA (1999) Oxidative stress. Curr Opin Microbiol 2(2):188–194
Suh SJ, Silosuh L, Woods DE, Hassett DJ, West SE, Ohman DE (1999) Effect of rpos mutation on the stress response and expression of virulence factors in Pseudomonas aeruginosa. J Bacteriol 181:3890–3897
Suhr M, Kim DS (1996) Identification of the snake venom substance that induces apoptosis. Biochem Biophys Res Commun 224(1):134–139
Treshalina HM, Lukasheva EV, Sedakova LA, Firsova GA, Guerassimova GK, Gogichaeva NV, Berezov TT (2000) Anticancer enzyme l-lysine α-oxidase. Appl Biochem Biotechnol 88(1):267–273
Yu Z, Qiao H (2012) Advances in non-snake venom l-amino acid oxidase. Appl Biochem Biotechnol 167:1–13
Yu Z, Zhou N, Zhao C, Qiu J (2013) In-gel determination of l-amino acid oxidase activity based on the visualization of Prussian blue-forming reaction. PLoS One 8(2):e55548
Yu Z, Wang Y, Zhao M, Qiu J, Lin J (2014a) Advances in detection methods of l-amino acid oxidase activity. Appl Biochem Biotechnol 174(1):13–27
Yu Z, Zhou N, Qiao H, Qiu J (2014b) Identification, cloning and expression of l-amino acid oxidase from marine Pseudoalteromonas sp. B3. Sci World J 201:979858
Yu Z, Wang J, Lin J, Zhao M, Qiu J (2015a) Exploring regulation genes involved in the expression of l-amino acid oxidase in Pseudoalteromonas sp. Rf-1. PLoS One 10(3):e0122741
Yu Z, Zhao M, Qiu J (2015b) Draft genome sequence of Pseudoalteromonas sp. strain R3, a red pigmented l-amino acid oxidase-producing bacterium. Genome Announc 3(6):e01339-15
Zhang YJ, Wang JH, Lee WH, Wang Q, Liu H, Zheng YT, Zhang Y (2003) Molecular characterization of Trimeresurus stejnegeri venom l-amino acid oxidase with potential anti-HIV activity. Biochem Biophys Res Commun 309(3):598–604
Acknowledgements
We thank Dr. Yili Huang at Zhejiang University for critical reading and revision on manuscript. This study was supported by National Natural Science Foundation of China (31670114) and Zhejiang Provincial Natural Science Foundation of China (LY16C010002), for which the authors are thankful.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
We declare that we have no conflict of interest.
Rights and permissions
About this article
Cite this article
Yu, Z., Yang, Y., Wang, Y. et al. Reactive oxygen species-scavenging system is involved in l-amino acid oxidase accumulation in Pseudoalteromonas sp. B3. 3 Biotech 7, 326 (2017). https://doi.org/10.1007/s13205-017-0976-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-017-0976-2