Abstract
Alpha-(1,2)-fucosyltransferase (FUT1) gene has some influence on economically important traits and disease resistance. DNA methylation plays an important role in human diseases but is relatively poorly studied in pigs by regulating the mRNA expression of genes. The aim of this study was to analyze the influence of promoter methylation on the expression of FUT1 gene. We used bisulfite sequencing PCR (BSP) and qPCR to analyze the methylation of the FUT1 5′-flanking region and FUT1 mRNA expression in the duodenum of Sutai piglets from newborn to weaning. FUT1 contains three CpG islands upstream of the start codon, of which two are located in the putative promoter region containing multiple promoter elements and transcription factor binding sites, such as CpG islands, a CAAT box, SP1, and EARLY-SEQ 1. The CpG island between nucleotides −1762 and −580 had a low degree of methylation, and its methylation level was significantly lower in 35-day-old piglets than 8- and 18-day-old piglets (P < 0.05). FUT1 mRNA expression was significantly higher in 35-day-old piglets than 8- and 18-day-old piglets (P < 0.05). Pearson’s correlation analysis showed that the methylation of the CpG island between nucleotides −1762 and −580 of FUT1 was significantly, negatively correlated with FUT1 mRNA expression (P < 0.05). These results demonstrate that differential methylation of CpG islands negatively regulates the expression of FUT1 in the porcine duodenum, suggesting a probable influence on the resistance of piglets to infection with ETEC F18.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Enterotoxigenic Escherichia coli is the pathogen most frequently responsible for diarrhea and edema in newborn and weaned piglets, and has become a major threat in the pig farming industry (Boldin 2008). Currently, the ETEC strain F18 is one of the most common and most hazardous E. coli pathogens in the pig farming industry (Van den Broeck et al. 2000). Genetic studies have revealed that the adhesion-mediated adhesion and colonization ability of ETEC F18 bacteria in piglets is dependent upon the presence of appropriate receptors for F18 cohesin in the brush borders on epithelial cells in the piglet intestine; when the F18 strain is bound to these receptors, the toxin produced by the bacteria can induce diarrhea and edema in pigs (Bertschinger and Pohlenz 1983; Imberechts et al. 1996). Using a candidate gene approach and linkage analysis, Vogeli et al. (1997) discovered that the α-(1,2) fucosyltransferase gene 1 (FUT1) is a candidate gene which can control the adhesion to ETEC F18 receptor. In a study in pigs in Switzerland, Meijerink et al. (1997) reported that the variation of FUT1 gene on M307 locus is correlated with resistance to ETEC F18, and successfully achieved resistance to ETEC F18 by breeding pigs with this marker. FUT1 and its M307 locus are one of the few resistance genes and resistance markers for controlling the expression of the receptor for ETEC F18 to be identified in pigs to date. Studies have shown that porcine FUT1 has some influence on economically important traits and disease resistance (Bao et al. 2012a, b; Zhang et al. 2007). Based on this research, we and other Chinese researchers have systematically analyzed the FUT1-coding region in nearly 30 Chinese native pig breeds and foreign pig breeds. The variation of FUT1 gene on M307 locus, which has been used as a genetic marker in foreign breeds, displays an extremely skewed distribution in native Chinese pig breeds compared to foreign breeds (Bao et al. 2008; Yan et al. 2003), and the presence or absence of the M307 locus does not significantly influence the expression level of FUT1 (Bao et al. 2012a, b). This makes the development of resistance to ETEC F18 in local Chinese pig breeds difficult and problematic. Therefore, screening for other candidate ETEC F18 resistance genes in local Chinese pig breeds is required, and additionally, the mechanism by which the non-coding regions of the FUT1 gene, especially the promoter region, offer resistance to ETEC F18 urgently requires investigation.
DNA methylation, a critical component of epigenetic control, plays an important role in cell differentiation, embryonic development, gene regulation, genomic imprinting, and X-chromosome inactivation (Reik et al. 2001; Ehrlich 2003; Li 2002; Bird 2002; Li et al. 1993; Heard 2004). Sullivan et al. (2012) revealed that hypermethylation of the insulin-like growth factor binding protein 7 (IGFBP7) promoter can lead to gene silencing. Many human diseases, such as coronary artery disease, Alzheimer’s disease, and cancer, are closely linked to altered promoter methylation (Kim et al. 2011; Friso et al. 2012; Silva et al. 2008); however, reports of the association between diseases and altered promoter methylation in pigs are relatively rare. Kang et al. (2007) suggested that dynamic methylation changes may play an important role in the establishment and maintenance of tissue-specific gene expression in cloned pigs. Xiao et al. (2012) analyzed the promoter methylation status and gene expression levels of the tyrosine protein kinase lyn (LYN) gene and demonstrated that methylation plays an important role in the regulation of porcine LYN gene transcription. Ling et al. (2009) revealed that the methylation status of the porcine adiponectin gene promoter dynamically fluctuates in a manner which correlates with adiponectin gene expression during development.
The promoter, an integral upstream regulatory region of a gene, controls the initiation of gene transcription as well as the gene expression level. The promoter regulates gene expression through a number of mechanisms, including promoter methylation (Juven-Gershon and Kadonaga 2010). Thus, intensive study of promoter methylation is essential to fully understand the mechanisms of gene regulation. As FUT1 is the most important candidate gene for the breeding of porcine ETEC F18 disease resistance, research on the potential methylation modifications which the FUT1 promoter undergoes will provide a new approach for investigating the mechanisms by which FUT1 confers resistance to ETEC F18. Nevertheless, data on the porcine FUT1 5′-flanking are absent in the authoritative database (GenBank); therefore, it is difficult to initiate studies on the FUT1 promoter. In a previous study, we cloned the FUT1 promoter sequence (using duodenum DNA of Sutai piglets by Genome Walking) and compared the sequences (see sequence U70883.2 in GenBank) via a high-throughput approach and by Blastn analysis. We retrieved the sequence 2000 bp upstream of the start codon of FUT1. In this study, we investigated the methylation status of the FUT1 5′-flanking and explored the influence of FUT1 promoter methylation on the gene expression levels of FUT1 in the duodenum of Sutai piglets from the newborn stage to weaning (8, 18, and 35 days old). This study provides a foundation for further studies of the mechanisms regulating porcine FUT1 expression.
Materials and methods
Experimental reagents
The EpiTect Bisulfite Kit was purchased from Qiagen (Valencia, CA, USA), TRIzol reagent from Invitrogen (Carlsbad, CA, USA), and Taq DNA polymerase, dNTPs, PCR purification kit, Plasmid DNA Extract kit, Competent Escherichia coli cell, pMD18-T vector, and SYBR Green Real-time PCR Mix from TaKaRa (Dalian, China). All the other chemicals were of analytical reagent grade.
Bioinformatic analysis and primer design
Analysis and identification of the CpG islands, transcription initiation site, and putative promoter region in the 5′-flanking of FUT1 were performed using the online tools MethPrimer (http://www.urogene.org/methprimer/index1.html), NNPP (http://www.fruitfly.org/seq_tools/promoter.html), PROMOTER 2.0 (http://www.cbs.dtu.dk/services/Promoter), FirstEF (http://rulai.cshl.org/tools/FirstEF/), and TFSEARCH (http://www.cbrc.jp/research/db/TFSEARCH.html). Based on this analysis and the location of the predicted CpG islands, bisulfite-sequencing PCR (BSP) primers were designed using Methyl Primer Express 1 software (Table 1). All primers were synthesized by Sangon (Shanghai, China).
Experimental animal
Sutai pig is a new breed of high-quality lean-meat-type pig bred by Sutai pig breeding center in Suzhou city. It is a breed developed from a Duroc (50%) × Meishan (50%) cross after 15 years of practice. In 1999, it was approved by National Committee of Livestock and Poultry Species for a new variety. All the pigs (Sutai pigs, 8, 18, and 35 days old, n = 5 each) included in the experiment were healthy and were, respectively, selected from five different families, with same feeding condition, similar birth weights, weaning weights, and body sizes. The pigs were purchased from Suzhou Sutai Pig Breeding Center (Suzhou, China) and all experiments were conducted at the Animal Hospital of Yangzhou University according to the regulation for the Administration of Affairs Concerning Experimental Animals (Ministry of Science and Technology, China, revised in June 2012) and approved by the experimental animal using permit with Permit No. SYXK (Su) 2012-0029.
Real-time PCR analysis
RNA was isolated from the duodenal tissues of the Sutai pigs using TRIzol reagent, according to the manufacturer’s instructions. Single-stranded cDNA was generated using the PrimeScript RT-PCR Kit following the manufacturer’s directions. Real-time quantitative PCR was performed using an ABI Prism 7500 sequence-detection system (Applied Biosystems, Foster City, CA, USA) with SYBR Green PCR Master Mix, according to the manufacturer’s instructions. The primer sequences for FUT1 were: F, 5′-TTTTAAGCCCCCAAACTGCC-3′ and R, 5′-TAAATCGACCCCATCAGCCTC-3′. The GAPDH primers used as internal control and thermocycler conditions are described in our previous study (Bao et al. 2012a, b), the expression of FUT1 in each sample was normalized to GAPDH. Triplicate PCR amplifications were performed for each sample. The fold change in relative gene expression was calculated using the standard 2−ΔΔCt method (Livak and Schmittgen 2001).
Promoter methylation analysis
Genomic DNA was extracted from porcine duodenal tissues by standard phenol/chloroform extraction and subjected to bisulfite conversion using the EpiTect Bisulfite Kit, according to the manufacturer’s instructions. Touchdown PCR was used to amplify the bisulfite-treated DNA (BST-DNA). The 50-μL reactions included 3.0 μL of DNA template, 3 μL of 10 × PCR buffer, 2 μL of Mg2+ (25 mmol), 1 μL of forward primer (10 μM), 1 μL of reverse primer (10 μM), 1 μL of dNTPs (10 mmol), 0.8 μL of Taq polymerase (5 U/μL) and 38.2 μL of water. The following reaction conditions were used: 98 °C for 4 min, then 20 cycles of 94 °C for 45 s, 66 °C for 45 s (which reduced by 0.5 °C each cycle), and 72 °C for 1 min; 20 cycles of 94 °C for 45 s, 56 °C for 45 s and 72 °C for 1 min, and a final extension at 72 °C for 10 min. The PCR products were subjected to electrophoresis on agarose gels, excised, purified, and inserted into the pMD18-T vector. The recombinant clones were used to transform E.coli TB1 cells. The positive recombinant clones were selected on LB agar plates containing 100 μg/mL ampicillin and confirmed by PCR and DNA sequencing (five positive recombinant clones were selected from each individual). The methylation rate was calculated as the number of CpG methylated loci/number of CpG loci.
Results
Analysis of the CpG islands, promoter elements, and transcription factor binding sites in the 5′-flanking of porcine FUT1
Analysis of the FUT1 gene 5′-flanking region (2000 bp upstream of the initiation codon) revealed that porcine FUT1 has an intron-cut 5′-flanking and FUT1 contains only one intron (Fig. 1). CpG islands in the porcine FUT1 5′-flanking were identified using the online software MethPromoter and Methyl Promoter Express software. The FUT1 5′-flanking contains three CpG islands: CpG island 1 is located between nucleotides −1779 and −886 with a length of 894 bp, CpG island 2 is located between nucleotides −796 and −619 with a length of 178 bp, and CpG island 3 is located between nucleotides −316 and −130 with a length of 187 bp. CpG islands 1 and 2 are located within the putative promoter region and CpG island 3 is located within intron 1 (Fig. 1). Online analysis using the tools NNPP, PROMOTER 2.0, FirstEF, and TFSEARCH showed that the putative FUT1 promoter region contains multiple promoter element characteristics and transcription factor binding sites, including CpG islands, a CAAT box, EARLY-SEQ 1, T-Ag, PuF, and SP1 (Fig. 2).
Prediction of cis-acting elements and transcription factor binding sites in the 5′-flanking region of porcine FUT1. In this core promoter region, the CpG island sequence was in the box and the non-CpG island sequence was in other area. All CG sites were highlighted with a yellow background and all methylated C sites were highlighted in bold red font. The underline represents a possible transcription factor binding site
Analysis of the methylation status of the putative promoter region of porcine FUT1
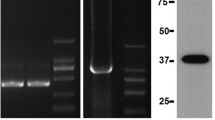
Five primer pairs were designed to examine the levels of methylation in the FUT1 5′-flanking (Fig. 1). The products produced by each primer pair from the amplification of cDNA extracted from the pig duodenum were examined by 1% agarose gel electrophoresis. The size of the amplified fragments corresponded with the expected product sizes for each primer pair and each primer pair amplified a single specific product which could be directly cloned and sequenced (Supplementary Fig. 1).
The methylation status of the putative FUT1 promoter region was examined in the duodenum of piglets of three different ages. As shown in Fig. 3 and Table 2, CpG methylation sites were ubiquitously present in the porcine putative FUT1 promoter region. The non-CpG island (from −2174 to −1762) was highly methylated, with an average methylation rate of 74.81%. The methylation status of the non-CpG island in the putative FUT1 promoter region was not significantly different among the three ages of piglets tested. The CpG island in the putative FUT1 promoter region (from −1762 to −580) had a low degree of methylation, with an average methylation rate of 3.82%. Although the degree of methylation of the CpG island in the putative FUT1 promoter region was low, its methylation status varied with the age of the piglets: the degree of methylation in the duodenum of 8-day-old piglets was 4.1%, which increased slightly (4.79%) but non-significantly compared to 18-day-old piglets. The degree of methylation drastically reduced in 35-day-old piglets (2.56%) and was significantly lower than that in 18-day-old and 8-day-old piglets (P < 0.05).
Analysis of the methylation status of the FUT1 5′-flanking region in the duodenum of piglets of different ages. Circles CpGs, black circles methylated CpGs, and white circles unmethylated CpGs; FUT1-1 to 5 the five primer pairs for bisulfite sequencing PCR. Every circle represents a CG, black circles methylated CG, and white circles unmethylated CG
Correlation between the methylation status of the putative promoter region and the expression levels of porcine FUT1
Quantitative real-time PCR was used to measure the expression of FUT1 in the duodenum of piglets at three different ages. The relative expression level of FUT1 decreased between 8-day-old and 18-day-old and then increased to the highest levels in 35-day-old piglets (P < 0.05; Table 2). Pearson’s correlation analysis indicated that the methylation status of the CpG island in the putative promoter region significantly, negatively correlated with the expression level of FUT1 in the duodenum as the piglets aged (Pearson’s correlation coefficient is −0.999, P < 0.05). There was no significant correlation between the methylation status of the non-CpG island in the putative promoter region and the expression of FUT1 in the porcine duodenum (Pearson’s correlation coefficient is 0.545, P > 0.05).
Discussion
FUT1 controls the expression of ETEC F18 and is one of the few candidate anti-diarrhea genes to have been identified in piglets (Vogeli et al. 1997; Meijerink et al. 1997). Many studies have studied the coding region of porcine FUT1; however, few studies have focused on the FUT1 5′-flanking region or promoter region due to an absence of sequence data on this region in authoritative databases. In a previous study, we retrieved the sequence of the region 2000 bp upstream of the FUT1 start codon using gene walking and high-throughput comparison (see sequence U70883.2 in GenBank), which provided a basis for further experiments. Promoter regions can be divided into CpG islands and non-CpG islands according to their CpG content. In this study, we first analyzed the 5′-flanking of FUT1 and found that the putative promoter region contains one non-CpG island (−2174 to −1762) and one CpG island (−1762 to −580).
Suzuki and Bird (2008) reported that the highly dynamic process of promoter methylation is an important regulation mechanism for normal developmental processes. Such dynamic methylation processes are particularly evident during embryonic development (Santos et al. 2002) and methylation plays a key role in the reprogramming of the genome in the early mammalian embryo (Thurston et al. 2007; Haaf 2006). In the present study, we observed that the methylation status of the putative promoter region of FUT1 altered dynamically as the piglets aged, with a large difference between the non-CpG island and CpG island. From the newborn stage to the weaning period in piglets, the non-CpG island remained highly methylated and its methylation status did not change significantly. The methylation status of the non-CpG island did not significantly correlate with the FUT1 expression level in the duodenum. This indicates that although the non-CpG island has a high degree of methylation, its methylation status does not significantly affect FUT1 expression. The CpG islands in the putative FUT1 promoter region had a low degree of methylation and its degree of methylation varied remarkably as the piglets aged; the degree of methylation in 35-day-old piglets was significantly lower than 8- and 18-day-old piglets. Real-time PCR demonstrated that the FUT1 expression level was significantly higher in 35-day-old piglets than 8- and 18-day-old piglets and the degree of methylation in the CpG islands was significantly, negatively correlated with the FUT1 mRNA expression level. This indicates that methylation of the CpG island in the putative FUT1 promoter region negatively regulates FUT1 gene expression.
Pigs of different ages usually display different degrees of sensitivity to disease, for example, newborn piglets are vulnerable to infection with E. coli strain K88, whereas 35-day-old piglets are susceptible to infection with ETEC F18 (Jones and Rutter 1972; Verdonck et al. 2002). In the present study, we observed that expression level of FUT1 was significantly higher in the duodenum of 35-day-old piglets than 8-day-old piglets, suggesting that the FUT1 expression level is associated with the susceptibility to infection by ETEC F18. Our preliminary studies and other studies suggest that the FUT1 expression level directly influences the sensitivity of piglets to ETEC F18 (Meijerink et al. 1997; Bao et al. 2012a, b); these findings suggest that methylation of the CpG island in the putative FUT1 promoter region may affect the resistance of piglets to ETEC F18. Besides, we found that methylation of CpG in FUT1-2 was a little different in 8 days, 18 days, and 35 days, which suggests that specific CpG sites may contribute more than others to regulate FUT1 mRNA expression. However, we found that these methylated CG sites were not located in the transcription factor binding sites of the core promoter region of FUT1 gene, so how they regulated the expression of FUT1 gene needed further studies.
In a previous study, we preliminarily explored the mechanisms regulating FUT1 expression and analyzed the expression level of the FUT1 M307 locus in different genotypes of Sutai pigs. The results showed that the mutation of the M307 locus (G/A) does not have a significant influence on the expression of FUT1 (Bao et al. 2012a, b). The results of this study further suggest that the regulatory mechanism controlling the response to ETEC F18 may possibly involve altered methylation of the CpG islands in the putative FUT1 promoter region.
There are two mechanisms by which promoter methylation affects gene regulation. First, the methylation of essential cis-acting elements in the promoter region influences the binding of transcription factors (Kass et al. 1997). Second, co-suppression factor complexes, including histone deacetylase, are recruited by the methylated cis-acting elements via methyl-CpG-binding proteins, which inhibit transcription (Nan et al. 1998; Magnaghi-Jaulin et al. 1998). Our bioinformatic analysis indicates that the putative FUT1 promoter region contains a number of transcription factor binding sites, such as EARLY-SEQ 1, PuF, and SP1-binding sites, within the CpG islands. SP1 is a key transcription factor required for mRNA transcription, and SP1-binding sites contain a CpG site (Holler et al. 1988). Methylation of CpG sites can directly influence the ability of SP1 to bind DNA and influence gene expression; likewise, EARLY-SEQ 1 also contains a CpG site. Methylation of the CpG sites within these transcription factor binding sites is likely to directly influence the expression of FUT1. Gonzalgo et al. (1998) reported that a certain ratio of CpG island methylation (>60%) can be sufficient to inhibit gene expression, whereas a lower ratio of methylation reduces the transcription and expression of genes. Thus, it is not necessary for all CpG sites to be methylated to inhibit gene expression; rather a regulatory effect can be achieved by controlling the degree of methylation in the binding sites of several transcription factors.
This study has a number of limitations. The methylation status of FUT1 should be examined in porcine tissues other than the duodenum, and larger numbers of samples should be tested to confirm the relationship between CpG island methylation and FUT1 expression. Further intensive analysis of the upstream control elements in the putative FUT1 promoter region is required to identify the key methylation sites which regulate FUT1 expression.
Conclusions
In the piglet duodenum, the low levels of methylation of the CpG island in the putative FUT1 promoter region significantly, negatively correlate with FUT1 expression as the piglets age. Methylation of the CpG island in the putative FUT1 promoter region may possibly play a role in regulating the resistance of pigs to ETEC F18.
References
Bao WB, Wu SL, Musa HH, Zhu GQ, Chen GH (2008) Genetic variation at the alpha-1-fucosyltransferase (FUT1) gene in Asian wild boar and Chinese and Western commercial pig breeds. J Anim Breed Genet 125:427–430
Bao WB, Ye L, Pan ZY, Zhu J, Du ZD, Zhu GQ, Huang XG, Wu SL (2012a) The effect of mutation at M307 in FUT1 gene on susceptibility of Escherichia coli F18 and gene expression in Sutai piglets. Mol Biol Rep 39:3131–3136
Bao WB, Ye L, Zhu J, Pan ZY, Zhu GQ, Huang XG, Wu SL (2012b) Evaluation of M307 of FUT1 gene as a genetic marker for disease resistance breeding of Sutai pigs. Mol Biol Rep 39:4223–4228
Bertschinger HU, Pohlenz J (1983) Bacterial colonization and morphology of the intestine in porcine Escherichia coli enterotoxemia (edema disease). Vet Pathol 20:99–110
Bird A (2002) DNA methylation patterns and epigenetic memory. Gene Dev 16:6–21
Boldin B (2008) Persistence and spread of gastro-intestinal infections: the case of enterotoxigenic Escherichia coli in piglets. B Math Biol 70:2077–2101
Ehrlich M (2003) Expression of various genes is controlled by DNA methylation during mammalian development. J Cell Biochem 88:899–910
Friso S, Lotto V, Choi SW, Girelli D, Pinotti M, Guarini P, Udali S, Pattini P, Pizzolo F, Martinelli N, Corrocher R, Bernardi F, Olivieri O (2012) Promoter methylation in coagulation F7 gene influences plasma FVII concentrations and relates to coronary artery disease. J Med Genet 49:192–199
Gonzalgo ML, Hayashida T, Bender CM, Pao MM, Tsai YC, Gonzales FA, Nguyen HD, Nguyen TT, Jones PA (1998) The role of DNA methylation in expression of the p19/p16 locus in human bladder cancer cell lines. Cancer Res 58:1245–1252
Haaf T (2006) Methylation dynamics in the early mammalian embryo: implications of genome reprogramming defects for development. Curr Top Microbiol 310:13–22
Heard E (2004) Recent advances in X-chromosome inactivation. Curr Opin Cell Biol 16:247–255
Holler M, Westin G, Jiricny J, Schaffner W (1988) Sp1 transcription factor binds DNA and activates transcription even when the binding site is CpG methylated. Gene Dev 2:1127–1135
Imberechts H, Wild P, Charlier G, De Greve H, Lintermans P, Pohl P (1996) Characterization of F18 fimbrial genes fedE and fedF involved in adhesion and length of enterotoxemic Escherichia coli strain 107/86. Microb Pathogenesis 21:183–192
Jones GW, Rutter JM (1972) Role of the K88 antigen in the pathogenesis of neonatal diarrhea caused by Escherichia coli in piglets. Infect Immun 6:918–927
Juven-Gershon T, Kadonaga JT (2010) Regulation of gene expression via the core promoter and the basal transcriptional machinery. Dev Biol 339:225–229
Kang JK, Park KW, Chung YG, You JS, Kim YK, Lee SH, Hong SP, Choi KM, Heo KN, Seol JG, Lee JH, Jin DI, Park CS, Seo JS, Lee HW, Han JW (2007) Coordinated change of a ratio of methylated H3lysine 4 or acetylated H3 to acetylated H4 and DNA methylation is associated with tissue-specific gene expression in cloned pig. Exp Mol Med 39:84–96
Kass SU, Pruss D, Wolffe AP (1997) How does DNA methylation repress transcription? Trends Genet 13:444–449
Kim DS, Lee JY, Lee SM, Choi JE, Cho S, Park JY (2011) Promoter methylation of the RGC32 gene in nonsmall cell lung cancer. Cancer 117:590–596
Li E (2002) Chromatin modification and epigenetic reprogramming in mammalian development. Nat Rev Genet 3:662–673
Li E, Beard C, Janisch AR (1993) Role for DNA methylation in genomic imprinting. Nature 366:362–365
Ling F, Li JQ, Wang C, Liu C, DU H, Xiao ZZ, Wang LL, Chen YS (2009) Methylation status of pig adiponectin promoter and its expression. Hereditas 31:1013–1019
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using realtime quantitative PCR and the 2−ΔΔCt method. Methods 25:402–408
Magnaghi-Jaulin AL, Groisman R, Naguibneva I, Robin P, Lorain S, Le Villain JP, Troalen F, Trouche D, Harel-Bellan A (1998) Retinoblastoma protein represses transcription by recruiting a histone deacetylase. Nature 391:601–605
Meijerink E, Fries R, Vögeli P, Masabanda J, Wigger G, Stricker C, Neuenschwander S, Bertschinger HU, Stranzinger G (1997) Two α(1,2) fucosyltransferase genes on porcine chromosome 6q11 are closely linked to the blood group inhibitor (S) and Escherichia coli F18 receptor (ECF18R) loci. Mamm Genome 8:736–741
Nan XS, Ng HH, Johnson CA, Laherty CD, Turner BM, Eisenman RN, Bird A (1998) Transcriptional repression by the methyl-CpG-binding protein MeCP2 involves a histone deacetylase complex. Nature 393:386–389
Reik W, Dean W, Walter J (2001) Epigenetic reprogramming in mammalian development. Science 293:1089–1093
Santos F, Hendrich B, Reik W, Dean W (2002) Dynamic reprogramming of DNA methylation in the early mouse embryo. Dev Biol 241:172–182
Silva PN, Gigek CO, Leal MF, Bertolucci PH, de Labio RW, Payão SL, Smith Mde A (2008) Promoter methylation analysis of SIRT3, SMARCA5, HTERT and CDH1 genes in aging and Alzheimer’s disease. J Alzheimers Dis 13:173–176
Sullivan L, Murphy TM, Barrett C, Loftus B, Thornhill J, Lawler M, Hollywooda D, Lynchd T, Perrya Antoinette S (2012) IGFBP7 promoter methylation and gene expression analysis in prostate cancer. J Urology 188:1354–1360
Suzuki MM, Bird A (2008) DNA methylation landscapes: provocative insights from epigenomics. Nat Rev Genet 9:465–476
Thurston A, Lucas ES, Allegrucci C, Steele W, Young LE (2007) Regionspecific DNA methylation in the preimplantation embryo as a target for genomic plasticity. Theriogenology 68(Suppl 1):S98–S106
Van den Broeck W, Cox E, Oudega B, Goddeeris BM (2000) The F4 fimbrial antigen of Escherichia coli and its receptors. Vet Microbiol 71:223–244
Verdonck F, Cox E, van Gog K, Van der Stede Y, Duchateau L, Deprez P, Goddeeris BM (2002) Different kinetic of antibody responses following infection of newly weaned pigs with an F4 enterotoxigenic Escherichia coli strain or an F18 verotoxigenic Escherichia coli strain. Vaccine 20:2995–3004
Vogeli P, Meijerink E, Fries R, Neuenschwander S, Vorländer N, Stranzinger G, Bertschinger HU (1997) A molecular test for the detection of E. coli F18 receptors: a breakthrough in the struggle against oedema disease and postweaning diarrhoea in swine. Schweiz Arch Tierh 139:479–484
Xiao Z, Wang C, Mo D, Li J, Chen Y, Zhang Z, Cong P (2012) Promoter CpG methylation status in porcine Lyn is associated with its expression levels. Gene 511:73–78
Yan XM, Ren J, Guo YM, Ding NS, Chen KF, Gao J, Ai HS, Chen CY, Ma JW, Huang LS (2003) Research on the genetic variations of al-fucosytransferase (FUT1) gene in 26 pig breeds. Acta Genetica Sinica 30:830–834
Zhang YH, Zhou ZX, Cao GQ (2007) FUT1 gene polymorphism and its correlation with litter size traits. Hereditas 29:52–56
Acknowledgements
This research was funded by the Natural Science Foundation of the Jiangsu Higher Education Institutions of China (14KJA230003), the National Natural Science Funds (31172183), the Genetically Modified Organisms Technology Major Project (No. 2014ZX0800601B), and the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest in the publication.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made.
The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
To view a copy of this licence, visit https://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dai, C., Sun, L., Xia, R. et al. Correlation between the methylation of the FUT1 promoter region and FUT1 expression in the duodenum of piglets from newborn to weaning. 3 Biotech 7, 247 (2017). https://doi.org/10.1007/s13205-017-0880-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-017-0880-9