Abstract
Wolbachia is a common group of intracellular bacteria found in arthropods and filarial nematodes. Since the past decade, they have attracted considerable interest owing to their various effects on hosts, which range from reproductive manipulation to mutualism. Moreover, they can influence the mitochondrial DNA pattern which do not reflect the real evolutionary history of the target species and may be incongruent with nuclear data. Previously, Wolbachia-manipulated mitochondrial DNA (mtDNA) patterns, namely mito-nuclear discordance and deep mitochondrial splits associated with specific Wolbachia infections, have been also discovered in the genus Maculinea. Here, we present a comprehensive study on Wolbachia infestation and the genetic diversity of all Maculinea species in the Carpathian Basin. The prevalence and the pattern of the infestation highly differ among Maculinea species. Maculinea alcon and Maculinea arion are infected in 100 %, each of these species with a single strain, but the infection level of Maculinea nausithous and Maculinea teleius is much lower, additionally, they are infected with multiple strains. The genetic diversity of Maculinea species proved to be generally low, only M. nausithous showed geographic pattern based on mitochondrial sequences and allozymes. In contrast with the previous studies, we could not detect mito-nuclear discordance or find evidence for Wolbachia-induced selective sweep. Based on our results, we cannot hold only Wolbachia responsible for the restricted genetic diversity of Maculinea in the Carpathian Basin. Probably several factors shape together the level and pattern of genetic variability in Maculinea butterflies.






Similar content being viewed by others
References
Aagaard, K., Hindar, K., Pullin, A. S., James, C. H., Hammarstedt, O., Balstad, T., et al. (2002). Phylogenetic relationships in brown argus butterflies (Lepidoptera: Lycaenidae: Aricia) from northwestern Europe. Biological Journal of the Linnean Society, 75(1), 27–37. doi:10.1046/j.1095-8312.2002.00004.x.
Als, T. D., Vila, R., Kandul, N. P., Nash, D. R., Yen, S.-H., Hsu, Y.-F., et al. (2004). The evolution of alternative parasitic life histories in large blue butterflies. Nature, 432(7015), 386–390. doi:10.1038/nature03020. http://www.nature.com/nature/journal/v432/n7015/suppinfo/nature03020_S1.html.
Andersen, A., Simcox, D. J., Thomas, J. A., & Nash, D. R. (2014). Assessing reintroduction schemes by comparing genetic diversity of reintroduced and source populations: a case study of the globally threatened Large Blue butterfly (Maculinea arion). Biological Conservation, 175, 34–41. doi:10.1016/j.biocon.2014.04.009.
Anderson, R. M., & May, R. M. (1982). Coevolution of hosts and parasites. Parasitology, 85(02), 411–426. doi:10.1017/S0031182000055360.
Aris-Brosou, S., & Excoffier, L. (1996). The impact of population expansion and mutation rate heterogeneity on DNA sequence polymorphism. Molecular Biology and Evolution, 13(3), 494–504.
Baldo, L., Dunning Hotopp, J. C., Jolley, K. A., Bordenstein, S. R., Biber, S. A., Choudhury, R. R., et al. (2006). Multilocus sequence typing system for the endosymbiont Wolbachia pipientis. Applied and Environmental Microbiology, 72(11), 7098–7110. doi:10.1128/aem.00731-06.
Bereczki, J., Pecsenye, K., Peregovits, L., & Varga, Z. (2005). Pattern of genetic differentiation in the Maculinea alcon species group (Lepidoptera, Lycaenidae) in Central Europe. Journal of Zoological Systematics and Evolutionary Research, 43(2), 157–165. doi:10.1111/j.1439-0469.2005.00305.157-165.
Bereczki, J., Tóth, J. P., Tóth, A., Bátori, E., Pecsenye, K. & Varga, Z. (2011). Genetic structure of phenologically differentiated Large Blue (Maculinea arion) populations (Lepidoptera: Lycaenidae) in the Carpathian Basin. European Journal of Entomology, 108(4), 519−527.
Bereczki, J., Tóth, J. P., Sramkó, G., & Varga, Z. (2014). Multilevel studies on the two phenological forms of large blue (Maculinea arion) (Lepidoptera: Lycaenidae). Journal of Zoological Systematics and Evolutionary Research, 52(1), 32–43. doi:10.1111/jzs.12034.
Burns, J. M. (1975). Isozymes in evolutionary systematics. In C. L. Markert (Ed.), Isozymes IV: genetics and evolution (pp. 49–62). New York: Academic.
Cavalli-Sforza, L. L., & Edwards, A. W. F. (1967). Phylogenetic analysis: models and estimation procedures. Evolution, 21, 550–570.
Davies, H. D., & Vinson, S. B. (1986). Passive evasion by eggs of braconid parasitoid Cardiochiles nigriceps of encapsulation in vitro by haemocytes of host Heliothis virescens. Possible role for fibrous layer in immunity. Journal of Insect Physiology, 32(12), 1003–1010. doi:10.1016/0022-1910(86)90119-8.
Dupuis, J. R., Roe, A. D., & Sperling, F. A. H. (2012). Multi-locus species delimitation in closely related animals and fungi: one marker is not enough. Molecular Ecology, 21(18), 4422–4436. doi:10.1111/j.1365-294X.2012.05642.x.
Dyson, E. A., Kamath, M. K., & Hurst, G. D. D. (2002). Wolbachia infection associated with all-female broods in Hypolimnas bolina (Lepidoptera: Nymphalidae): evidence for horizontal transmission of a butterfly male killer. Heredity, 88(3), 166–171.
Earl, D., & vonHoldt, B. (2012). STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conservation Genetics Resources, 4(2), 359–361. doi:10.1007/s12686-011-9548-7.
Evanno, G., Regnaut, S., & Goudet, J. (2005). Detecting the number of clusters of individuals using the software structure: a simulation study. Molecular Ecology, 14(8), 2611–2620. doi:10.1111/j.1365-294X.2005.02553.x.
Fine, P. E. M. (1975). Vectors and vertical transmission: an epidemiologic perspective. Annals of the New York Academy of Sciences, 266(1), 173–194. doi:10.1111/j.1749-6632.1975.tb35099.x.
Fujii, Y., Kageyama, D., Hoshizaki, S., Ishikawa, H., & Sasaki, T. (2001). Transfection of Wolbachia in Lepidoptera: the feminizer of the adzuki bean borer Ostrinia scapulalis causes male killing in the Mediterranean flour moth Ephestia kuehniella. Proceedings of the Royal Society of London, Series B: Biological Sciences, 268(1469), 855–859. doi:10.1098/rspb.2001.1593.
Galtier, N., Nabholz, B., Glémin, S., & Hurst, G. D. D. (2009). Mitochondrial DNA as a marker of molecular diversity: a reappraisal. Molecular Ecology, 18(22), 4541–4550. doi:10.1111/j.1365-294X.2009.04380.x.
Hammer, Ø., Harper, D. A. T., & Ryan, P. D. (2001). PAST: paleontological statistics software package for education and data analysis. Paleontologica Electronica, 4(1), 9.
Hilgenboecker, K., Hammerstein, P., Schlattmann, P., Telschow, A., & Werren, J. H. (2008). How many species are infected with Wolbachia? – a statistical analysis of current data. FEMS Microbiology Letters, 281(2), 215–220. doi:10.1111/j.1574-6968.2008.01110.x.
Hoffmann, A. A., Hercus, M., & Dagher, H. (1998). Population dynamics of the Wolbachia infection causing cytoplasmic incompatibility in Drosophila melanogaster. Genetics, 148(1), 221–231.
Hu, J., Zhu, X.-X., & Fu, W.-J. (2003). Passive evasion of encapsulation in Macrocentrus cingulum Brischke (Hymenoptera: Braconidae), a polyembryonic parasitoid of Ostrinia furnacalis Guenée (Lepidoptera: Pyralidae). Journal of Insect Physiology, 49(4), 367–375. doi:10.1016/S0022-1910(03)00021-0.
Jiggins, F. M. (2003). Male-killing Wolbachia and mitochondrial DNA: selective sweeps, hybrid introgression and parasite population dynamics. Genetics, 164(1), 5–12.
Kodandaramaiah, U., Simonsen, T. J., Bromilow, S., Wahlberg, N., & Sperling, F. (2013). Deceptive single-locus taxonomy and phylogeography: Wolbachia-associated divergence in mitochondrial DNA is not reflected in morphology and nuclear markers in a butterfly species. Ecology and Evolution, 3(16), 5167–5176. doi:10.1002/ece3.886.
Librado, P., & Rozas, J. (2009). DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics, 25(11), 1451–1452. doi:10.1093/bioinformatics/btp187.
Nei, M. (1987). Molecular evolutionary genetics. New York: Columbia University Press.
Nei, M., & Li, W. H. (1979). Mathematical model for studying genetic variation in terms of restriction endonucleases. Proceedings of the National Academy of Sciences, 76(10), 5269–5273.
Nice, C. C., Gompert, Z., Forister, M. L., & Fordyce, J. A. (2009). An unseen foe in arthropod conservation efforts: the case of Wolbachia infections in the Karner blue butterfly. Biological Conservation, 142(12), 3137–3146. doi:10.1016/j.biocon.2009.08.020.
Paradis, E. (2010). PEGAS: an R package for population genetics with an integrated-modular approach. Bioinformatics, 26(3), 419–420. doi:10.1093/bioinformatics/btp696.
Patricelli, D., Sielezniew, M., Ponikwicka-Tyszko, D., Ratkiewicz, M., Bonelli, S., Barbero, F., et al. (2013). Contrasting genetic structure of rear edge and continuous range populations of a parasitic butterfly infected by Wolbachia. BMC Evolutionary Biology, 13(1), 14.
Peakall, R. O. D., & Smouse, P. E. (2006). Genalex 6: genetic analysis in Excel. Population genetic software for teaching and research. Molecular Ecology Notes, 6(1), 288–295. doi:10.1111/j.1471-8286.2005.01155.x.
Pech, P., Fric, Z., Konvička, M., & Zrzavý, J. (2004). Phylogeny of Maculinea blues (Lepidoptera: Lycaenidae) based on morphological and ecological characters: evolution of parasitic myrmecophily. Cladistics, 20(4), 362–375. doi:10.1111/j.1096-0031.2004.00031.x.
Pecsenye, K., Bereczki, J., Tihanyi, B., Tóth, A., Peregovits, L., & Varga, Z. (2007). Genetic differentiation among the Maculinea species (Lepidoptera: Lycaenidae) in eastern Central Europe. Biological Journal of the Linnean Society, 91(1), 11–21.
Pritchard, J. K., Stephens, M., & Donnelly, P. (2000). Inference of population structure using multilocus genotype data. Genetics, 155(2), 945–959.
QGIS Development Team (2014). QGIS Geographic Information System. Open Source Geospatial Foundation Project.
R Development Core Team. (2014). R: a language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing.
Ritter, S., Michalski, S. G., Settele, J., Wiemers, M., Fric, Z. F., Sielezniew, M., et al. (2013). Wolbachia infections mimic cryptic speciation in two parasitic butterfly species, Phengaris teleius and P. nausithous (Lepidoptera: Lycaenidae). Plos One, 8(11), e78107. doi:10.1371/journal.pone.0078107.
Ronquist, F., Teslenko, M., van der Mark, P., Ayres, D. L., Darling, A., Höhna, S., et al. (2012). MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Systematic Biology, 61(3), 539–542. doi:10.1093/sysbio/sys029.
Schmitt, T., & Hewitt, G. M. (2004). The genetic pattern of population threat and loss: a case study of butterflies. Molecular Ecology, 13(1), 21–31. doi:10.1046/j.1365-294X.2004.02020.x.
Schmitt, T., Giessl, A., & Seitz, A. (2003). Did Polyommatus icarus (Lepidoptera: Lycaenidae) have distinct glacial refugia in southern Europe? Evidence from population genetics. Biological Journal of the Linnean Society, 80(3), 529–538. doi:10.1046/j.1095-8312.2003.00261.x.
Shaw, M. R., Stefanescu, C., & Van Nouhuys, S. (2009). Parasitoids of European butterflies. In J. Settele, T. G. Shreeve, M. Konvicka, & H. Van Dyck (Eds.), Ecology of butterflies in Europe (pp. 130–156). Cambridge: Cambridge University Press.
Shoemaker, D., Keller, G., & Ross, K. G. (2003). Effects of Wolbachia on mtDNA variation in two fire ant species. Molecular Ecology, 12(7), 1757–1771. doi:10.1046/j.1365-294X.2003.01864.x.
Sielezniew, M., Rutkowski, R., Ponikwicka-Tyszko, D., Ratkiewicz, M., Dziekańska, I., & Švitra, G. (2012). Differences in genetic variability between two ecotypes of the endangered myrmecophilous butterfly Phengaris (=Maculinea) alcon– the setting of conservation priorities. Insect Conservation and Diversity, 5(3), 223–236. doi:10.1111/j.1752-4598.2011.00163.x.
Smee, M. R. (2011). Population ecology and genetics of the marsh fritillary butterfly Euphydryas aurinia. Exeter: University of Exeter.
Sneath, P. H., & Sokal, R. R. (1973). Numerical taxonomy. San Francisco: W. H. Freeman.
Stephens, M., & Donnelly, P. (2003). A comparison of Bayesian methods for haplotype reconstruction from population genotype data. The American Journal of Human Genetics, 73(5), 1162–1169. doi:10.1086/379378.
Stephens, M., Smith, N. J., & Donnelly, P. (2001). A new statistical method for haplotype reconstruction from population data. The American Journal of Human Genetics, 68(4), 978–989. doi:10.1086/319501.
Tajima, F. (1983). Evolutionary relationship of DNA sequences in finite populations. Genetics, 105(2), 437–460.
Tajima, F. (1989). Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics, 123(3), 585–595.
Tamura, K., Stecher, G., Peterson, D., Filipski, A., & Kumar, S. (2013). MEGA6: molecular evolutionary genetics analysis version 6.0. Molecular Biology and Evolution, 30(12), 2725–2729. doi:10.1093/molbev/mst197.
Tartally, A. (2008). Myrmecophily of Maculinea butterflies in the Carpathian Basin (Lepidoptera: Lycaenidae). PhD, University of Debrecen
Thomas, J. A., Simcox, D. J., & Clarke, R. T. (2009). Successful conservation of a threatened Maculinea butterfly. Science, 325(5936), 80–83. doi:10.1126/science.1175726.
Turelli, M. (1994). Evolution of incompatibility-inducing microbes and their hosts. Evolution, 48(5), 1500–1513. doi:10.2307/2410244.
Turelli, M., & Hoffmann, A. A. (1991). Rapid spread of an inherited incompatibility factor in California Drosophila. Nature, 353(6343), 440–442. doi:10.1038/353440a0.
Turelli, M., & Hoffmann, A. A. (1995). Cytoplasmic incompatibility in Drosophila simulans: dynamics and parameter estimates from natural populations. Genetics, 140(4), 1319–1338.
Ugelvig, L. V., Vila, R., Pierce, N. E., & Nash, D. R. (2011). A phylogenetic revision of the Glaucopsyche section (Lepidoptera: Lycaenidae), with special focus on the Phengaris–Maculinea clade. Molecular Phylogenetics and Evolution, 61(1), 237–243. doi:10.1016/j.ympev.2011.05.016.
Wahlberg, N., & Wheat, C. W. (2008). Genomic outposts serve the phylogenomic pioneers: designing novel nuclear markers for genomic DNA extractions of Lepidoptera. Systematic Biology, 57(2), 231–242. doi:10.1080/10635150802033006.
Watterson, G. A. (1975). On the number of segregating sites in genetical models without recombination. Theoretical Population Biology, 7(2), 256–276. doi:10.1016/0040-5809(75)90020-9.
Werren, J. H., & Windsor, D. M. (2000). Wolbachia infection frequencies in insects: evidence of a global equilibrium? Proceedings of the Royal Society of London, Series B: Biological Sciences, 267(1450), 1277–1285. doi:10.1098/rspb.2000.1139.
Werren, J. H., Baldo, L., & Clark, M. E. (2008). Wolbachia: master manipulators of invertebrate biology. Nature Reviews Microbiology, 6(10), 741–751. doi:10.1038/nrmicro1969.
Wynhoff, I. (1998). Lessons from the reintroduction of Maculinea teleius and M. nausithous in the Netherlands. Journal of Insect Conservation, 2(1), 47–57. doi:10.1023/A:1009692723056.
Yamamura, N. (1993). Vertical transmission and evolution of mutualism from parasitism. Theoretical Population Biology, 44(1), 95–109. doi:10.1006/tpbi.1993.1020.
Zug, R., & Hammerstein, P. (2014). Bad guys turned nice? A critical assessment of Wolbachia mutualisms in arthropod hosts. Biological Reviews. doi:10.1111/brv.12098.
Acknowledgments
The study was supported by the Hungarian Scientific Research Fund (OTKA K109223 and K84071), János Bolyai Scholarship of the Hungarian Academy of Sciences, EU EESD-ENV-2OO2-NAS (Mac Man) project and Social Renewal Operational Programme (TÁMOP-4.2.2/B-10/1-2010-0024). Grateful acknowledgments are due to S. Szabó, A. Tartally, T. Korompai, Z. Ilonczay, A. Ambrus, L. Peregovits, L. Rákosy, F. Rebousek and R. Verovnik for collecting samples. The technical assistance of V. Mester, T. Tóth, K. Kecsmár, H. Boros and B. Prokaj in the molecular work is very much respected. The support of the Nature Conservation Authorities is also greatly appreciated.
Data accessibility
DNA sequences: GenBank accessions: HG326619-HG326646, KM517249-KM517565
Sampling locations and data uploaded as online Supporting Information.
Allozyme genotypes: Dryad doi: 10.5061/dryad.v8q8r
Wolbachia WSP and MLST sequence information uploaded into Wolbachia MLST database.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Additional supporting information may be found in the online version of this article at the publisher’s website.
Online Resource 1
Sampling sites. Abb. – abbreviations of localities. Region – the geographic region from which the samples originate. The table includes sampling time, the number of Maculinea individuals tested for Wolbachia in parentheses and the number of individuals used in allozyme studies in bold. (PDF 238 kb)
Online Resource 2
GenBank accession numbers of the sequenced mitochondrial and nuclear genes of Maculinea butterflies as well as 16S rRNA genes of Wolbachia found in Maculinea species. ID – individual identifiers of Maculinea specimens. The infected individuals are indicated with W. Specimens from which Wolbachia strain identification was carried out are indicated with ‘+’. WSP – Wolbachia surface protein, MLST – Multilocus Sequence Typing based on five genes (gatB, coxA, hcpA, ftsZ, fbpA). (PDF 63 kb)
Online Resource 3
Results of Bayesian inference analyses of single nuclear gene datasets in Maculinea. (a) GAPDH, (b) MDH, (c) wg. (PDF 31 kb)
Online Resource 4
Results of the Bayesian-clustering Structure analysis based on 11 allozyme loci. The most probable K values are indicated in the case of each Maculinea species. See the abbreviations of populations in Online Resource 1. M. alcon: ‘pneumonanthe type’ populations are in bold. M. arion: ‘spring type’ populations are in bold. Populations belonging to the same geographic region are joined together on the upper part of each barplot. (PDF 209 kb)
Online Resource 5
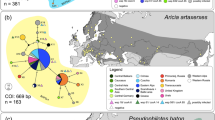
COI haplotype networks for Maculinea nausithous (a) and M. teleius (b) including the sequences of Ritter et al. (2013). Colours show the geographic regions from where haplotypes originate. Circle size is proportional to haplotypes frequency. Dots on lines linking haplotypes indicate the number of mutations. A separate Wolbachia clade is present in both species but there are numerous infected individuals outside this clade. (PDF 98 kb)
Rights and permissions
About this article
Cite this article
Bereczki, J., Rácz, R., Varga, Z. et al. Controversial patterns of Wolbachia infestation in the social parasitic Maculinea butterflies (Lepidoptera: Lycaenidae). Org Divers Evol 15, 591–607 (2015). https://doi.org/10.1007/s13127-015-0217-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13127-015-0217-7