Abstract
Human lung cell lines are utilized widely for investigating tumor biology, experimental therapy, anticancer drug screening and biomarkers identification. However, the consistency of drug responses of these established cell lines and non-small cell lung cancer (NSCLC) is uncertain. In this study, we assessed the drug response consistency between lung cell lines and NSCLC tumors in The Cancer Genome Atlas by hierarchical clustering using copy number variations in driver genes, and profiled the molecular patterns and correlations in cell lines. We found that some frequently used cell lines of NSCLC subtypes were not clustered with their matched subtypes of tumor. Mutation profiles in the oxidative stress response and squamous differentiation pathway in lung cell lines were in concordance with lung squamous cell carcinoma. Furthermore, lung cell lines and tumors in the same sub-cluster had very similar responses to certain drugs but some were inconsistent, suggesting that clustering through copy number variation data could capture part of the suitability of lung cell lines. The analysis of these results could aid investigators in evaluating drug response models and eventually enabling personalized treatment recommendations for individual patients with NSCLC.
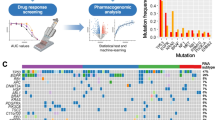
Graphical abstract




Similar content being viewed by others
Data Availability
The data and material are presented within the manuscript.
Code Availability
The codes used in this manuscript were conducted in R packages.
References
Bray F, Ferlay J, Soerjomataram I et al (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 68:394–424. https://doi.org/10.3322/caac.21492
Graham S, Shaban M, Qaiser T et al (2018) Classification of lung cancer histology images using patch-level summary statistics. In: Med Imaging 2018: Digit Pathol. https://doi.org/10.1117/12.2293855
Greenlee RT, Murray T, Bolden S, Wingo PA (2000) Cancer statistics, 2000. CA Cancer J Clin 50:7–33. https://doi.org/10.3322/canjclin.50.1.7
Hutter C, Zenklusen JC (2018) The cancer genome atlas: creating lasting value beyond its data. Cell 173:283–285. https://doi.org/10.1016/j.cell.2018.03.042
Zhang J, Bajari R, Andric D et al (2019) The international cancer genome consortium data portal. Nat Biotechnol 37:367–369. https://doi.org/10.1038/s41587-019-0055-9
Hammerman PS, Lawrence MS, Voet D et al (2012) Comprehensive genomic characterization of squamous cell lung cancers. Nature 489:519–525. https://doi.org/10.1038/nature11404
Collisson EA, Campbell JD, Brooks AN et al (2014) Comprehensive molecular profiling of lung adenocarcinoma. Nature 511:543–550. https://doi.org/10.1038/nature13385
Herbst RS, Morgensztern D, Boshoff C (2018) The biology and management of non-small cell lung cancer. Nature 553:446–454. https://doi.org/10.1038/nature25183
Kim HS, Mitsudomi T, Soo RA, Cho BC (2013) Personalized therapy on the horizon for squamous cell carcinoma of the lung. Lung Cancer 80:249–255. https://doi.org/10.1016/j.lungcan.2013.02.015
Yuan M, Huang L-L, Chen J-H et al (2019) The emerging treatment landscape of targeted therapy in non-small-cell lung cancer. Signal Transduct Target Ther 4:61–61. https://doi.org/10.1038/s41392-019-0099-9
Goodspeed A, Heiser LM, Gray JW, Costello JC (2016) Tumor-derived cell lines as molecular models of cancer pharmacogenomics. Mol Cancer Res 14:3–13. https://doi.org/10.1158/1541-7786.MCR-15-0189
Barretina J, Caponigro G, Stransky N et al (2012) The Cancer cell line encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483:603–607. https://doi.org/10.1038/nature11003
Yang W, Soares J, Greninger P et al (2013) Genomics of drug sensitivity in cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res 41:D955–D961. https://doi.org/10.1093/nar/gks1111
Tate JG, Bamford S, Jubb HC et al (2019) COSMIC: the catalogue of somatic mutations in cancer. Nucleic Acids Res 47:D941–D947. https://doi.org/10.1093/nar/gky1015
Mouradov D, Sloggett C, Jorissen RN et al (2014) Colorectal cancer cell lines are representative models of the main molecular subtypes of primary cancer. Cancer Res 74:3238–3247. https://doi.org/10.1158/0008-5472.CAN-14-0013
Sinha R, Winer AG, Chevinsky M et al (2017) Analysis of renal cancer cell lines from two major resources enables genomics-guided cell line selection. Nat Commun 8:15165. https://doi.org/10.1038/ncomms15165
Landa I, Pozdeyev N, Korch C et al (2019) Comprehensive genetic characterization of human thyroid cancer cell lines: a validated panel for preclinical studies. Clin Cancer Res 25:3141–3151. https://doi.org/10.1158/1078-0432.CCR-18-2953
Najgebauer H, Yang M, Francies HE et al (2020) CELLector: genomics-guided selection of cancer in vitro models. Cell Syst 10:424-432.e6. https://doi.org/10.1016/j.cels.2020.04.007
Zhao N, Liu Y, Wei Y et al (2017) Optimization of cell lines as tumour models by integrating multi-omics data. Brief Bioinform 18:515–529. https://doi.org/10.1093/bib/bbw082
Hynds RE, Frese KK, Pearce DR et al (2021) Progress towards non-small-cell lung cancer models that represent clinical evolutionary trajectories. Open Biol 11:200247. https://doi.org/10.1098/rsob.200247
Stransky N, Ghandi M, Kryukov GV et al (2015) Pharmacogenomic agreement between two cancer cell line data sets. Nature 528:84–87. https://doi.org/10.1038/nature15736
Goldman M, Craft B, Brooks A et al (2019) The UCSC Xena platform for public and private cancer genomics data visualization and interpretation. bioRxiv. https://doi.org/10.1101/326470
Ding Z, Zu S, Gu J (2016) Evaluating the molecule-based prediction of clinical drug responses in cancer. Bioinformatics 32:2891–2895. https://doi.org/10.1093/bioinformatics/btw344
Skidmore ZL, Wagner AH, Lesurf R et al (2016) GenVisR: genomic visualizations in R. Bioinformatics 32:3012–3014. https://doi.org/10.1093/bioinformatics/btw325
Akbani R, Akdemir KC, Aksoy BA et al (2015) Genomic classification of cutaneous melanoma. Cell 161:1681–1696. https://doi.org/10.1016/j.cell.2015.05.044
Nazari Z, Kang D, Asharif MR et al (2015) A new hierarchical clustering algorithm. In: 2015 Int Conf Intell Inform Biomed Sci (ICIIBMS). https://doi.org/10.1109/ICIIBMS.2015.7439517
P S, Gupta S, (2011) A comparative study on distance measuring approaches for clustering. Int J Res Comput Sci 2:29–31. https://doi.org/10.7815/ijorcs.21.2011.011
Ward JH (1963) Hierarchical grouping to optimize an objective function. J Am Stat Assoc 58:236–244. https://doi.org/10.1080/01621459.1963.10500845
Xu Q, Zhang Q, Liu J, Luo B (2020) Efficient synthetical clustering validity indexes for hierarchical clustering. Expert Syst Appl 151:113367. https://doi.org/10.1016/j.eswa.2020.113367
Nana FA, Lecocq M, Ladjemi MZ et al (2019) Therapeutic potential of focal adhesion kinase inhibition in small cell lung cancer. Mol Cancer Ther 18:17–27. https://doi.org/10.1158/1535-7163.MCT-18-0328
Hecker L (2018) Mechanisms and consequences of oxidative stress in lung disease: therapeutic implications for an aging populace. Am J Physiol-Lung Cell Mol Physiol 314:1642–1653. https://doi.org/10.1152/ajplung.00275.2017
Inamura K (2017) Lung cancer: understanding its molecular pathology and the 2015 WHO classification. Front Oncol 7:193. https://doi.org/10.3389/fonc.2017.00193
Funding
This work was supported by the grants from the National Natural Science Foundation of China (62102004), the Natural Science Young Foundation of Anhui (2008085QF293), the Natural Science Young Foundation of Anhui Agricultural University (2019zd12), the Introduction and Stabilization of Talent Project of Anhui Agricultural University (yj2019-32), and the Graduate Innovation Fund of Anhui Agricultural University (2021yjs-53).
Author information
Authors and Affiliations
Contributions
YS: conceptualization, data curation, methodology, visualization, and writing-original draft. YX: data curation, software, and methodology. XH: data curation and visualization. YZ: methodology and writing-review and editing. ZY: conceptualization, supervision, writing-review and editing, and funding acquisition. All the authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Shen, Y., Xiang, Y., Huang, X. et al. Pharmacogenomic Cluster Analysis of Lung Cancer Cell Lines Provides Insights into Preclinical Model Selection in NSCLC. Interdiscip Sci Comput Life Sci 14, 712–721 (2022). https://doi.org/10.1007/s12539-022-00517-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12539-022-00517-z