Abstract
In December 2019, Wuhan city in the Hubei province of China reported for the first time a cluster of patients infected with a novel coronavirus, since then there has been an outburst of this disease across the globe affecting millions of human inhabitants. Severe acute respiratory syndrome coronavirus type-2 (SARS-CoV-2), is a member of beta coronavirus family which upon exposure caused a highly infectious disease called novel coronavirus disease-2019 (COVID-19). COVID-19, a probably bat originated disease was declared by World Health Organization (WHO) as a global pandemic in March 2020. Since then, despite rigorous global containment and quarantine efforts, the disease has affected nearly 56,261,952 laboratory confirmed human population and caused deaths of over 1,349,506 lives worldwide. Virus passes in majority through respiratory droplets and then enters lung epithelial cells by binding to angiotensin converting enzyme 2 (ACE2) receptor and there it undergoes replication and targeting host cells causing severe pathogenesis. Majority of human population exposed to SARS-CoV-2 having fully functional immune system undergo asymptomatic infection while 5–10% are symptomatic and only 1–2% are critically affected and requires ventilation support. Older people or people with co-morbidities are severely affected by COVID-19. These categories of patients also display cytokine storm due to dysfunctional immune response which brutally destroys the affected organs and may lead to death in some. Real time PCR is still considered as standard method of diagnosis along with other serology, radiological and biochemical investigations. Till date, no specific validated medication is available for the treatment of COVID-19 patients. Thus, this review provides detailed knowledge about the different landscapes of disease incidence, etiopathogenesis, involvement of various organs, diagnostic criteria’s and treatment guidelines followed for management of COVID-19 infection since its inception. In conclusion, extensive research to recognize novel pathways and their cross talk to combat this virus in precarious settings is our future positive hope.
Similar content being viewed by others
Introduction
An outbreak of highly infectious novel coronavirus disease-2019 (COVID-19) emerged in December 2019 in Wuhan city in the Hubei province of China which rapidly spread across worldwide and was later declared pandemic by WHO on March 11, 2020 [1]. The disease was caused by a novel coronavirus, severe acute respiratory syndrome coronavirus type-2 (SARS-CoV-2) classified on the same day by International Committee on Taxonomy of Viruses [2]. Since then, disease is continuously spreading and as per the WHO report till 20th November, it has nearly targeted 56,261,952 infected cases and cost over 1,349,506 human lives [3]. In India, first case of COVID-19 was reported on January 30, 2020, which originated from China. According to Indian Council of Medical Research (ICMR) and Ministry of Health and Family Welfare (MOHFW), total 9,004,365 confirmed cases were reported, whereas 132,162 deaths occurred till November 20, 2020 [4, 5].
Shanghai Public Health Clinical Center and School of Public Health researchers on January 7, 2020 jointly revealed the origin of this coronavirus from a seafood market in Wuhan city [6]. Number of cases started increasing since then, even in people not having any exposure to animal market, and hence human-to-human transmission was also reported to occur [7]. Coronavirus (CoVs) encompass a large virus family affecting mainly humans and different animal species including cats, cattles, camels and also bats [8]. Not so often, but animal coronaviruses have also infected human, recently introduced SARS-CoV2 has joined the list of middle east respiratory syndrome coronavirus (MERS-CoV) and severe acute respiratory syndrome coronavirus (SARS-CoV). SARS-CoV2, a beta coronavirus, has probably originated from bats just like MERS-CoV and SARS-CoV [9]. Human coronaviruses in majority causes respiratory and enteric infections and SARS-CoV-2 infection causes flu like symptoms ranging from fever, cough, headache, asthenia etc. Once the virus reaches to respiratory epithelial cells, it enters by binding to angiotensin converting enzyme-2 through its spike proteins and is then endocytosed inside the host cells. Once inside the cells, the virus starts replicating and targets variety of host cells thereby causing severe pathologies. However, SARS-CoV-2 infection affects people of all ages, in some high-risk individuals, such as older people or those having co-morbidities, the virus might cause severe infections such as interstitial pneumonia, acute respiratory distress syndrome (ARDS), progressing to multi organ failure and ultimately causing respiratory failure leading to death [10]. Cytokine storm is one of the characteristics of COVID-19 infection majorly occurring in persons having dysfunctional immune responses leading to secretion of pro-inflammatory cytokines such IL-16, IL-1β, TNF-α, IL-2, IL-7, MCP1, G-CSF, IP-10, IL-10, MIP1α and others. This review, focusses on current incidence, etiopathogenesis, clinical characteristics with multiorgan involvements associated infections, diagnostic criteria’s, its management and present and future prospects of clinical trials to prevent, manage and control COVID-19.
Structural Aspects of Coronavirus
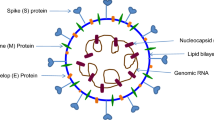
Coronavirus (CoVs) are the largest group of viruses belongs to the Nidovirales order that comprises three families i.e. arteriviridae, roniviridae and coronaviridae [11]. On the basis of genome structure, coronaviridae comprises of four genera- alpha coronavirus, beta coronavirus, gamma coronavirus and delta coronavirus. Alpha coronavirus and beta coronavirus transmission are limited to mammals and in humans they cause respiratory illnesses such as SARS and MERS, whereas gamma coronaviruses and delta coronaviruses infect birds as well as mammals [12, 13]. Coronaviruses are 60 to 140 nm in diameter and have 26–32 kilobases positive sense single stranded RNA genome connected to a nucleoprotein surrounded by capsid [14]. A typical CoV has atleast six open reading frames (ORF) and all proteins including structural and accessory are encoded by a single guide RNA [15]. Two major overlapping open reading frames (ORFs) i.e. ORF1a and ORF1b comprise of two-third of the whole genome and the third ORF codes for four structural proteins i.e. spike (S), envelope (E), membrane (M), and nucleocapsid (N). SARS-CoV-2 genome encodes, a very large polyproteins pp1ab, four structural proteins and six accessory proteins (3a, 6, 7a, 7b, 8, and 10) which is equivalent to the structure of SARS-CoV and MERS-CoV [16, 17]. (Figs. 1 and 2).
The Spike (S) proteins has been divided into functional subunits i.e. N-terminal S1 subunit and a C-terminal S2 subunit. The size of each monomer of S protein is 180 kDa and is involved in attachment and membrane fusion. Depending on the virus, both N and C terminal domain are involved in the membrane attachment and entry of viral genome inside the host cells. Apart from this, an additional furin like cleavage site is present at the junction of S1 and S2 subunits of S protein in SARS-CoV-2, which is absent in the genome of SARS-CoV. This additional subunit has been speculated to be involved in higher infectivity of SARS-CoV-2 than SARS-COV [18]. Further, using phylogenetic tree investigation that SARS-CoV-2 genome shares 79% genetic similarity with SARS-CoV [19] whereas it demonstrates 98% genetic similarities with bat coronavirus RaTG13 [20] and is 85.98% identical to pangolin-CoV genome [21] thereby suggesting its zoonotic origin.
Further, whole genome sequencing analyses have revealed that mutations occurring in the spike protein are important in the evolution of SARS-CoV-2. Phylogenetic and alignment study of 591 novel coronaviruses of different clades from Group I to Group V identified several mutations and related amino acid changes and majority were distributed over spike protein [22]. In a major study involving investigation of 3617 available whole genome sequence of SARS-COV-2 maintained in NCBI globally identified that E protein possess several non-synonymous mutations [23]. Additionally, from five different isolates identified in the state of West Bengal (Eastern state of India), India, two major mutations one at position 723 and other at 1124 were identified in the S2 domain of Spike (S) protein. Another mutation at 614 position in S1 domain of S protein was also identified which is similar to mutation found in the isolates of state of Gujarat (Western state of India). These mutations were crucial in modulating the affinity of receptor binding [24]. Thus, these whole genome study revealed that SARS-CoV-2 is continuously evolving virus with crucial mutation identified in the receptor binding regions.
SARS CoV-2 Transmission and Its Etiopathogenesis
Virus spreads mainly from person to person through respiratory droplets or bits of liquid, mostly through sneezing or coughing [25,26,27,28]. According to National Institutes of Health (NIH), SARS-CoV-2 virus is stable in aerosols and on surfaces for several hours to days depending upon the surface materials and also ambient environmental conditions. Virus has been reported to be detectable up to three hours in aerosols, up to four hours on copper, up to 24 h on cardboard and up to two to three days on plastic and stainless steel [29]. It has also been testified in stool and may contaminate water supply possibly and subsequently results in aerosol and/or feco-oral based route of transmission [30]. Another study reported the presence of SARS-CoV-2 virus in the tears and conjunctival secretions [31].
Virus can pass through nasal and larynx mucous membranes and enter into the lungs through respiratory tract. First step of any viral infection is the binding of some viral proteins with receptors expressed by host cells followed by fusion with host cell membrane. ACE2 has been speculated to be the ultimate target of spike protein of SARS-CoV-2. Initial target of virus include lung epithelial cells, where virus attach through its spikes to cellular angiotensin converting enzymes 2 (ACE2) receptor [32].
ACE2 is a membrane bound metalloproteinase and was discovered in 2000. It is a homolog of ACE, though its activity cannot be stopped by angiotensin-converting enzyme inhibitors (ACEIs) [33]. Further study reported that SARS-CoV-2 has nearly 10–20 folds higher affinity for ACE2 than SARS-CoV. ACE2 is also significantly expressed in other tissues like heart, renal, and gastrointestinal tract [34, 35]. It is an important regulator of renin-angiotensin system (RAS) and renin angiotensin aldosterone system (RAAS). RAS is an indispensable system of human body which is regulated and counter-regulated by two main pathways: classical RAS axis ACE-angiotensin II- angiotensin I receptor pathway, and counter-regulatory RAS axis ACE2- angiotensin 1–7- MasR-based pathway, which plays a negative role in regulation [36, 37]. In classical RAS pathway, angiotensinogen is cleaved into angiotensin (Ang) by renin, which is then cleaved by ACE into angiotensin II. This Ang II after binding to type 1 angiotensin II receptor (AT1R) mediate functions like increases secretion of aldosterone and anti-diuretic hormone, increases blood pressure by decreasing sensitivity to baroreflex, increases vasoconstriction, decreases production of nitric oxide (NO), increases fibrosis, reactive oxygen species production and inflammation. Whereas ang II can also be cleaved by ACE2 in counter-regulated pathway into Ang-(1–9) and Ang-(1–7), Ang (1–9) which then binds to AT2R and triggers NO production thus mediating vasodilation and reducing blood pressure. In addition to that, it also reduces inflammation, fibrosis and cardiac hypertrophy. Further, Ang-(1–7) binds to proto- oncogene Mas receptor (MasR) and reduces the blood pressure and reverses all its function performed during classical RAS pathway.
SARS-CoV-2 attaches to the receptor of host cell through S1 domains of its spike glycoprotein (S), and this spike protein is proteolytically cleaved by transmembrane serine protease 2 (TMPRSS2) into S1 and S2 subunits and then S2 induces membrane fusion and virus internalization by endocytosis [38]. S1 regulates virus-host range and cellular tropism with the receptor binding domain (RBD), where S2 helps in virus cell membrane fusion with the help of two tandem domains, heptad repeats 1 and 2 (HR1 & HR2) [39, 40]. After the fusion it releases viral genome RNA into cytoplasm via endocytosis and replication and transcription of virus occur in cytoplasm. Positive RNA genome acts as a messenger RNA (mRNA). Once viral genome enters into host cell, it translates into pp1a and pp1ab, two large precursor polyproteins [41] and these polyproteins by ORF 1a-encoded viral proteinases, 3C-like proteinases (3CLpro) and papain-like proteinase (PLpro) get processed into 16 mature nonstructural proteins (nsp1–nsp16) [9, 42]. During viral RNA replication and transcription these nonstructural proteins (nsps) perform vital functions [43] (Fig. 3). This viral replication and cell to cell transmission plays important role in the suppression of ACE2 expression. This suppression of ACE2 leads to decrease in Ang (1–7) synthesis and enhanced levels of Ang II which drives the Ang II–AT1R dependent inflammatory pathways in the lungs and causes parenchymal injury [44]. SARS-CoV-2 enhanced the apoptosis and p53 signaling pathways in lymphocytes thereby causing the lymphopenic situation in COVID-19 patients [45]. Additionally, coagulopathy has also been observed in COVID-19 infection, as elevated levels of plasmin(ogen) has been observed in patients. This plasmin along with other proteases might play an important role in the cleavage of furin site in the S protein of SARS-CoV-2, thus increasing its virulence and infectivity and is also associated with hyperfibrinolysis [46]. Among the existing coronaviruses, RNA recombination along with its potential proof-reading capacity has been involved in the development and occurrence of novel coronaviruses [47]. Organs presenting higher ACE2 expression are presented as possible targets of SARS-CoV-2 virus infection such as lung being an important and primary infection target organ. Further, ACE2 expression and its organ wise distribution is significantly associated with COVID-19 clinical symptoms [25, 48]. (Fig. 4).
Symptoms And Etiological Factors Associated With Pathogenesis
Nearly, 80% of the SARS-CoV-2 infected patients display mild symptoms, whereas 15% cases come in severe category and rest 5% fall under critical category requiring ventilation support [49]. The course of infection of SARS-CoV-2 spans from mild disease limiting itself to upper respiratory, non-severe pneumonia, could be severe pneumonia involving ARDS, multiple organ failure and ultimately death [50]. Common clinical symptoms include fever, dry cough, headache, sore throat, breathlessness, diarrhea, vomiting and abdominal pain [51, 52]. Olfactory and taste desensitization have also been reported in COVID-19 [53].
People who have some chronic medical comorbidities such as cardiovascular disease, any lung problems or diabetes or who are older and have higher risk of severe illness (charted out in Tables 1 and 2). Children who are infected with SARS-CoV-2 generally display mild symptoms which might be due to altered ACE2 activity and active innate immune response. Whereas adult patients display suppressed adaptive immunity and dysfunctional immune response. Further, it is also observed that women are less vulnerable to viral infection compared to men which may be due to incongruent immune system, steroid hormone or sex chromosome associated factors. In the next few paragraphs, we will observe that how infection with SARS-CoV-2 affects different organs of the body and also on the dysfunctional immune response.
Immune Response With Cytokine Storm in COVID-19
Immune system of an individual is highly responsible to depict asymptomatic or symptomatic clinico manifestations of COVID-19. Geriatric population with poor immune function along with comorbidities is at increased risk with more susceptibility to various viral and bacterial infections due to less efficient, less coordinated and slower immune response. Cytokine storm noted in critical cases of COVID-19 is a response of an uncontrolled immune mechanism with an uncontrolled release of cytokines. Multiple factors are involved in triggering cytokine storm which includes virus, bacterial components, sepsis, super antigens, toxins, chimeric antigen receptor T cells and others [60]. It is a life threatening condition leading to detrimental changes including capillaries leakage, edema, tissue toxicity, organ failure and even shock. Increased levels of IL-6 were significantly observed with clinical manifestation in critical COVID-19 cases [61]. A trend has been observed where, SARS-CoV-2 infection rapidly activates CD4 + T lymphocytes thereby forming pathogenic T helper (Th1) cells and release various cytokines including GM-CSF. This cytokine environment, induces an inflammatory environment with production of CD14 + , CD16 + monocytes along with enhanced secretion of IL-6. Finally, pathogenic T-cells along with monocytes enter pulmonary circulation where monocytes induce to become macrophages [62]. Acute respiratory distress syndrome (ARDS) progression in SARS was generally manifested along with increased circulatory levels of pro-inflammatory cytokines including interleukin-1β (IL-1β), IL-6, CXC-chemokine ligand 2 (CCL2) and CXCL10. COVID-19 cases have been associated with increased plasma levels of pro-inflammatory mediators, including IL1-β, IL1RA, IL7, IL8, basic FGF2, IFNγ, MCP1, MIP1α, MIP1β, TNFα, and others. Studies have documented levels of IL2, IL7, IL10, GCSF, IP10, MCP1, MIP1α, and TNFα were associated with disease severity in COVID-19 [32, 63]. (Fig. 5).
Lung Injury in COVID-19
Coronavirus infection frequently occurs in patients with pre-existing lung disease such as asthma and COPD [64, 65]. Lung tissue has high renin-angiotensin system (RAS) activity which is chief site of Ang II synthesis. Ang II is an effective pulmonary vasoconstrictor [66]. Ang II over activity also promotes pulmonary edema and impairs lung function [67]. Acute respiratory distress syndrome (ARDS) is the most severe form of acute lung injury. ACE2 expression is very high in lung tissue and is associated with protection of lung tissue injury induced by sepsis [67]. Decreased expression of ACE2 is inversely correlated with angiotensin II levels thereby enhanced its production. Further, this angiotensin II stimulates its type 1a receptor, which then enhances lung vascular permeability and leads to its pathological phenotype [14].
Single-cell RNA sequencing data of 43,134 human lung cells reported that only 0.64% of total cells expressed ACE2 and out of which 83% of total ACE2 expression was present on type II apical surface of epithelial cells (AT2) [41]. Other cellular types including type I epithelial cells, airway epithelial cells, endothelial and others also expresses ACE2 but at levels considerably lower than AT2 cells [68]. Thus, these results indicated AT2 cells as primary targets of SARS-CoV-2 in lungs. Hypoxia is one of the clinical symptoms associated with various pulmonary diseases. During initial phase of hypoxia, enhanced expression of ACE2 was determine in pulmonary smooth muscle cells whereas during later phase. Hypoxia-Inducible Factor-1α (HIF-1α), induces ACE expression (one of the targets of HIF-1α) with concomitant decreased in the expression of ACE2 [69].
Kidney Injury in COVID-19
Kidneys are presented as another major targets of SARS-CoV-2 viral infection as virus was detected in the urine samples of few COVID-19 positive cases [25]. In these patients, acute kidney injury is presented as a critical complication targeting renal intrinsic cells including proximal tubular epithelial cell thus leading to kidney dysfunction. Kidney expresses very high levels of ACE2, which is very high especially in the renal cortex [70]. During acute kidney disease [71] and other models of chronic kidney diseases, ACE2 expression levels are severely compromised which then deregulated the homeostasis of RAS and causes debilitating pathological changes in the kidneys [72].
Gastrointestinal Tract (GIT) in COVID-19
Some COVID-19 patients have depicted gastrointestinal symptoms like diarrhea, vomiting and abdominal pain. Previously reported studies suggested relation between coronavirus and gastrointestinal tract tropism [60]. In coronavirus infection, tropism of gastrointestinal may explain an incidence of diarrhea. SARS-CoV RNA might be detected in SARS patients stool specimens [73]. SARS-CoV-2 detected in fecal samples was most likely due to virus entering the blood from lungs and then traveling from blood to intestines [74]. This infected fecal sample can promote fomites-based transmission, especially because of generation of infectious aerosol from toilet plume. Literature has reported very high expression of ACE2 receptor in GIT, particularly in small and large intestines [75, 76]. Positive staining of viral nucleocapsid was detected in cytoplasm of gastric, duodenal and rectal epithelium cells [77]. These evidences provided indispensable information about mode of entry of virus into host cells and further unveiled the possible route of transmission. Some studies reported viral RNA presence in the stool of COVID-19 patients [78,79,80]. In another study of 73 COVID-19 patients, 39 (53.4%) were positive for SARS-CoV-2 RNA in stool, where duration of viral positivity ranged from 1 to 12 days. Even more, 17 (23.3%) patients that were negative for respiratory samples showed positive results with viral RNA in stool [75].
Liver Injury in COVID-19
Patients with COVID-19 have been reported to have high levels of serum Alanine Transaminase (ALT) and Aspartate Transaminase (AST) [48, 77]. In a study, 56 COVID-19 patients, 54% patients have high levels of serum Gamma-Glutamyl Transferase (GGT) [81]. Chai et al. stated that ACE2 expressed in both liver cells and bile duct cells [82]. Nevertheless, expression of ACE2 in bile duct cells was higher as compared to liver cells. Epithelial cells of bile duct play important roles in liver regeneration and immune response [83]. According to these results in COVID-19, patient’s liver injury occurred may be due to damage of bile duct cells, but not liver cells by virus infection.
Cardiovascular Diseases Associated with COVID-19
Respiratory failure is the main cause of death from COVID-19 patients, mostly in older adults and those who have weaker immune system [84, 85]. Most common cardiovascular (CV) complication is acute myocardial injury. SARS-CoV-2 enters human cells through ACE2 receptor. In normal healthy and in various disease conditions, ACE2 helps in neurohormonal regulation of CV system. When SARS-CoV-2 binds with ACE2R it alters the ACE2 signalling pathways that can lead to acute myocardial and lung injuries [86]. Roughly, 8–12% cases of acute myocardial injury reported significant elevation of cardiac troponin I (cTnI) [86]. In a meta-analysis from China [66, 87], incidences of acute cardiac injury were reported to be 8%; however, in another study, they included only those patients who were discharged from hospital or dead and found 17% incidence of cTnI elevation [88]. Clinical investigations suggested that patients with heart diseases, hypertension or diabetes who were treated with ACE2 increasing drugs including inhibitors and blockers showed increased expression of ACE2 and these were at higher risk of getting SARS-CoV-2 infection [89] (Table 3).
Diabetes Mellitus Associated with COVID-19
ACE2 modifies neutral amino acid transporters expression on epithelial cells surface and growth of pancreatic islet cells along with insulin secretion by pancreatic β-cells [70]. A study involving a mice model of adenovirus mediated human ACE2 expression showed enhanced production of insulin along with decreased apoptosis of pancreatic islets [96]. In addition to that, ACE2 also improves the overall endothelial function of pancreatic islet microvascular unit [97]. Some studies stated that diabetes mellitus (DM) was associated with more severe disease, acute respiratory distress syndrome and increased mortality [25, 98]. In a study by Guan et al., patients with DM showed higher disease severity (16.2%) in contrast, patients without DM showed lesser disease severity (5.7%) [25]. Further, an obvious conclusion was made, where patients having DM higher in age was compared to non-Diabetes mellitus patients. Thus, as age advances it emerged as crucial prognostic factor in determining course of COVID-19 disease. Simultaneously, patients with DM showed lowered ACE2 expression possibly due to glycosylation, which explained increased comorbidity with severe lung injury and ARDS in COVID-19 patients [99, 100]. COVID-19 patients with diabetes as coexisting disorders had a worse clinical outcome.
Diagnosis of COVID-19 Infection
Currently, multiple approaches are efficiently being used for the diagnosis of COVID-19 infection [101]. Two broadly classified techniques include, Real time reverse transcriptase polymerase chain reaction (rRT-PCR) based detection and serology-based detection. RT-PCR technique involves conversion of viral RNA genome into DNA, which is further amplified using specific primers set against defined targeted regions of viral genome. Major targeted regions of SARS-CoV-2 genome include nucleocapsid (N) gene, envelope (E) gene and ORF1ab gene regions [102]. RT-PCR based method displays high sensitivity (85–90%) and high specificity for the COVID-19 diagnosis as it is dedicated to the direct amplification of viral genetic material and the turnaround time (TAT) is about 2.5–3.5 h. This technique is quantitative in nature. Sample types for RT-PCR include nasopharyngeal swabs and oropharyngeal swabs. The RT-PCR results generally become positive after 2–8 days of infection and it is able to process large batches of samples. However, there are some limitations of this technique apart from skilled manpower, mutation in the viral genome might give false negative results, and further possible limitation includes improper collection, transportation and handling contributing to false negative results [103]. Rapid PCR by cartridge system (CBNAAT) are also available which are equally efficient and has significantly short response time [104]. Additionally, loop mediated isothermal amplification (LAMP) is another version of CBNAAT, which is highly specific isothermal amplification technique. In this method, transcription of RNA to DNA is then followed by amplification done using 4–6 primers set against multiple targeted regions [105]. This method has got several advantages over RT-PCR in being specific, user friendly in detection, speed (turnaround time is 20 min) and less background. Recently, Sree Chitra Thirunal Medical Institute and Technology in collaboration with Agappe Diagnostics Ltd. developed Chitra Gene LAMP-N that detects two different regions of N gene. In addition to above, CDC has approved a one-step rRT-PCR test kit to quantitatively determine viral molecules in > 90 samples within 45 min [106]. Additionally, Specific High Sensitivity Enzymatic Reporter Unlocking (SHERLOCK) employs the CRISPR-Cas system for the detection of viral RNA genome with greater specificity [107]. Similar, technique was employed by Indian scientists who have developed FELUDA, a test kit for SARS-CoV-2 genome detection also employs CRISPR-Cas9 system [108].
Another, crucial diagnostic method is serology-based detection of SARS-CoV-2 infection among COVID-19 patients. This method employs detection of antigens/antibodies in the human blood. COVID-19 infection leads to the generation of IgM antibodies and then IgG antibodies inside the host body. Initially after infection, the titer of IgM increases within a week time. Later on, titer of IgG increases from day 4 post infection and reaches to its peak by day 14. Further, the levels of IgM degrade very rapidly whereas IgG antibodies titer persist in the body for longer duration thus helps in determination of active infection. Apart from antibodies detection, antigen detection (S and N protein) is also being explored for early detection of viral infection. Three major strategies are being employed for detection of Ags/Abs including lateral flow immunoassay, ELISA, and chemilumescence.
Further apart from above two standardized techniques for detection of COVID-19 infection, other parameters such as high sensitivity CRP levels, lactate dehydrogenase, alanine transaminase, erythrocyte sedimentation rate have been observed [109]. Further, decreased lymphocyte count i.e. depletion of CD4 + and CD8 + cells and decreased IFN-γ expression in CD4 + T cells are linked with severe COVID-19, illustrating the cytokine storm. Another crucial observation is the presence of intravascular coagulation associated with increased D-dimer and fibrinogen levels in some COVID-19 infection. Lastly, chest X-ray (CXR) and computed tomography (CT) scan are considered as an essential tool in the detection of COVID-19 pneumonia during this pandemic.
Treatment Strategies for COVID-19 Infection
ACE2 for Therapeutics
Despite significant efforts made towards development of vaccines and therapeutic drugs, no significant advances have been observed till today. Currently available drugs are generally categorized according to their targets, one acts on mode of entry of virus and other acts on inhibition of enzymes responsible for viral genome replication. Combination of lisinopril and losartan treatment in normotensive Lewis rats abolished increase in ACE2 mRNA levels observed individually but retained losartan induced rise in ACE2 activity in heart [110]. Ang II can regulate ACE2 expression through AT1R. Healthy hearts and kidneys displayed high levels of ACE2 mRNA and protein expression, with moderate expression of ACE [82]. RAS over activation in CVD increases AT1R stimulation by Ang II, promoting ERK1/2 and p38 MAPK signalling pathways to downregulate ACE2 while upregulating ACE expression [111]. Promoting ACE2/Ang 1–7/Mas signalling by rhACE2 or Ang 1–7 receptors agonist AVE 0991 can have valuable therapeutic effects in CVD and lung disease from diverse aetiologies [112]. Ang 1–7 receptors agonist AVE 0991 has been shown to exert cardiorenal and pulmonary protective effects, [113] and treatment with rhACE2 improved symptoms of acute lung injury, CVD, and kidney injury in various preclinical models [67, 70, 114]. Maintaining ACE2 levels in patients with or predisposed to common CVD states such as diabetes, hypertension, and obesity wards off the advancement of these comorbidities in instances where, patient contracts SARS-CoV-2 by maintaining levels of ACE2/Ang1–7/MasR negative counter-regulation.
Vitamin D Response in COVID-19
Vitamin D (calcitriol or vitamin D3) plays an important role in regulation of serum calcium concentration. Although less highlighted, several reviews have addressed role of vitamin D in prevention of viral infections [115, 116]. Multiple mechanisms are employed by vitamin D in reducing risk of microbial infection and diseases. In our previous papers, we have shown that various polymorphic variants of vitamin D receptor gene modulate the metabolism of blood lead levels along with circulatory levels of vitamin D [117, 118]. Such as in reducing risk of common cold, vitamin D employs three different categories of mechanisms: physical barrier, cellular natural immunity, and adaptive immunity [119, 120]. Some studies reported that administration of an oral dose of 50,000 IU of vitamin D reduces risks of influenza. Vitamin D adequacy also reduces severity of pneumonia, which is associated with coronavirus infection. Taking higher doses of vitamin D supplement or sun exposure is reported to boost immunity and reduce risk. Vitamin D helps in maintenance of various cellular junctions including tight junction, gap junction and adherence junction which are otherwise evaded by several viruses to get inside cells and thereby causing infections [121]. In response to the bacterial and viral infections, innate immune system produces both pro and anti-inflammatory cytokines. Vitamin D improves cellular immunity by minimizing these abrupt increases in cytokines produced mainly by innate immunity [7]. Vitamin D supplementation also stimulates antioxidant genes expression including glutathione reductase and glutamate–cysteine ligase modifier subunit [122]. This increased secretion of glutathione further spares ascorbic acid (vitamin C) which got significant antimicrobial properties [123] and is suggested to be used in prevention and treatment of COVID-19 [124]. SARS-CoV-2 attached to host cells through ACE2R and vitamin D might reduce ACE2R. Recent studies have established that RAS is a major target of vitamin D. Inverse relationship between circulating 25(OH)D levels (a physiological quantifiable form) and plasma renin activity in hypertensive subjects was reported more than two decades ago; however, significance of the same was recognized only after discovery that 1,25(OH)2 D3 is a negative endocrine regulator of renin production [125].
Other drugs and their targets
Further, a broad range of antiviral drugs have also been used in various clinical trials against COVID-19 [126]. RNA dependant RNA polymerase, a crucial protease required during viral RNA replication is also an important therapeutic target. Remdesivir, a broad-spectrum antiviral drug initially used for Ebola virus treatment is an analogue adenosine. It effectively blocks viral infection and replication in vitro and in animals and has shown some promising effect against SARS-CoV-2 [127]. Favipiravir, purine nucleic acid analogue initially used for the treatment of influenza is an effective RNA dependant RNA polymerase inhibitor [128]. In addition to above, combination of lopinavir and ritonavir has also been tested for the treatment of COVID-19 infected patients with little clinical significance [129].
Another potential drug is chloroquine, an anti-malarial and anti-parasitic drug, is a promising anti-viral known for neutralising the acidic endosomal pH and thereby blocking the endosomal mediated viral entry [130]. Chloroquine demonstrates potent anti-viral effect by inhibiting SARS-CoV-2 in-vitro with a 90% effective concentration of 6.9 µM. Hydroxychloroquine, an analogue of chloroquine is a less toxic and more potent drug. Chloroquine exhibits potent anti-inflammatory and immuno modulatory effects which might play an important role in reduction of COVID-19 infection probably by hindering cytokine storm [131]. Combination of chloroquine with azithromycin has provided some clinical benefits in COVID-19 infection [132].
Convalescent plasma therapy has long been given to patients infected with deadly viruses like SARS-CoV, H1N1, Spanish flu, Ebola and the MERS. The techniques involved identification of patients who have already infected with COVID-19 and have completely recovered 14 days prior to be successful donors. Further, they must have sufficient titer of neutralizing antibodies [133]. Various clinical trials have highlighted that the therapy significantly improved the patient’s clinical outcomes and were able to successfully clear the infection and infected cells by activating complement system and phagocytosis [134, 135].
Cytokine storm is one of the characteristic phenomena of SARS-CoV-2 infected patients. Initial evidences revealed that targeting anti-inflammatory molecules such as IL-6, IL-1R and TNF-α might be important in reducing the inflammation and ultimately inflammatory response. Currently, clinical trials have shown that tocilizumab (anit-IL-6), a humanized antibody significantly improves the clinical outcome of COVID-19 cases [4]. Similarly, Anakinra (anti-IL-1R) is also important in decreasing the inflammation [136].
Corticosteroids are well known anti-inflammatory molecules are effective for the treatment of a variety of inflammatory diseases [137]. They behaved like a double edged sword as on one hand they reduce inflammation while on the other hand they inhibit immune response thereby delaying the infection clearance [137]. In COVID-19, various clinical trials have outlined the role of corticosteroids such as one retrospective observational study revealed that 72% of ICU patients with COVID-19 are receiving glucocorticoids treatment [48]. Further, another study revealed that treatment of COVID-19 patients having ARDS, with steroids significantly decreased the mortality rate compared to those that did not receive steroids (46% vs 68%) [100].
Mesenchymal stem cells (MSCs) are known to play important roles in immune modulation either by secreting cytokines or directly interacting with target cells. MSCs are available at multiple sources including blood, bone, adipose tissue, placenta, umbilical cord etc. [138]. A clinical trial in COVID-19 patients (n=17) revealed that the treatment cleared inflammatory cytokines secreting T cells population whereas dendritic cell percentage and IL-10 levels were increased. Further, absence of ACE2 and TMPRSS2 in the MSCs further highlights their importance in the COVID-19 treatment [139].
Recently, studies are focussing on curbing the complement system pathway, a component of innate immune system, in reducing hyperinflammatory and hypercoagulation stage in severe COVID-19 patients [140]. Furthermore, Ibrahim et al. (2020) reported that intravenous injection of N-acetyl cysteine in small cases of COVID-19, demonstrated significantly decreased inflammation and clinical improvement along with markedly reduced CRP levels in all patients and ferritin in 9/10 patients [141]. Lastly, other options such as Imatinib a tyrosine kinase inhibitor and colchicine, an anti-inflammatory drug have also been explored in few COVID-19 cases [142, 143].
Present and Future Prospects of Ongoing Clinical trial
Currently, various clinical trials are ongoing to test specificity and efficacy of vaccines and antibodies precisely targeting SARS-CoV-2. Multiple pharmaceutical drugs that have kept therapeutic promises in the treatment of COVID-19 includes human immunoglobulin, chloroquine, hydroxychloroquine, remdesivir, favipiravir, ritonavir, lopinavir, arbidol etc. The role of these therapeutic drugs shown in Table 4.
Conclusion
Current review addressed relevant information of SARS-CoV-2, incidence, etiopathogenesis, clinical characteristics with multiorgan involvements in associated comorbid diseases, present and future prospects of clinical trials to prevent, manage and control COVID-19 pandemic infection. Information depicted in present review may serve to fill existing lacunae in knowledge on pathogenesis of COVID-19. Moreover, aspects of SARS-COV-2 etio-clinio-pathogenesis and replication depicted inside infected cells might help in development of present and future prospects of ongoing clinical trials. New updates poured reticent on deadly virus from all over world are considered in present review to enhance knowledge and clear out concepts for present future prospects of clinico-trials to ascertain success in the possible drugs development for COVID-19 treatment. Extensive research to recognize novel pathways and their cross talk to combat this virus in precarious settings is our future positive hope.
References
Wang C, Horby PW, Hayden FG, Gao GF. A novel coronavirus outbreak of global health concern. The Lancet. 2020;395:470–3. https://doi.org/10.1016/S0140-6736(20)30185-9.
Jarvis JLM, Drug firms mobilize to combat coronavirus outbreak, C EN. (2020) 98, 5.
World Health Organization. Coronavirus disease (COVID-19) situation reports 114., (2020). https://www.who.int/publications-detail/infection-prevention-and-control-during-health-care-when-novel-coronavirus-(ncov)- infection-is-suspected-20200125.
M. of H. and F.W. www.mohfw.gov.in. Retrieved 14, May 2020, n.d. www.mohfw.gov.in. Retrieved 21 April 2020.
Mitra P, Misra S, Sharma P. COVID-19 pandemic in India: what lies ahead. Indian J Clin Biochem. 2020;35:257–9. https://doi.org/10.1007/s12291-020-00886-6.
Lu R, Zhao X, Li J, Niu P, Yang B, Wu H, Wang W, Song H, Huang B, Zhu N, Bi Y, Ma X, Zhan F, Wang L, Hu T, Zhou H, Hu Z, Zhou W, Zhao L, Chen J, Meng Y, Wang J, Lin Y, Yuan J, Xie Z, Ma J, Liu WJ, Wang D, Xu W, Holmes EC, Gao GF, Wu G, Chen W, Shi W, Tan W. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. The Lancet. 2020;395:565–74. https://doi.org/10.1016/S0140-6736(20)30251-8.
Huang C, Wang Y, Li X, Ren L, Zhao J, Hu Y, Zhang L, Fan G, Xu J, Gu X, Cheng Z, Yu T, Xia J, Wei Y, Wu W, Xie X, Yin W, Li H, Liu M, Xiao Y, Gao H, Guo L, Xie J, Wang G, Jiang R, Gao Z, Jin Q, Wang J, Cao B. Clinical features of patients infected with 2019 novel coronavirus in Wuhan. China, The Lancet. 2020;395(2020):497–506. https://doi.org/10.1016/S0140-6736(20)30183-5.
Singhal T. A review of coronavirus disease-2019 (COVID-19). Indian J Pediatr. 2020;87:281–6. https://doi.org/10.1007/s12098-020-03263-6.
Guo Y-R, Cao Q-D, Hong Z-S, Tan Y-Y, Chen S-D, Jin H-J, Tan K-S, Wang D-Y, Yan Y. The origin, transmission and clinical therapies on coronavirus disease 2019 (COVID-19) outbreak—an update on the status. Mil Med Res. 2020;7:11. https://doi.org/10.1186/s40779-020-00240-0.
Mitra P, Suri S, Goyal T, Misra R, Singh K, Garg MK, Misra S, Sharma Abhilasha P. Association of comorbidities with coronavirus disease 2019: a review. Ann Natl Acad Med Sci India. 2020;56:102–11. https://doi.org/10.1055/s-0040-1714159.
Hui DS, Azhar EI, Madani TA, Ntoumi F, Kock R, Dar O, Ippolito G, Mchugh TD, Memish ZA, Drosten C, Zumla A, Petersen E. The continuing 2019-nCoV epidemic threat of novel coronaviruses to global health—The latest 2019 novel coronavirus outbreak in Wuhan, China. Int J Infect Dis. 2020;91:264–6. https://doi.org/10.1016/j.ijid.2020.01.009.
Forni D, Cagliani R, Clerici M, Sironi M. Molecular evolution of human coronavirus genomes. Trends Microbiol. 2017;25:35–48. https://doi.org/10.1016/j.tim.2016.09.001.
Yin Y, Wunderink RG. MERS, SARS and other coronaviruses as causes of pneumonia: MERS, SARS and coronaviruses. Respirology. 2018;23:130–7. https://doi.org/10.1111/resp.13196.
de Wit E, van Doremalen N, Falzarano D, Munster VJ. SARS and MERS: recent insights into emerging coronaviruses. Nat Rev Microbiol. 2016;14:523–34. https://doi.org/10.1038/nrmicro.2016.81.
Mousavizadeh L, Ghasemi S. Genotype and phenotype of COVID-19: Their roles in pathogenesis. J Microbiol Immunol Infect. 2020. https://doi.org/10.1016/j.jmii.2020.03.022.
Cui J, Li F, Shi Z-L. Origin and evolution of pathogenic coronaviruses. Nat Rev Microbiol. 2019;17:181–92. https://doi.org/10.1038/s41579-018-0118-9.
Wang N, Shang J, Jiang S, Du L. Subunit vaccines against emerging pathogenic human coronaviruses. Front Microbiol. 2020. https://doi.org/10.3389/fmicb.2020.00298.
Coutard B, Valle C, de Lamballerie X, Canard B, Seidah NG, Decroly E. The spike glycoprotein of the new coronavirus 2019-nCoV contains a furin-like cleavage site absent in CoV of the same clade. Antiviral Res. 2020;176:104742. https://doi.org/10.1016/j.antiviral.2020.104742.
Coronaviridae Study Group of the International Committee on Taxonomy of Viruses, The species Severe acute respiratory syndrome-related coronavirus: classifying 2019-nCoV and naming it SARS-CoV-2, Nat. Microbiol. 2020; 5: 536–544. https://doi.org/10.1038/s41564-020-0695-z.
Zhou P, Yang X-L, Wang X-G, Hu B, Zhang L, Zhang W, Si H-R, Zhu Y, Li B, Huang C-L, Chen H-D, Chen J, Luo Y, Guo H, Jiang R-D, Liu M-Q, Chen Y, Shen X-R, Wang X, Zheng X-S, Zhao K, Chen Q-J, Deng F, Liu L-L, Yan B, Zhan F-X, Wang Y-Y, Xiao G-F, Shi Z-L. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020;579:270–3. https://doi.org/10.1038/s41586-020-2012-7.
Chitranshi N, Gupta VK, Rajput R, Godinez A, Pushpitha K, Shen T, Mirzaei M, You Y, Basavarajappa D, Gupta V, Graham SL. Evolving geographic diversity in SARS-CoV2 and in silico analysis of replicating enzyme 3CLpro targeting repurposed drug candidates. J Transl Med. 2020;18:278. https://doi.org/10.1186/s12967-020-02448-z.
Tiwari M, Mishra D. Investigating the genomic landscape of novel coronavirus (2019-nCoV) to identify non-synonymous mutations for use in diagnosis and drug design. J Clin Virol. 2020;128:104441. https://doi.org/10.1016/j.jcv.2020.104441.
Hassan SS, Choudhury PP, Roy B. SARS-CoV2 envelope protein: non-synonymous mutations and its consequences. Genomics. 2020;112:3890–2. https://doi.org/10.1016/j.ygeno.2020.07.001.
Begum F, Mukherjee D, Thagriki D, Das S, Tripathi PP, Banerjee AK, Ray U. Analyses of spike protein from first deposited sequences of SARS-CoV2 from West Bengal, India. F1000Research. 2020;9:371. https://doi.org/10.12688/f1000research.23805.1.
Guan Z, Ni Y, Hu W, Liang C, Ou J, He L, Liu H, Shan C, Lei DSC, Hui B, Du L, Li G, Zeng K-Y, Yuen R, Chen C, Tang T, Wang P, Chen J, Xiang S, Li J, Wang Z, Liang Y, Peng L, Wei Y, Liu Y, Hu P, Peng J, Wang J, Liu Z, Chen G, Li Z, Zheng S, Qiu J, Luo C, Ye S, Zhu NZ. Clinical characteristics of coronavirus disease 2019 in China. N Engl J Med. 2020. https://doi.org/10.1056/NEJMoa2002032.
Chowell G, Abdirizak F, Lee S, Lee J, Jung E, Nishiura H, Viboud C. Transmission characteristics of MERS and SARS in the healthcare setting: a comparative study. BMC Med. 2015;13:210. https://doi.org/10.1186/s12916-015-0450-0.
Kang CK, Song K-H, Choe PG, Park WB, Bang JH, Kim ES, Park SW, Kim HB, Kim NJ, Cho S, Lee J, Oh M. Clinical and epidemiologic characteristics of spreaders of middle east respiratory syndrome coronavirus during the 2015 Outbreak in Korea. J Korean Med Sci. 2017;32:744. https://doi.org/10.3346/jkms.2017.32.5.744.
Ralph R, Lew J, Zeng T, Francis M, Xue B, Roux M, Toloue Ostadgavahi A, Rubino S, Dawe NJ, Al-Ahdal MN, Kelvin DJ, Richardson CD, Kindrachuk J, Falzarano D, Kelvin AA. 2019-nCoV (Wuhan virus), a novel Coronavirus: human-to-human transmission, travel-related cases, and vaccine readiness. J Infect Dev Ctries. 2020;14:3–17. https://doi.org/10.3855/jidc.12425.
van Doremalen N, Bushmaker T, Morris DH, Holbrook MG, Gamble A, Williamson BN, Tamin A, Harcourt JL, Thornburg NJ, Gerber SI, Lloyd-Smith JO, de Wit E, Munster VJ. Aerosol and surface stability of SARS-CoV-2 as compared with SARS-CoV-1. N Engl J Med. 2020;382:1564–7. https://doi.org/10.1056/NEJMc2004973.
Hindson J. COVID-19: faecal–oral transmission? Nat Rev Gastroenterol Hepatol. 2020. https://doi.org/10.1038/s41575-020-0295-7.
Xia J, Tong J, Liu M, Shen Y, Guo D. Evaluation of coronavirus in tears and conjunctival secretions of patients with SARS-CoV-2 infection. J Med Virol. 2020;92:589–94. https://doi.org/10.1002/jmv.25725.
Rothan HA, Byrareddy SN. The epidemiology and pathogenesis of coronavirus disease (COVID-19) outbreak. J Autoimmun. 2020;109:102433. https://doi.org/10.1016/j.jaut.2020.102433.
Donoghue M, Hsieh F, Baronas E, Godbout K, Gosselin M, Stagliano N, Donovan M, Woolf B, Robison K, Jeyaseelan R, Breitbart RE, Acton S. A novel angiotensin-converting enzyme-related carboxypeptidase (ACE2) converts angiotensin i to angiotensin 1–9. Circ Res. 2000. https://doi.org/10.1161/01.RES.87.5.e1.
Turner AJ, Hiscox JA, Hooper NM. ACE2: from vasopeptidase to SARS virus receptor. Trends Pharmacol Sci. 2004;25:291–4. https://doi.org/10.1016/j.tips.2004.04.001.
Ohtsuki M, Morimoto S, Izawa H, Ismail TF, Ishibashi-Ueda H, Kato Y, Horii T, Isomura T, Suma H, Nomura M, Hishida H, Kurahashi H, Ozaki Y. Angiotensin converting enzyme 2 gene expression increased compensatory for left ventricular remodeling in patients with end-stage heart failure. Int J Cardiol. 2010;145:333–4. https://doi.org/10.1016/j.ijcard.2009.11.057.
Tikellis C, Bernardi S, Burns WC. Angiotensin-converting enzyme 2 is a key modulator of the renin–angiotensin system in cardiovascular and renal disease. Curr Opin Nephrol Hypertens. 2011;20:62–8. https://doi.org/10.1097/MNH.0b013e328341164a.
Paz Ocaranza M, Riquelme JA, García L, Jalil JE, Chiong M, Santos RAS, Lavandero S. Counter-regulatory renin–angiotensin system in cardiovascular disease. Nat Rev Cardiol. 2020;17:116–29. https://doi.org/10.1038/s41569-019-0244-8.
Zhang N, Jiang S, Du L. Current advancements and potential strategies in the development of MERS-CoV vaccines. Expert Rev Vaccines. 2014;13:761–74. https://doi.org/10.1586/14760584.2014.912134.
Xia S, Zhu Y, Liu M, Lan Q, Xu W, Wu Y, Ying T, Liu S, Shi Z, Jiang S, Lu L. Fusion mechanism of 2019-nCoV and fusion inhibitors targeting HR1 domain in spike protein. Cell Mol Immunol. 2020. https://doi.org/10.1038/s41423-020-0374-2.
Yu F, Du L, Ojcius DM, Pan C, Jiang S. Measures for diagnosing and treating infections by a novel coronavirus responsible for a pneumonia outbreak originating in Wuhan, China. Microbes Infect. 2020;22:74–9. https://doi.org/10.1016/j.micinf.2020.01.003.
de Wilde AH, Snijder EJ, Kikkert M, van Hemert MJ (2017) Host factors in coronavirus replication. In: Tripp RA, Tompkins SM (Eds), Roles Host Gene Non-Coding RNA Expr. Virus Infect., Springer International Publishing, Cham, 2017: pp 1–42. https://doi.org/10.1007/82_2017_25.
Song Z, Xu Y, Bao L, Zhang L, Yu P, Qu Y, Zhu H, Zhao W, Han Y, Qin C. From SARS to MERS, thrusting coronaviruses into the spotlight. Viruses. 2019;11:59. https://doi.org/10.3390/v11010059.
Narayanan K, Ramirez SI, Lokugamage KG, Makino S. Coronavirus nonstructural protein 1: common and distinct functions in the regulation of host and viral gene expression. Virus Res. 2015;202:89–100. https://doi.org/10.1016/j.virusres.2014.11.019.
South AM, Tomlinson L, Edmonston D, Hiremath S, Sparks MA. Controversies of renin–angiotensin system inhibition during the COVID-19 pandemic. Nat Rev Nephrol. 2020;16:305–7. https://doi.org/10.1038/s41581-020-0279-4.
Xiong Y, Liu Y, Cao L, Wang D, Guo M, Jiang A, Guo D, Hu W, Yang J, Tang Z, Wu H, Lin Y, Zhang M, Zhang Q, Shi M, Liu Y, Zhou Y, Lan K, Chen Y. Transcriptomic characteristics of bronchoalveolar lavage fluid and peripheral blood mononuclear cells in COVID-19 patients. Emerg Microbes Infect. 2020;9:761–70. https://doi.org/10.1080/22221751.2020.1747363.
Ji H-L, Zhao R, Matalon S, Matthay MA. Elevated plasmin(ogen) as a common risk factor for COVID-19 susceptibility. Physiol Rev. 2020;100:1065–75. https://doi.org/10.1152/physrev.00013.2020.
Ji W, Wang W, Zhao X, Zai J, Li X. Cross-species transmission of the newly identified coronavirus 2019-nCoV. J Med Virol. 2020;92:433–40. https://doi.org/10.1002/jmv.25682.
Wang D, Hu B, Hu C, Zhu F, Liu X, Zhang J, Wang B, Xiang H, Cheng Z, Xiong Y, Zhao Y, Li Y, Wang X, Peng Z. Clinical characteristics of 138 hospitalized patients with 2019 novel coronavirus-infected pneumonia in Wuhan, China. JAMA. 2020;323:1061. https://doi.org/10.1001/jama.2020.1585.
Rokni M, Ghasemi V, Tavakoli Z. Immune responses and pathogenesis of SARS-CoV -2 during an outbreak in Iran: comparison with SARS and MERS. Rev Med Virol. 2020. https://doi.org/10.1002/rmv.2107.
Chavez S, Long B, Koyfman A, Liang SY. Coronavirus disease (COVID-19): a primer for emergency physicians. Am J Emerg Med. 2020. https://doi.org/10.1016/j.ajem.2020.03.036.
Chen G, Wu D, Guo W, Cao Y, Huang D, Wang H, Wang T, Zhang X, Chen H, Yu H, Zhang X, Zhang M, Wu S, Song J, Chen T, Han M, Li S, Luo X, Zhao J, Ning Q. Clinical and immunological features of severe and moderate coronavirus disease 2019. J Clin Invest. 2020;130:2620–9. https://doi.org/10.1172/JCI137244.
Wong SH, Lui RN, Sung JJ. Covid-19 and the digestive system. J Gastroenterol Hepatol. 2020;35:744–8. https://doi.org/10.1111/jgh.15047.
Lechien JR, Chiesa-Estomba CM, De Siati DR, Horoi M, Le Bon SD, Rodriguez A, Dequanter D, Blecic S, El Afia F, Distinguin L, Chekkoury-Idrissi Y, Hans S, Delgado IL, Calvo-Henriquez C, Lavigne P, Falanga C, Barillari MR, Cammaroto G, Khalife M, Leich P, Souchay C, Rossi C, Journe F, Hsieh J, Edjlali M, Carlier R, Ris L, Lovato A, De Filippis C, Coppee F, Fakhry N, Ayad T, Saussez S. Olfactory and gustatory dysfunctions as a clinical presentation of mild-to-moderate forms of the coronavirus disease (COVID-19): a multicenter European study. Eur Arch Otorhinolaryngol. 2020;277:2251–61. https://doi.org/10.1007/s00405-020-05965-1.
Xu Z, Shi L, Wang Y, Zhang J, Huang L, Zhang C, Liu S, Zhao P, Liu H, Zhu L, Tai Y, Bai C, Gao T, Song J, Xia P, Dong J, Zhao J, Wang F-S. Pathological findings of COVID-19 associated with acute respiratory distress syndrome. Lancet Respir Med. 2020;8:420–2. https://doi.org/10.1016/S2213-2600(20)30076-X.
Liu Y, Yang Y, Zhang C, Huang F, Wang F, Yuan J, Wang Z, Li J, Li J, Feng C, Zhang Z, Wang L, Peng L, Chen L, Qin Y, Zhao D, Tan S, Yin L, Xu J, Zhou C, Jiang C, Liu L. Clinical and biochemical indexes from 2019-nCoV infected patients linked to viral loads and lung injury. Sci China Life Sci. 2020;63:364–74. https://doi.org/10.1007/s11427-020-1643-8.
Chen N, Zhou M, Dong X, Qu J, Gong F, Han Y, Qiu Y, Wang J, Liu Y, Wei Y, Xia J, Yu T, Zhang X, Zhang L. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. The Lancet. 2020;395:507–13. https://doi.org/10.1016/S0140-6736(20)30211-7.
Zhu N, Zhang D, Wang W, Li X, Yang B, Song J, Zhao X, Huang B, Shi W, Lu R, Niu P, Zhan F, Ma X, Wang D, Xu W, Wu G, Gao GF, Tan W. A novel coronavirus from patients with pneumonia in China. N Engl J Med. 2019;382(2020):727–33. https://doi.org/10.1056/NEJMoa2001017.
Lin L, Jiang X, Zhang Z, Huang S, Zhang Z, Fang Z, Gu Z, Gao L, Shi H, Mai L, Liu Y, Lin X, Lai R, Yan Z, Li X, Shan H. Gastrointestinal symptoms of 95 cases with SARS-CoV-2 infection. Gut. 2020. https://doi.org/10.1136/gutjnl-2020-321013.
Fan Z, Chen L, Li J, Tian C, Zhang Y, Huang S, Liu Z, Cheng J. Clinical features of COVID-19 related liver damage. Gastroenterology. 2020. https://doi.org/10.1101/2020.02.26.20026971.
Nazinitsky A, Rosenthal KS. Cytokine storms: systemic disasters of infectious diseases. Infect Dis Clin Pract. 2010;18:188–92. https://doi.org/10.1097/IPC.0b013e3181d2ee41.
Liu T, Zhang J, Yang Y, Zhang L, Ma H, Li Z, Zhang J, Cheng J, Zhang X, Wu G, Yi J. The potential role of IL-6 in monitoring coronavirus disease 2019. Infect Dis (except HIV/AIDS). 2020. https://doi.org/10.1101/2020.03.01.20029769.
Tang X, Wu C, Li X, Song Y, Yao X, Wu X, Duan Y, Zhang H, Wang Y, Qian Z, Cui J, Lu J. On the origin and continuing evolution of SARS-CoV-2. Natl Sci Rev. 2020. https://doi.org/10.1093/nsr/nwaa036.
Lingeswaran M, Goyal T, Ghosh R, Suri S, Mitra P, Misra S, Sharma P. Inflammation, Immunity and Immunogenetics in COVID-19: a narrative review. Indian J Clin Biochem. 2020;35:260–73. https://doi.org/10.1007/s12291-020-00897-3.
El-Sahly HM, Atmar RL, Glezen WP, Greenberg SB. Spectrum of clinical illness in hospitalized patients with “common cold” virus infections. Clin Infect Dis. 2000;31:96–100. https://doi.org/10.1086/313937.
Stolz D, Papakonstantinou E, Grize L, Schilter D, Strobel W, Louis R, Schindler C, Hirsch HH, Tamm M. Time-course of upper respiratory tract viral infection and COPD exacerbation. Eur Respir J. 2019;54:1900407. https://doi.org/10.1183/13993003.00407-2019.
Kiely D. Haemodynamic and endocrine effects of type 1 angiotensin II receptor blockade in patients with hypoxaemic cor pulmonale. Cardiovasc Res. 1997;33:201–8. https://doi.org/10.1016/S0008-6363(96)00180-0.
Imai Y, Kuba K, Rao S, Huan Y, Guo F, Guan B, Yang P, Sarao R, Wada T, Leong-Poi H, Crackower MA, Fukamizu A, Hui C-C, Hein L, Uhlig S, Slutsky AS, Jiang C, Penninger JM. Angiotensin-converting enzyme 2 protects from severe acute lung failure. Nature. 2005;436:112–6. https://doi.org/10.1038/nature03712.
Zhao Y, Zhao Z, Wang Y, Zhou Y, Ma Y, Zuo W. Single-cell RNA expression profiling of ACE2, the receptor of SARS-CoV-2. Bioinformatics. 2020. https://doi.org/10.1101/2020.01.26.919985.
Zhang R, Wu Y, Zhao M, Liu C, Zhou L, Shen S, Liao S, Yang K, Li Q, Wan H. Role of HIF-1α in the regulation ACE and ACE2 expression in hypoxic human pulmonary artery smooth muscle cells. Am J Physiol-Lung Cell Mol Physiol. 2009;297:631–40. https://doi.org/10.1152/ajplung.90415.2008.
Kuba K, Imai Y, Ohto-Nakanishi T, Penninger JM. Trilogy of ACE2: a peptidase in the renin–angiotensin system, a SARS receptor, and a partner for amino acid transporters. Pharmacol Ther. 2010;128:119–28. https://doi.org/10.1016/j.pharmthera.2010.06.003.
Fang F, Liu GC, Zhou X, Yang S, Reich HN, Williams V, Hu A, Pan J, Konvalinka A, Oudit GY, Scholey JW, John R. Loss of ACE2 exacerbates murine renal ischemia-reperfusion injury. PLoS ONE. 2013;8:e71433. https://doi.org/10.1371/journal.pone.0071433.
Soler MJ, Wysocki J, Batlle D. ACE2 alterations in kidney disease. Nephrol Dial Transplant. 2013;28:2687–97. https://doi.org/10.1093/ndt/gft320.
Zhang W, Du R-H, Li B, Zheng X-S, Yang X-L, Hu B, Wang Y-Y, Xiao G-F, Yan B, Shi Z-L, Zhou P. Molecular and serological investigation of 2019-nCoV infected patients: implication of multiple shedding routes. Emerg Microbes Infect. 2020;9:386–9. https://doi.org/10.1080/22221751.2020.1729071.
Xu Y, Li X, Zhu B, Liang H, Fang C, Gong Y, Guo Q, Sun X, Zhao D, Shen J, Zhang H, Liu H, Xia H, Tang J, Zhang K, Gong S. Characteristics of pediatric SARS-CoV-2 infection and potential evidence for persistent fecal viral shedding. Nat Med. 2020;26:502–5. https://doi.org/10.1038/s41591-020-0817-4.
Xiao F, Tang M, Zheng X, Liu Y, Li X, Shan H. Evidence for gastrointestinal infection of SARS-CoV-2. Gastroenterology. 2020. https://doi.org/10.1053/j.gastro.2020.02.055.
Harmer D, Gilbert M, Borman R, Clark KL. Quantitative mRNA expression profiling of ACE 2, a novel homologue of angiotensin converting enzyme. FEBS Lett. 2002;532:107–10. https://doi.org/10.1016/S0014-5793(02)03640-2.
Zhao D, Yao F, Wang L, Zheng L, Gao Y, Ye J, Guo F, Zhao H, Gao R. A comparative study on the clinical features of COVID-19 pneumonia to other pneumonias. Clin Infect Dis. 2020. https://doi.org/10.1093/cid/ciaa247.
Holshue ML, DeBolt C, Lindquist S, Lofy KH, Wiesman J, Bruce H, Spitters C, Ericson K, Wilkerson S, Tural A, Diaz G, Cohn A, Fox L, Patel A, Gerber SI, Kim L, Tong S, Lu X, Lindstrom S, Pallansch MA, Weldon WC, Biggs HM, Uyeki TM, Pillai SK. First case of 2019 novel coronavirus in the United States. N Engl J Med. 2019;382(2020):929–36. https://doi.org/10.1056/NEJMoa2001191.
Tang A, Tong Z, Wang H, Dai Y, Li K, Liu J, Wu W, Yuan C, Yu M, Li P, Yan J. Detection of novel coronavirus by RT-PCR in stool specimen from asymptomatic child, China. Emerg Infect Dis. 2020. https://doi.org/10.3201/eid2606.200301.
Young BE, Ong SWX, Kalimuddin S, Low JG, Tan SY, Loh J, Ng O-T, Marimuthu K, Ang LW, Mak TM, Lau SK, Anderson DE, Chan KS, Tan TY, Ng TY, Cui L, Said Z, Kurupatham L, Chen MI-C, Chan M, Vasoo S, Wang L-F, Tan BH, Lin RTP, Lee VJM, Leo Y-S, Lye DC. for the Singapore 2019 novel coronavirus outbreak research team, epidemiologic features and clinical course of patients infected with SARS-CoV-2 in Singapore. JAMA. 2020. https://doi.org/10.1001/jama.2020.3204.
Zhang C, Shi L, Wang F-S. Liver injury in COVID-19: management and challenges. Lancet Gastroenterol Hepatol. 2020;5:428–30. https://doi.org/10.1016/S2468-1253(20)30057-1.
Chai X, Hu L, Zhang Y, Han W, Lu Z, Ke A, Zhou J, Shi G, Fang N, Fan J, Cai J, Fan J, Lan F. Specific ACE2 expression in cholangiocytes may cause liver damage after 2019-nCoV infection. Genomics. 2020. https://doi.org/10.1101/2020.02.03.931766.
Banales JM, Huebert RC, Karlsen T, Strazzabosco M, LaRusso NF, Gores GJ. Cholangiocyte pathobiology. Nat Rev Gastroenterol Hepatol. 2019;16:269–81. https://doi.org/10.1038/s41575-019-0125-y.
dong chen, Li X, qifa song, Hu C, Su F, Dai J (2020) Hypokalemia and clinical implications in patients with coronavirus disease 2019 (COVID-19). Infect Dis (except HIV/AIDS); 2020. https://doi.org/10.1101/2020.02.27.20028530.
Li Y, Bai W, Hashikawa T. The neuroinvasive potential of SARS-CoV2 may play a role in the respiratory failure of COVID-19 patients. J Med Virol. 2020;92:552–5. https://doi.org/10.1002/jmv.25728.
Lippi G, Plebani M. Laboratory abnormalities in patients with COVID-2019 infection. Clin Chem Lab Med CCLM. 2020. https://doi.org/10.1515/cclm-2020-0198.
Li B, Yang J, Zhao F, Zhi L, Wang X, Liu L, Bi Z, Zhao Y. Prevalence and impact of cardiovascular metabolic diseases on COVID-19 in China. Clin Res Cardiol. 2020. https://doi.org/10.1007/s00392-020-01626-9.
Zhou F, Yu T, Du R, Fan G, Liu Y, Liu Z, Xiang J, Wang Y, Song B, Gu X, Guan L, Wei Y, Li H, Wu X, Xu J, Tu S, Zhang Y, Chen H, Cao B. Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: a retrospective cohort study. The Lancet. 2020;395:1054–62. https://doi.org/10.1016/S0140-6736(20)30566-3.
Zheng Y-Y, Ma Y-T, Zhang J-Y, Xie X. COVID-19 and the cardiovascular system. Nat Rev Cardiol. 2020;17:259–60. https://doi.org/10.1038/s41569-020-0360-5.
Hoffmann M, Kleine-Weber H, Krüger N, Müller M, Drosten C, Pöhlmann S. The novel coronavirus 2019 (2019-nCoV) uses the SARS-coronavirus receptor ACE2 and the cellular protease TMPRSS2 for entry into target cells. Mol Biol. 2019. https://doi.org/10.1101/2020.01.31.929042.
Hashimoto T, Perlot T, Rehman A, Trichereau J, Ishiguro H, Paolino M, Sigl V, Hanada T, Hanada R, Lipinski S, Wild B, Camargo SMR, Singer D, Richter A, Kuba K, Fukamizu A, Schreiber S, Clevers H, Verrey F, Rosenstiel P, Penninger JM. ACE2 links amino acid malnutrition to microbial ecology and intestinal inflammation. Nature. 2012;487:477–81. https://doi.org/10.1038/nature11228.
South AM, Diz DI, Chappell MC. COVID-19, ACE2, and the cardiovascular consequences. Am J Physiol-Heart Circ Physiol. 2020;318:1084–90. https://doi.org/10.1152/ajpheart.00217.2020.
Gheblawi M, Wang K, Viveiros A, Nguyen Q, Zhong J-C, Turner AJ, Raizada MK, Grant MB, Oudit GY. Angiotensin converting enzyme 2: SARS-CoV-2 receptor and regulator of the renin-angiotensin system. Circ Res. 2020;1:200. https://doi.org/10.1161/CIRCRESAHA.120.317015.
Yuan Y-M, Luo L, Guo Z, Yang M, Ye R-S, Luo C. Activation of renin–angiotensin–aldosterone system (RAAS) in the lung of smoking-induced pulmonary arterial hypertension (PAH) rats. J Renin Angiotensin Aldosterone Syst. 2015;16:249–53. https://doi.org/10.1177/1470320315576256.
Lin M, Gao P, Zhao T, He L, Li M, Li Y, Shui H, Wu X. Calcitriol regulates angiotensin-converting enzyme and angiotensin converting-enzyme 2 in diabetic kidney disease. Mol Biol Rep. 2016;43:397–406. https://doi.org/10.1007/s11033-016-3971-5.
Bindom SM, Hans CP, Xia H, Boulares AH, Lazartigues E. Angiotensin I-converting enzyme type 2 (ACE2) gene therapy improves glycemic control in diabetic mice. Diabetes. 2010;59:2540–8. https://doi.org/10.2337/db09-0782.
Lu C-L, Wang Y, Yuan L, Li Y, Li X-Y. The angiotensin-converting enzyme 2/angiotensin (1–7)/Mas axis protects the function of pancreatic β cells by improving the function of islet microvascular endothelial cells. Int J Mol Med. 2014;34:1293–300. https://doi.org/10.3892/ijmm.2014.1917.
Yang X, Yu Y, Xu J, Shu H, Xia J, Liu H, Wu Y, Zhang L, Yu Z, Fang M, Yu T, Wang Y, Pan S, Zou X, Yuan S, Shang Y. Clinical course and outcomes of critically ill patients with SARS-CoV-2 pneumonia in Wuhan, China: a single-centered, retrospective, observational study. Lancet Respir Med. 2020. https://doi.org/10.1016/S2213-2600(20)30079-5.
Tikellis C, Thomas MC. Angiotensin-converting enzyme 2 (ACE2) is a key modulator of the renin angiotensin system in health and disease. Int J Pept. 2012;2012:1–8. https://doi.org/10.1155/2012/256294.
Wu C, Chen X, Cai Y, Xia J, Zhou X, Xu S, Huang H, Zhang L, Zhou X, Du C, Zhang Y, Song J, Wang S, Chao Y, Yang Z, Xu J, Zhou X, Chen D, Xiong W, Xu L, Zhou F, Jiang J, Bai C, Zheng J, Song Y. Risk factors associated with acute respiratory distress syndrome and death in patients with coronavirus disease 2019 pneumonia in Wuhan, China. JAMA Intern Med. 2020. https://doi.org/10.1001/jamainternmed.2020.0994.
Arun Krishnan R, Elizabeth Thomas R, Sukumaran A, Paul JK, Vasudevan DM. COVID-19: current trends in invitro diagnostics. Indian J Clin Biochem. 2020;35:285–9. https://doi.org/10.1007/s12291-020-00906-5.
Chu DKW, Pan Y, Cheng SMS, Hui KPY, Krishnan P, Liu Y, Ng DYM, Wan CKC, Yang P, Wang Q, Peiris M, Poon LLM. Molecular diagnosis of a novel coronavirus (2019-nCoV) causing an outbreak of pneumonia. Clin Chem. 2020;66:549–55. https://doi.org/10.1093/clinchem/hvaa029.
Tahamtan A, Ardebili A. Real-time RT-PCR in COVID-19 detection: issues affecting the results. Expert Rev Mol Diagn. 2020;20:453–4. https://doi.org/10.1080/14737159.2020.1757437.
Thabet L, Mhalla S, Naija H, Jaoua MA, Hannachi N, Fki- Berrajah L, Toumi A, Karray-Hakim H. SARS-CoV-2 infection virological diagnosis. Tunis Med. 2020;98(4):304–8.
Notomi T. Loop-mediated isothermal amplification of DNA. Nucleic Acids Res. 2000;28:63e–63. https://doi.org/10.1093/nar/28.12.e63.
Centers for Disease Control and Prevention. 2020. Coronavirus, Disease 2019 (COVID-19)., Centers for Disease Control and Prevention. 2020. Coronavirus Disease 2019 (COVID-19)., (n.d.).
Kellner MJ, Koob JG, Gootenberg JS, Abudayyeh OO, Zhang F. SHERLOCK: nucleic acid detection with CRISPR nucleases. Nat Protoc. 2019;14:2986–3012. https://doi.org/10.1038/s41596-019-0210-2.
Azhar M, Phutela R, Ansari AH, Sinha D, Sharma N, Kumar M, Aich M, Sharma S, Rauthan R, Singhal K, Lad H, Patra PK, Makharia G, Chandak GR, Chakraborty D, Maiti S. Rapid, field-deployable nucleobase detection and identification using FnCas9. Mol Biol. 2020. https://doi.org/10.1101/2020.04.07.028167.
Wang Y, Wang Y, Chen Y, Qin Q. Unique epidemiological and clinical features of the emerging 2019 novel coronavirus pneumonia (COVID-19) implicate special control measures. J Med Virol. 2020;92:568–76. https://doi.org/10.1002/jmv.25748.
Ferrario CM, Jessup J, Chappell MC, Averill DB, Brosnihan KB, Tallant EA, Diz DI, Gallagher PE. Effect of angiotensin-converting enzyme inhibition and angiotensin II receptor blockers on cardiac angiotensin-converting enzyme 2. Circulation. 2005;111:2605–10. https://doi.org/10.1161/CIRCULATIONAHA.104.510461.
Koka V, Huang XR, Chung ACK, Wang W, Truong LD, Lan HY. Angiotensin II up-regulates angiotensin i-converting enzyme (ACE), but down-regulates ACE2 via the AT1-ERK/p38 MAP kinase pathway. Am J Pathol. 2008;172:1174–83. https://doi.org/10.2353/ajpath.2008.070762.
Patel VB, Zhong J-C, Grant MB, Oudit GY. Role of the ACE2/angiotensin 1–7 axis of the renin-angiotensin system in heart failure. Circ Res. 2016;118:1313–26. https://doi.org/10.1161/CIRCRESAHA.116.307708.
Jia H. Pulmonary angiotensin-converting enzyme 2 (ACE2) and inflammatory lung disease. SHOCK. 2016;46:239–48. https://doi.org/10.1097/SHK.0000000000000633.
Oudit GY, Zhong J, Basu R, Guo D, Penninger JM, Kassiri Z. Angiotensin converting enzyme 2 suppresses pathological hypertrophy, myocardial fibrosis and diastolic dysfunction. J Card Fail. 2010;16:S16. https://doi.org/10.1016/j.cardfail.2010.06.054.
Wei R, Christakos S. Mechanisms underlying the regulation of innate and adaptive immunity by vitamin D. Nutrients. 2015;7:8251–60. https://doi.org/10.3390/nu7105392.
Gombart AF, Pierre A, Maggini S. A review of micronutrients and the immune system-working in harmony to reduce the risk of infection. Nutrients. 2020;12:236. https://doi.org/10.3390/nu12010236.
Himani R, Kumar JA, Ansari AA, Mahdi D, Sharma B, Karunanand SK. Datta, blood lead levels in occupationally exposed workers involved in battery factories of Delhi-NCR Region: effect on vitamin D and calcium metabolism. Indian J Clin Biochem. 2020;35:80–7. https://doi.org/10.1007/s12291-018-0797-z.
Himani K, Raman K, Busi D. Sudip Kumar, Association of vitamin D receptor (VDR) gene polymorphism with blood lead levels in occupationally lead-exposed male battery workers in Delhi—National capital region, India. Indian J Biochem Biophys IJBB. 2020;57:236–44.
Rondanelli M, Miccono A, Lamburghini S, Avanzato I, Riva A, Allegrini P, Faliva MA, Peroni G, Nichetti M, Perna S. Self-care for common colds: the pivotal role of vitamin D, Vitamin C, zinc, and Echinacea in Three main immune interactive clusters (physical barriers, innate and adaptive immunity) involved during an episode of common colds—practical advice on dosages and on the time to take these nutrients/botanicals in order to prevent or treat common colds. Evid Based Complement Alternat Med. 2018;2018:1–36. https://doi.org/10.1155/2018/5813095.
Kumar Himani R, Haq A, Wimalawansa SJ, Sharma A. Putative roles of vitamin D in modulating immune response and immunopathology associated with COVID-19. Virus Res. 2020. https://doi.org/10.1016/j.virusres.2020.198235.
Grant WB, Lahore H, McDonnell SL, Baggerly CA, French CB, Aliano JL, Bhattoa HP. Evidence that vitamin D supplementation could reduce risk of influenza and COVID-19 infections and deaths. Nutrients. 2020;12:988. https://doi.org/10.3390/nu12040988.
Lei G-S, Zhang C, Cheng B-H, Lee C-H. Mechanisms of action of vitamin D as supplemental therapy for pneumocystis pneumonia. Antimicrob Agents Chemother. 2017;61(e01226–17):e01226-e1317. https://doi.org/10.1128/AAC.01226-17.
Mousavi S, Bereswill S, Heimesaat MM. Immunomodulatory and antimicrobial effects of vitamin C. Eur J Microbiol Immunol. 2019;9:73–9. https://doi.org/10.1556/1886.2019.00016.
Colunga Biancatelli RML, Berrill M, Marik PE. The antiviral properties of vitamin C. Expert Rev Anti Infect Ther. 2020;18:99–101. https://doi.org/10.1080/14787210.2020.1706483.
Li YC. Molecular mechanism of vitamin D in the cardiovascular system. J Investig Med. 2011;59:868–71. https://doi.org/10.2310/JIM.0b013e31820ee448.
Das SK. The pathophysiology, diagnosis and treatment of corona virus disease 2019 (COVID-19). Indian J Clin Biochem. 2020;35:385–96. https://doi.org/10.1007/s12291-020-00919-0.
Reina J. Remdesivir, la esperanza antiviral frente al SARS-CoV-2. Rev Esp Quimioter. 2020;33:176–9. https://doi.org/10.37201/req/028.2020.
Du Y, Chen X. Favipiravir: pharmacokinetics and concerns about clinical trials for 2019-nCoV infection. Clin Pharmacol Ther. 2020;108:242–7. https://doi.org/10.1002/cpt.1844.
Srinivas P, Sacha GL, Koval C. Antivirals for COVID-19. Cleve Clin J Med. 2020. https://doi.org/10.3949/ccjm.87a.ccc030.
Liu J, Cao R, Xu M, Wang X, Zhang H, Hu H, Li Y, Hu Z, Zhong W, Wang M. Hydroxychloroquine, a less toxic derivative of chloroquine, is effective in inhibiting SARS-CoV-2 infection in vitro. Cell Discov. 2020;6:16. https://doi.org/10.1038/s41421-020-0156-0.
Zhou D, Dai S-M, Tong Q. COVID-19: a recommendation to examine the effect of hydroxychloroquine in preventing infection and progression. J Antimicrob Chemother. 2020;75:1667–70. https://doi.org/10.1093/jac/dkaa114.
Gautret P, Lagier J-C, Parola P, Hoang VT, Meddeb L, Mailhe M, Doudier B, Courjon J, Giordanengo V, Vieira VE, Dupont HT, Honoré S, Colson P, Chabrière E, La Scola B, Rolain J-M, Brouqui P, Raoult D. Hydroxychloroquine and azithromycin as a treatment of COVID-19: results of an open-label non-randomized clinical trial. Int J Antimicrob Agents. 2020;1:200. https://doi.org/10.1016/j.ijantimicag.2020.105949.
Bloch EM, Shoham S, Casadevall A, Sachais BS, Shaz B, Winters JL, van Buskirk C, Grossman BJ, Joyner M, Henderson JP, Pekosz A, Lau B, Wesolowski A, Katz L, Shan H, Auwaerter PG, Thomas D, Sullivan DJ, Paneth N, Gehrie E, Spitalnik S, Hod EA, Pollack L, Nicholson WT, Pirofski L, Bailey JA, Tobian AAR. Deployment of convalescent plasma for the prevention and treatment of COVID-19. J Clin Invest. 2020;130:2757–65. https://doi.org/10.1172/JCI138745.
Zhang B, Liu S, Tan T, Huang W, Dong Y, Chen L, Chen Q, Zhang L, Zhong Q, Zhang X, Zou Y, Zhang S. Treatment with convalescent plasma for critically ill patients with severe acute respiratory syndrome coronavirus 2 infection. Chest. 2020;158:e9–13. https://doi.org/10.1016/j.chest.2020.03.039.
Cunningham AC, Goh HP, Koh D. Treatment of COVID-19: old tricks for new challenges. Crit Care. 2020;24:91. https://doi.org/10.1186/s13054-020-2818-6.
Monteagudo LA, Boothby A, Gertner E. Continuous intravenous anakinra infusion to calm the cytokine storm in macrophage activation syndrome. ACR Open Rheumatol. 2020;2:276–82. https://doi.org/10.1002/acr2.11135.
Ramamoorthy S, Cidlowski JA. Corticosteroids. Rheum Dis Clin N Am. 2016;42:15–31. https://doi.org/10.1016/j.rdc.2015.08.002.
Golchin A, Farahany TZ, Khojasteh A, Soleimanifar F, Ardeshirylajimi A. The clinical trials of mesenchymal stem cell therapy in skin diseases: an update and concise review. Curr Stem Cell Res Ther. 2019;14:22–33. https://doi.org/10.2174/1574888X13666180913123424.
Leng Z, Zhu R, Hou W, Feng Y, Yang Y, Han Q, Shan G, Meng F, Du D, Wang S, Fan J, Wang W, Deng L, Shi H, Li H, Hu Z, Zhang F, Gao J, Liu H, Li X, Zhao Y, Yin K, He X, Gao Z, Wang Y, Yang B, Jin R, Stambler I, Lim LW, Su H, Moskalev A, Cano A, Chakrabarti S, Min K-J, Ellison-Hughes G, Caruso C, Jin K, Zhao RC. Transplantation of ACE2- mesenchymal stem cells improves the outcome of patients with COVID-19 pneumonia. Aging Dis. 2020;11:216. https://doi.org/10.14336/AD.2020.0228.
Satyam A, Tsokos GC. Curb complement to cure COVID-19. Clin Immunol. 2020;221:108603. https://doi.org/10.1016/j.clim.2020.108603.
Ibrahim H, Perl A, Smith D, Lewis T, Kon Z, Goldenberg R, Yarta K, Staniloae C, Williams M. Therapeutic blockade of inflammation in severe COVID-19 infection with intravenous N-acetylcysteine. Clin Immunol. 2020;219:108544. https://doi.org/10.1016/j.clim.2020.108544.
Morales-Ortega A, Bernal-Bello D, Llarena-Barroso C, Frutos-Pérez B, Duarte-Millán MÁ, García de Viedma-García V, Farfán-Sedano AI, Canalejo-Castrillero E, Ruiz-Giardín JM, Ruiz-Ruiz J, San Martín-López JV. Imatinib for COVID-19: a case report. Clin Immunol. 2020;218:108518. https://doi.org/10.1016/j.clim.2020.108518.
Treating COVID-19 with colchicine in community healthcare setting | Elsevier Enhanced Reader, (n.d.). https://doi.org/10.1016/j.clim.2020.108490.
Savarino A, Boelaert JR, Cassone A, Majori G, Cauda R. Effects of chloroquine on viral infections: an old drug against today’s diseases. Lancet Infect Dis. 2003;3:722–7. https://doi.org/10.1016/S1473-3099(03)00806-5.
Sheahan TP, Sims AC, Leist SR, Schäfer A, Won J, Brown AJ, Montgomery SA, Hogg A, Babusis D, Clarke MO, Spahn JE, Bauer L, Sellers S, Porter D, Feng JY, Cihlar T, Jordan R, Denison MR, Baric RS. Comparative therapeutic efficacy of remdesivir and combination lopinavir, ritonavir, and interferon beta against MERS-CoV. Nat Commun. 2020;11:222. https://doi.org/10.1038/s41467-019-13940-6.
Agostini ML, Andres EL, Sims AC, Graham RL, Sheahan TP, Lu X, Smith EC, Case JB, Feng JY, Jordan R, Ray AS, Cihlar T, Siegel D, Mackman RL, Clarke MO, Baric RS, Denison MR. Coronavirus susceptibility to the antiviral Remdesivir (GS-5734) is mediated by the viral polymerase and the proofreading exoribonuclease. MBio. 2018. https://doi.org/10.1128/mBio.00221-18.
Furuta Y, Komeno T, Nakamura T. Favipiravir (T-705), a broad spectrum inhibitor of viral RNA polymerase. Proc Jpn Acad Ser B. 2017;93:449–63. https://doi.org/10.2183/pjab.93.027.
Cao B, Wang Y, Wen D, Liu W, Wang J, Fan G, Ruan L, Song B, Cai Y, Wei M, Li X, Xia J, Chen N, Xiang J, Yu T, Bai T, Xie X, Zhang L, Li C, Yuan Y, Chen H, Li H, Huang H, Tu S, Gong F, Liu Y, Wei Y, Dong C, Zhou F, Gu X, Xu J, Liu Z, Zhang Y, Li H, Shang L, Wang K, Li K, Zhou X, Dong X, Qu Z, Lu S, Hu X, Ruan S, Luo S, Wu J, Peng L, Cheng F, Pan L, Zou J, Jia C, Wang J, Liu X, Wang S, Wu X, Ge Q, He J, Zhan H, Qiu F, Guo L, Huang C, Jaki T, Hayden FG, Horby PW, Zhang D, Wang C. A trial of Lopinavir-Ritonavir in adults hospitalized with severe covid-19. N Engl J Med. 2020;1:200. https://doi.org/10.1056/NEJMoa2001282.
Chandwani A, Shuter J. Lopinavir/ritonavir in the treatment of HIV-1 infection: a review. Ther Clin Risk Manag. 2008;4:1023–33. https://doi.org/10.2147/tcrm.s3285.
Jiang F, Deng L, Zhang L, Cai Y, Cheung CW, Xia Z. Review of the clinical characteristics of coronavirus disease 2019 (COVID-19). J Gen Intern Med. 2020. https://doi.org/10.1007/s11606-020-05762-w.
Wagstaff KM, Sivakumaran H, Heaton SM, Harrich D, Jans DA. Ivermectin is a specific inhibitor of importin α/β-mediated nuclear import able to inhibit replication of HIV-1 and dengue virus. Biochem J. 2012;443:851–6. https://doi.org/10.1042/BJ20120150.
Yang SNY, Atkinson SC, Wang C, Lee A, Bogoyevitch MA, Borg NA, Jans DA. The broad spectrum antiviral ivermectin targets the host nuclear transport importin α/β1 heterodimer. Antiviral Res. 2020;177:104760. https://doi.org/10.1016/j.antiviral.2020.104760.
Caly L, Druce JD, Catton MG, Jans DA, Wagstaff KM. The FDA-approved drug ivermectin inhibits the replication of SARS-CoV-2 in vitro. Antiviral Res. 2020;178:104787. https://doi.org/10.1016/j.antiviral.2020.104787.
Balayan T, Horvath H, Rutherford GW. Ritonavir-boosted Darunavir plus two nucleoside reverse transcriptase inhibitors versus other regimens for initial antiretroviral therapy for people with hiv infection: a systematic review. AIDS Res Treat. 2017;2017:1–9. https://doi.org/10.1155/2017/2345617.
Harrison C. Coronavirus puts drug repurposing on the fast track. Nat Biotechnol. 2020;38:379–81. https://doi.org/10.1038/d41587-020-00003-1.
Khan A, Benthin C, Zeno B, Albertson TE, Boyd J, Christie JD, Hall R, Poirier G, Ronco JJ, Tidswell M, Hardes K, Powley WM, Wright TJ, Siederer SK, Fairman DA, Lipson DA, Bayliffe AI, Lazaar AL. A pilot clinical trial of recombinant human angiotensin-converting enzyme 2 in acute respiratory distress syndrome. Crit Care. 2017;21:234. https://doi.org/10.1186/s13054-017-1823-x.
Kadam RU, Wilson IA. Structural basis of influenza virus fusion inhibition by the antiviral drug Arbidol. Proc Natl Acad Sci. 2017;114:206–14. https://doi.org/10.1073/pnas.1617020114.
Chen C, Zhang Y, Huang J, Yin P, Cheng Z, Wu J, Chen S, Zhang Y, Chen B, Lu M, Luo Y, Ju L, Zhang J, Wang X. Favipiravir versus arbidol for COVID-19: a randomized clinical trial. Infect Dis (except HIV/AIDS). 2020. https://doi.org/10.1101/2020.03.17.20037432.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interests
Authors declare that they have no conflict of interest to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Rathi, H., Burman, V., Datta, S.K. et al. Review on COVID-19 Etiopathogenesis, Clinical Presentation and Treatment Available with Emphasis on ACE2. Ind J Clin Biochem 36, 3–22 (2021). https://doi.org/10.1007/s12291-020-00953-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12291-020-00953-y