Abstract
Purpose
Papillary thyroid cancers (PTCs) are the most common type of thyroid cancers, in which BRAFV600E is the most prevalent driver mutation. It is known that BRAFV600E-positive PTCs are clinically and molecularly heterogenous in terms of aggressiveness and prognosis. The molecular mechanisms of this heterogeneity were evaluated.
Methods
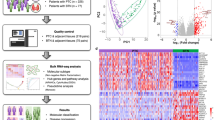
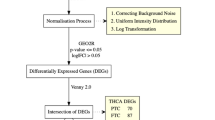
The publicly available RNA-seq data for 26 classical (c) and 5 follicular variant (fv) PTCs with BRAFV600E mutation and the BRAF-like expression signature in the BRAFV600E-RAS score (BRS) and their respective normal adjacent tissues were downloaded, and analyzed for differentially expressed genes (DEGs). The DEGs were then analyzed with the Gene Ontology annotation and the KEGG pathway dataset.
Results
We found four lines of evidence for heterogeneity of cPTCs. First, the cluster dendrogram and principle component analyses could not completely distinguish the cancer tissues from normal tissues. Second, the DEGs identified in each sample were highly diverse from one another. Third, although the DEGs were enriched in many terms containing the word “extracellular” (“extracellular region”, “extracellular space”, and so on) when analyzed as groups, the degree of this enrichment was variable when analyzed individually. Fourth, there are only a few intersections in the over-/underexpressed genes annotated with the terms containing the word “extracellular” among the samples examined. Essentially same results were obtained with BRAF-like, fvPTCs with BRAFV600E. Nevertheless, some frequently over-/underexpressed genes were detected, of which LIPH (lipase H) expression was found to be prognostic and its high expression was favorable for PTCs.
Conclusion
Groups of BRAF-like, BRAFV600E-positive cPTCs and fvPTCs that are homogenous in regard to histopathology, driver mutation and BRS were found to be highly heterogenous in terms of gene expression patterns. Yet, among the genes that were annotated with the terms containing the word “extracellular” and frequently over-/underexpressed, LIPH is a favorable prognostic marker for PTCs.

Similar content being viewed by others
References
T. Carling, R. Udelsman, Thyroid cancer. Annu Rev. Med 65, 125–137 (2014). https://doi.org/10.1146/annurev-med-061512-105739
D. Tumino, F. Frasca, K. Newbold, Updates on the management of advanced, metastatic, and radioiodine refractory differentiated thyroid cancer. Front. Endocrinol. 8, 312 (2017). https://doi.org/10.3389/fendo.2017.00312
B.R. Haugen, E.K. Alexander, K.C. Bible, G.M. Doherty, S.J. Mandel, Y.E. Nikiforov, F. Pacini, G.W. Randolph, A.M. Sawka, M. Schlumberger, K.G. Schuff, S.I. Sherman, J.A. Sosa, D.L. Steward, R.M. Tuttle, L. Wartofsky, 2015 American Thyroid Association Management guidelines for adult patients with thyroid nodules and differentiated thyroid cancer: The American Thyroid Association Guidelines Task Force on Thyroid Nodules and Differentiated Thyroid Cancer. Thyroid 26(1), 1–133 (2016). https://doi.org/10.1089/thy.2015.0020
The Cancer Genome Atlas Research, N., Integrated genomic characterization of papillary thyroid carcinoma. Cell 159(3), 676–690 (2014). https://doi.org/10.1016/j.cell.2014.09.050
G. Riesco-Eizaguirre, P. Santisteban, Endocrine tumours: advances in the molecular pathogenesis of thyroid cancer: lessons from the cancer genome. Eur. J. Endocrinol. 175(5), R203–R217 (2016). https://doi.org/10.1530/eje-16-0202
K. Kim, S. Jeon, T. M. Kim, C. K. Jung, Immune gene signature delineates a subclass of papillary thyroid cancer with unfavorable clinical outcomes. Cancers (Basel) 10(12) (2018). https://doi.org/10.3390/cancers10120494
S.K. Yoo, S. Lee, S.J. Kim, H.G. Jee, B.A. Kim, H. Cho, Y.S. Song, S.W. Cho, J.K. Won, J.Y. Shin, J. Park do, J.I. Kim, K.E. Lee, Y.J. Park, J.S. Seo, Comprehensive analysis of the transcriptional and mutational landscape of follicular and papillary thyroid cancers. PLoS Genet 12(8), e1006239 (2016). https://doi.org/10.1371/journal.pgen.1006239
S.W. Wingett, S. Andrews, FastQ Screen: a tool for multi-genome mapping and quality control. F1000Res 7, 1338 (2018). https://doi.org/10.12688/f1000research.15931.2
D. Kim, B. Langmead, S.L. Salzberg, HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12(4), 357–360 (2015). https://doi.org/10.1038/nmeth.3317
Y. Liao, G.K. Smyth, W. Shi, featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30(7), 923–930 (2014). https://doi.org/10.1093/bioinformatics/btt656
M.D. Robinson, D.J. McCarthy, G.K. Smyth, edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26(1), 139–140 (2010). https://doi.org/10.1093/bioinformatics/btp616
J. Han, M. Chen, Y. Wang, B. Gong, T. Zhuang, L. Liang, H. Qiao, Identification of biomarkers based on differentially expressed genes in papillary thyroid carcinoma. Sci. Rep. 8(1), 9912 (2018). https://doi.org/10.1038/s41598-018-28299-9
Y. Huang, M. Prasad, W.J. Lemon, H. Hampel, F.A. Wright, K. Kornacker, V. LiVolsi, W. Frankel, R.T. Kloos, C. Eng, N.S. Pellegata, A. de la Chapelle, Gene expression in papillary thyroid carcinoma reveals highly consistent profiles. Proc. Natl. Acad. Sci. USA 98(26), 15044–15049 (2001). https://doi.org/10.1073/pnas.251547398
Y. Xu, J. Chen, Z. Yang, L. Xu, Identification of RNA expression profiles in thyroid cancer to construct a competing endogenous RNA (ceRNA) network of mRNAs, long Noncoding RNAs (lncRNAs), and microRNAs (miRNAs). Med. Sci. Monit.: Int. Med. J. Exp. Clin. Res. 25, 1140–1154 (2019). https://doi.org/10.12659/msm.912450
I. Reyes, N. Reyes, R. Suriano, C. Iacob, N. Suslina, A. Policastro, A. Moscatello, S. Schantz, R.K. Tiwari, J. Geliebter, Gene expression profiling identifies potential molecular markers of papillary thyroid carcinoma. Cancer Biomark. 24(1), 71–83 (2019). https://doi.org/10.3233/cbm-181758
L. Liu, C. He, Q. Zhou, G. Wang, Z. Lv, J. Liu, Identification of key genes and pathways of thyroid cancer by integrated bioinformatics analysis. J. Cell. Physiol. (2019). https://doi.org/10.1002/jcp.28932
I. Landa, T. Ibrahimpasic, L. Boucai, R. Sinha, J.A. Knauf, R.H. Shah, S. Dogan, J.C. Ricarte-Filho, G.P. Krishnamoorthy, B. Xu, N. Schultz, M.F. Berger, C. Sander, B.S. Taylor, R. Ghossein, I. Ganly, J.A. Fagin, Genomic and transcriptomic hallmarks of poorly differentiated and anaplastic thyroid cancers. J. Clin. Investig. 126(3), 1052–1066 (2016). https://doi.org/10.1172/jci85271
O.L. Griffith, A. Melck, S.J. Jones, S.M. Wiseman, Meta-analysis and meta-review of thyroid cancer gene expression profiling studies identifies important diagnostic biomarkers. J. Clin. Oncol. 24(31), 5043–5051 (2006). https://doi.org/10.1200/jco.2006.06.7330
M.C. Barros-Filho, F.A. Marchi, C.A. Pinto, S.R. Rogatto, L.P. Kowalski, High diagnostic accuracy based on CLDN10, HMGA2, and LAMB3 transcripts in papillary thyroid carcinoma. J. Clin. Endocrinol. Metab. 100(6), E890–E899 (2015). https://doi.org/10.1210/jc.2014-4053
H.S. Kim, D.H. Kim, J.Y. Kim, N.H. Jeoung, I.K. Lee, J.G. Bong, E.D. Jung, Microarray analysis of papillary thyroid cancers in Korean. Korean J. Intern. Med. 25(4), 399–407 (2010). https://doi.org/10.3904/kjim.2010.25.4.399
Q.X. Wang, E.D. Chen, Y.F. Cai, Q. Li, Y.X. Jin, W.X. Jin, Y.H. Wang, Z.C. Zheng, L. Xue, O.C. Wang, X.H. Zhang, A panel of four genes accurately differentiates benign from malignant thyroid nodules. J. Exp. Clin. Cancer Res. 35(1), 169 (2016). https://doi.org/10.1186/s13046-016-0447-3
S.P. Cheng, M.J. Chen, M.N. Chien, C.H. Lin, J.J. Lee, C.L. Liu, Overexpression of teneurin transmembrane protein 1 is a potential marker of disease progression in papillary thyroid carcinoma. Clin. Exp. Med. 17(4), 555–564 (2017). https://doi.org/10.1007/s10238-016-0445-y
D. Luo, H. Chen, P. Lu, X. Li, M. Long, X. Peng, M. Huang, K. Huang, S. Lin, L. Tan, Y. Zhu, Z. Chen, N. Ouyang, H. Li, CHI3L1 overexpression is associated with metastasis and is an indicator of poor prognosis in papillary thyroid carcinoma. Cancer Biomark. 18(3), 273–284 (2017). https://doi.org/10.3233/cbm-160255
Y. Zhang, Z. Zhang, J. Ma, J. Pu, P. Hou, Q. Yang, High-accuracy detection of preoperative thyroid nodules using combination of BRAF(V600E) mutation and TMPRSS4 mRNA level. Arch. Med. Res. 49(6), 365–372 (2018). https://doi.org/10.1016/j.arcmed.2018.11.003
J. Tang, D. Kong, Q. Cui, K. Wang, D. Zhang, Q. Yuan, X. Liao, Y. Gong, G. Wu, Bioinformatic analysis and identification of potential prognostic microRNAs and mRNAs in thyroid cancer. PeerJ 6, e4674 (2018). https://doi.org/10.7717/peerj.4674
H. Ishimine, R. Zhou, K. Sumitomo, Y. Ito, Y. Seki, Y. Yoshida, A. Kurisaki, Lipase member H frequently overexpressed in human esophageal adenocarcinomas. Tumour Biol. 37(2), 2075–2081 (2016). https://doi.org/10.1007/s13277-015-3985-y
Y. Li, X. Zhou, Q. Zhang, E. Chen, Y. Sun, D. Ye, O. Wang, X. Zhang, J. Lyu, Lipase member H is a downstream molecular target of hypoxia inducible factor-1alpha and promotes papillary thyroid carcinoma cell migration in BCPAP and KTC-1 cell lines. Cancer Manag. Res. 11, 931–941 (2019). https://doi.org/10.2147/cmar.S183355
M. Sponziello, F. Rosignolo, M. Celano, V. Maggisano, V. Pecce, R.F. De Rose, G.E. Lombardo, C. Durante, S. Filetti, G. Damante, D. Russo, S. Bulotta, Fibronectin-1 expression is increased in aggressive thyroid cancer and favors the migration and invasion of cancer cells. Mol. Cell. Endocrinol. 431, 123–132 (2016). https://doi.org/10.1016/j.mce.2016.05.007
Y. Jin, W. Jin, Z. Zheng, E. Chen, Q. Wang, Y. Wang, O. Wang, X. Zhang, GABRB2 plays an important role in the lymph node metastasis of papillary thyroid cancer. Biochem. Biophys. Res. Commun. 492(3), 323–330 (2017). https://doi.org/10.1016/j.bbrc.2017.08.114
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Nagayama, Y., Mishima, H. Heterogenous nature of gene expression patterns in BRAF-like papillary thyroid carcinomas with BRAFV600E. Endocrine 66, 607–613 (2019). https://doi.org/10.1007/s12020-019-02063-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12020-019-02063-z