Abstract
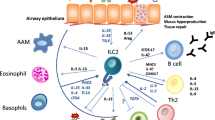
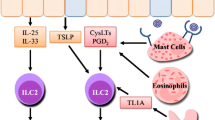
Asthma is a chronic immunological disease affecting all age groups, but often starting in childhood. Although it has long been ascribed to a single pathology, recent studies have highlighted its heterogeneity due to the potential involvement of various pathogenic mechanisms. Here, we present our current understanding of the role of innate-like T (ILT) cells in asthma pathogenesis. These cells constitute a specific family mainly comprising γδT, invariant natural killer (iNKT) and mucosal-associated invariant (MAIT) T cells. They all share the ability to massively secrete a wide range of cytokines in a T-cell receptor (TCR)-dependent or -independent manner. ILT cells are prevalent in mucosal tissues, including airways, where their innate and adaptive immune functions consist primarily in protecting tissue integrity. However, ILT cells may also have detrimental effects leading to asthma symptoms. The immune mechanisms through which this pathogenic effect occurs will be discussed in this overview.



Similar content being viewed by others
References
Reddel HK, FitzGerald JM, Bateman ED, Bacharier LB, Becker A, Brusselle G, Buhl R, Cruz AA, Fleming L, Inoue H, Ko FW, Krishnan JA, Levy ML, Lin J, Pedersen SE, Sheikh A, Yorgancioglu A, Boulet LP (2019) GINA 2019: a fundamental change in asthma management: treatment of asthma with short-acting bronchodilators alone is no longer recommended for adults and adolescents. Eur Respir J 53(6). https://doi.org/10.1183/13993003.01046-2019
Lötvall J, Akdis CA, Bacharier LB, Bjermer L, Casale TB, Custovic A, Lemanske RF, Wardlaw AJ, Wenzel SE, Greenberger PA (2011) Asthma endotypes: a new approach to classification of disease entities within the asthma syndrome. J Allergy Clin Immunol 127(2):355–360. https://doi.org/10.1016/j.jaci.2010.11.037
Kaur R, Chupp G (2019) Phenotypes and endotypes of adult asthma: moving toward precision medicine. J Allergy Clin Immunol 144(1):1–12. https://doi.org/10.1016/j.jaci.2019.05.031
Pavord ID, Beasley R, Agusti A, Anderson GP, Bel E, Brusselle G, Cullinan P, Custovic A, Ducharme FM, Fahy JV, Frey U, Gibson P, Heaney LG, Holt PG, Humbert M, Lloyd CM, Marks G, Martinez FD, Sly PD, von Mutius E, Wenzel S, Zar HJ, Bush A (2018) After asthma: redefining airways diseases. Lancet 391(10118):350–400. https://doi.org/10.1016/S0140-6736(17)30879-6
Wenzel SE (2006) Asthma: defining of the persistent adult phenotypes. Lancet 368(9537):804–813. https://doi.org/10.1016/S0140-6736(06)69290-8
Wenzel SE, Schwartz LB, Langmack EL, Halliday JL, Trudeau JB, Gibbs RL, Chu HW (1999) Evidence that severe asthma can be divided pathologically into two inflammatory subtypes with distinct physiologic and clinical characteristics. Am J Respir Crit Care Med 160(3):1001–1008. https://doi.org/10.1164/ajrccm.160.3.9812110
Wilson RH, Whitehead GS, Nakano H, Free ME, Kolls JK, Cook DN (2009) Allergic sensitization through the airway primes Th17-dependent neutrophilia and airway hyperresponsiveness. Am J Respir Crit Care Med 180(8):720–730. https://doi.org/10.1164/rccm.200904-0573OC
Bullens DM, Truyen E, Coteur L, Dilissen E, Hellings PW, Dupont LJ, Ceuppens JL (2006) IL-17 mRNA in sputum of asthmatic patients: linking T cell driven inflammation and granulocytic influx? Respir Res 7:135. https://doi.org/10.1186/1465-9921-7-135
Chesné J, Braza F, Chadeuf G, Mahay G, Cheminant MA, Loy J, Brouard S, Sauzeau V, Loirand G, Magnan A (2015) Prime role of IL-17A in neutrophilia and airway smooth muscle contraction in a house dust mite-induced allergic asthma model. J Allergy Clin Immunol 135(6):1643-1643.e1643. https://doi.org/10.1016/j.jaci.2014.12.1872
Lindén A (2001) Role of interleukin-17 and the neutrophil in asthma. Int Arch Allergy Immunol 126(3):179–184. https://doi.org/10.1159/000049511
Evasovic JM, Singer CA (2019) Regulation of IL-17A and implications for TGF-β1 comodulation of airway smooth muscle remodeling in severe asthma. Am J Physiol Lung Cell Mol Physiol 316(5):L843–L868. https://doi.org/10.1152/ajplung.00416.2018
Agache I (2019) Severe asthma phenotypes and endotypes. Semin Immunol:101301. https://doi.org/10.1016/j.smim.2019.101301
Choy DF, Hart KM, Borthwick LA, Shikotra A, Nagarkar DR, Siddiqui S, Jia G, Ohri CM, Doran E, Vannella KM, Butler CA, Hargadon B, Sciurba JC, Gieseck RL, Thompson RW, White S, Abbas AR, Jackman J, Wu LC, Egen JG, Heaney LG, Ramalingam TR, Arron JR, Wynn TA, Bradding P (2015) TH2 and TH17 inflammatory pathways are reciprocally regulated in asthma. Sci Transl Med 7(301):301ra129. https://doi.org/10.1126/scitranslmed.aab3142
Godfrey DI, Uldrich AP, McCluskey J, Rossjohn J, Moody DB (2015) The burgeoning family of unconventional T cells. Nat Immunol 16(11):1114–1123. https://doi.org/10.1038/ni.3298
Salio M, Silk JD, Jones EY, Cerundolo V (2014) Biology of CD1- and MR1-restricted T cells. Annu Rev Immunol 32:323–366. https://doi.org/10.1146/annurev-immunol-032713-120243
Crosby CM, Kronenberg M (2018) Tissue-specific functions of invariant natural killer T cells. Nat Rev Immunol 18(9):559–574. https://doi.org/10.1038/s41577-018-0034-2
Lantz O, Bendelac A (1994) An invariant T cell receptor alpha chain is used by a unique subset of major histocompatibility complex class I-specific CD4+ and CD4-8- T cells in mice and humans. J Exp Med 180(3):1097–1106
Porcelli S, Yockey CE, Brenner MB, Balk SP (1993) Analysis of T cell antigen receptor (TCR) expression by human peripheral blood CD4-8- alpha/beta T cells demonstrates preferential use of several V beta genes and an invariant TCR alpha chain. J Exp Med 178(1):1–16. https://doi.org/10.1084/jem.178.1.1
Tilloy F, Treiner E, Park SH, Garcia C, Lemonnier F, de la Salle H, Bendelac A, Bonneville M, Lantz O (1999) An invariant T cell receptor alpha chain defines a novel TAP-independent major histocompatibility complex class Ib-restricted alpha/beta T cell subpopulation in mammals. J Exp Med 189(12):1907–1921. https://doi.org/10.1084/jem.189.12.1907
Treiner E, Duban L, Bahram S, Radosavljevic M, Wanner V, Tilloy F, Affaticati P, Gilfillan S, Lantz O (2003) Selection of evolutionarily conserved mucosal-associated invariant T cells by MR1. Nature 422(6928):164–169. https://doi.org/10.1038/nature01433
Kalyan S, Kabelitz D (2013) Defining the nature of human γδ T cells: a biographical sketch of the highly empathetic. Cell Mol Immunol 10(1):21–29. https://doi.org/10.1038/cmi.2012.44
Nielsen MM, Witherden DA, Havran WL (2017) γδ T cells in homeostasis and host defence of epithelial barrier tissues. Nat Rev Immunol 17(12):733–745. https://doi.org/10.1038/nri.2017.101
Hayday AC (2009) Gammadelta T cells and the lymphoid stress-surveillance response. Immunity 31(2):184–196. https://doi.org/10.1016/j.immuni.2009.08.006
Hayday A, Tigelaar R (2003) Immunoregulation in the tissues by gammadelta T cells. Nat Rev Immunol 3(3):233–242. https://doi.org/10.1038/nri1030
Bonneville M, O'Brien RL, Born WK (2010) Gammadelta T cell effector functions: a blend of innate programming and acquired plasticity. Nat Rev Immunol 10(7):467–478. https://doi.org/10.1038/nri2781
Adams EJ, Gu S, Luoma AM (2015) Human gamma delta T cells: evolution and ligand recognition. Cell Immunol 296(1):31–40. https://doi.org/10.1016/j.cellimm.2015.04.008
Wu YL, Ding YP, Tanaka Y, Shen LW, Wei CH, Minato N, Zhang W (2014) γδ T cells and their potential for immunotherapy. Int J Biol Sci 10(2):119–135. https://doi.org/10.7150/ijbs.7823
Gu S, Nawrocka W, Adams EJ (2014) Sensing of pyrophosphate metabolites by Vγ9Vδ2 T cells. Front Immunol 5:688. https://doi.org/10.3389/fimmu.2014.00688
Vantourout P, Laing A, Woodward MJ, Zlatareva I, Apolonia L, Jones AW, Snijders AP, Malim MH, Hayday AC (2018) Heteromeric interactions regulate butyrophilin (BTN) and BTN-like molecules governing γδ T cell biology. Proc Natl Acad Sci U S A 115(5):1039–1044. https://doi.org/10.1073/pnas.1701237115
Ribot JC, deBarros A, Pang DJ, Neves JF, Peperzak V, Roberts SJ, Girardi M, Borst J, Hayday AC, Pennington DJ, Silva-Santos B (2009) CD27 is a thymic determinant of the balance between interferon-gamma- and interleukin 17-producing gammadelta T cell subsets. Nat Immunol 10(4):427–436. https://doi.org/10.1038/ni.1717
Chen L, He W, Kim ST, Tao J, Gao Y, Chi H, Intlekofer AM, Harvey B, Reiner SL, Yin Z, Flavell RA, Craft J (2007) Epigenetic and transcriptional programs lead to default IFN-gamma production by gammadelta T cells. J Immunol 178(5):2730–2736
He W, Hao J, Dong S, Gao Y, Tao J, Chi H, Flavell R, O'Brien RL, Born WK, Craft J, Han J, Wang P, Zhao L, Wu J, Yin Z (2010) Naturally activated V gamma 4 gamma delta T cells play a protective role in tumor immunity through expression of eomesodermin. J Immunol 185(1):126–133. https://doi.org/10.4049/jimmunol.0903767
Gao Y, Yang W, Pan M, Scully E, Girardi M, Augenlicht LH, Craft J, Yin Z (2003) Gamma delta T cells provide an early source of interferon gamma in tumor immunity. J Exp Med 198(3):433–442. https://doi.org/10.1084/jem.20030584
Girardi M, Oppenheim DE, Steele CR, Lewis JM, Glusac E, Filler R, Hobby P, Sutton B, Tigelaar RE, Hayday AC (2001) Regulation of cutaneous malignancy by gammadelta T cells. Science 294(5542):605–609. https://doi.org/10.1126/science.1063916
Wu D, Yan WM, Wang HW, Huang D, Luo XP, Ning Q (2018) γδ T cells contribute to the outcome of murine fulminant viral hepatitis via effector cytokines TNF-α and IFN-γ. Curr Med Sci 38(4):648–655. https://doi.org/10.1007/s11596-018-1926-x
Haas JD, González FH, Schmitz S, Chennupati V, Föhse L, Kremmer E, Förster R, Prinz I (2009) CCR6 and NK1.1 distinguish between IL-17A and IFN-gamma-producing gammadelta effector T cells. Eur J Immunol 39(12):3488–3497. https://doi.org/10.1002/eji.200939922
Puel A, Cypowyj S, Bustamante J, Wright JF, Liu L, Lim HK, Migaud M, Israel L, Chrabieh M, Audry M, Gumbleton M, Toulon A, Bodemer C, El-Baghdadi J, Whitters M, Paradis T, Brooks J, Collins M, Wolfman NM, Al-Muhsen S, Galicchio M, Abel L, Picard C, Casanova JL (2011) Chronic mucocutaneous candidiasis in humans with inborn errors of interleukin-17 immunity. Science 332(6025):65–68. https://doi.org/10.1126/science.1200439
Conti HR, Peterson AC, Brane L, Huppler AR, Hernández-Santos N, Whibley N, Garg AV, Simpson-Abelson MR, Gibson GA, Mamo AJ, Osborne LC, Bishu S, Ghilardi N, Siebenlist U, Watkins SC, Artis D, McGeachy MJ, Gaffen SL (2014) Oral-resident natural Th17 cells and γδ T cells control opportunistic Candida albicans infections. J Exp Med 211(10):2075–2084. https://doi.org/10.1084/jem.20130877
Cho JS, Pietras EM, Garcia NC, Ramos RI, Farzam DM, Monroe HR, Magorien JE, Blauvelt A, Kolls JK, Cheung AL, Cheng G, Modlin RL, Miller LS (2010) IL-17 is essential for host defense against cutaneous Staphylococcus aureus infection in mice. J Clin Invest 120(5):1762–1773. https://doi.org/10.1172/JCI40891
Sumaria N, Roediger B, Ng LG, Qin J, Pinto R, Cavanagh LL, Shklovskaya E, Fazekas de St Groth B, Triccas JA, Weninger W (2011) Cutaneous immunosurveillance by self-renewing dermal gammadelta T cells. J Exp Med 208(3):505–518. https://doi.org/10.1084/jem.20101824
Papotto PH, Ribot JC, Silva-Santos B (2017) IL-17. Nat Immunol 18(6):604–611. https://doi.org/10.1038/ni.3726
Papotto PH, Reinhardt A, Prinz I, Silva-Santos B (2018) Innately versatile: γδ17 T cells in inflammatory and autoimmune diseases. J Autoimmun 87:26–37. https://doi.org/10.1016/j.jaut.2017.11.006
Rei M, Gonçalves-Sousa N, Lança T, Thompson RG, Mensurado S, Balkwill FR, Kulbe H, Pennington DJ, Silva-Santos B (2014) Murine CD27(−) Vγ6(+) γδ T cells producing IL-17A promote ovarian cancer growth via mobilization of protumor small peritoneal macrophages. Proc Natl Acad Sci U S A 111(34):E3562–E3570. https://doi.org/10.1073/pnas.1403424111
Zuany-Amorim C, Ruffié C, Hailé S, Vargaftig BB, Pereira P, Pretolani M (1998) Requirement for gammadelta T cells in allergic airway inflammation. Science 280(5367):1265–1267
Lahn M, Kanehiro A, Takeda K, Joetham A, Schwarze J, Köhler G, O'Brien R, Gelfand EW, Born W, Kanehio A (1999) Negative regulation of airway responsiveness that is dependent on gammadelta T cells and independent of alphabeta T cells. Nat Med 5(10):1150–1156. https://doi.org/10.1038/13476
Gueders MM, Paulissen G, Crahay C, Quesada-Calvo F, Hacha J, Van Hove C, Tournoy K, Louis R, Foidart JM, Noël A, Cataldo DD (2009) Mouse models of asthma: a comparison between C57BL/6 and BALB/c strains regarding bronchial responsiveness, inflammation, and cytokine production. Inflamm Res 58(12):845–854. https://doi.org/10.1007/s00011-009-0054-2
Murdoch JR, Lloyd CM (2010) Resolution of allergic airway inflammation and airway hyperreactivity is mediated by IL-17-producing {gamma}{delta}T cells. Am J Respir Crit Care Med 182(4):464–476. https://doi.org/10.1164/rccm.200911-1775OC
Koenecke C, Chennupati V, Schmitz S, Malissen B, Förster R, Prinz I (2009) In vivo application of mAb directed against the gammadelta TCR does not deplete but generates “invisible” gammadelta T cells. Eur J Immunol 39(2):372–379. https://doi.org/10.1002/eji.200838741
Schramm CM, Puddington L, Yiamouyiannis CA, Lingenheld EG, Whiteley HE, Wolyniec WW, Noonan TC, Thrall RS (2000) Proinflammatory roles of T-cell receptor (TCR)gammadelta and TCRalphabeta lymphocytes in a murine model of asthma. Am J Respir Cell Mol Biol 22(2):218–225. https://doi.org/10.1165/ajrcmb.22.2.3620
Huang Y, Jin N, Roark CL, Aydintug MK, Wands JM, Huang H, O'Brien RL, Born WK (2009) The influence of IgE-enhancing and IgE-suppressive gammadelta T cells changes with exposure to inhaled ovalbumin. J Immunol 183(2):849–855. https://doi.org/10.4049/jimmunol.0804104
Hahn YS, Taube C, Jin N, Sharp L, Wands JM, Aydintug MK, Lahn M, Huber SA, O'Brien RL, Gelfand EW, Born WK (2004) Different potentials of gamma delta T cell subsets in regulating airway responsiveness: V gamma 1+ cells, but not V gamma 4+ cells, promote airway hyperreactivity, Th2 cytokines, and airway inflammation. J Immunol 172(5):2894–2902
Belkadi A, Dietrich C, Machavoine F, Victor JR, Leite-de-Moraes M (2018) γδ T cells amplify Blomia tropicalis-induced allergic airway disease. Allergy. https://doi.org/10.1111/all.13618
Huang Y, Aydintug MK, Loomis J, Macleod MK, McKee AS, Kirchenbaum G, Jakubzick CV, Kedl RM, Sun D, Jacobelli J, O'Brien RL, Born WK (2013) Antigen-specific regulation of IgE antibodies by non-antigen-specific γδ T cells. J Immunol 190(3):913–921. https://doi.org/10.4049/jimmunol.1202230
de Oliveira MG, de Lima Lira AA, da Ressureição SF, Inoue AHS, Santos LS, Nakamatsu BY, Duarte AJDS, Leite-de-Moraes M, Victor JR (2019) Maternal IgG impairs the maturation of offspring intrathymic IL-17-producing γδT cells: implications for murine and human allergies. Clin Exp Allergy. https://doi.org/10.1111/cea.13393
Krejsek J, Král B, Vokurková D, Derner V, Tousková M, Paráková Z, Kopecký O (1998) Decreased peripheral blood gamma delta T cells in patients with bronchial asthma. Allergy 53(1):73–77. https://doi.org/10.1111/j.1398-9995.1998.tb03776.x
Zhao Y, Yang J, Gao YD (2011) Altered expressions of helper T cell (Th)1, Th2, and Th17 cytokines in CD8(+) and γδ T cells in patients with allergic asthma. J Asthma 48(5):429–436. https://doi.org/10.3109/02770903.2011.570403
Spinozzi F, Agea E, Bistoni O, Forenza N, Monaco A, Bassotti G, Nicoletti I, Riccardi C, Grignani F, Bertotto A (1996) Increased allergen-specific, steroid-sensitive gamma delta T cells in bronchoalveolar lavage fluid from patients with asthma. Ann Intern Med 124(2):223–227
Urboniene D, Babusyte A, Lötvall J, Sakalauskas R, Sitkauskiene B (2013) Distribution of γδ and other T-lymphocyte subsets in patients with chronic obstructive pulmonary disease and asthma. Respir Med 107(3):413–423. https://doi.org/10.1016/j.rmed.2012.11.012
Krug N, Erpenbeck VJ, Balke K, Petschallies J, Tschernig T, Hohlfeld JM, Fabel H (2001) Cytokine profile of bronchoalveolar lavage-derived CD4(+), CD8(+), and gammadelta T cells in people with asthma after segmental allergen challenge. Am J Respir Cell Mol Biol 25(1):125–131. https://doi.org/10.1165/ajrcmb.25.1.4194
Pellicci DG, Hammond KJ, Uldrich AP, Baxter AG, Smyth MJ, Godfrey DI (2002) A natural killer T (NKT) cell developmental pathway iInvolving a thymus-dependent NK1.1(−)CD4(+) CD1d-dependent precursor stage. J Exp Med 195(7):835–844
Benlagha K, Kyin T, Beavis A, Teyton L, Bendelac A (2002) A thymic precursor to the NK T cell lineage. Science 296(5567):553–555
Benlagha K, Wei DG, Veiga J, Teyton L, Bendelac A (2005) Characterization of the early stages of thymic NKT cell development. J Exp Med 202(4):485–492. https://doi.org/10.1084/jem.20050456
Gapin L, Matsuda JL, Surh CD, Kronenberg M (2001) NKT cells derive from double-positive thymocytes that are positively selected by CD1d. Nat Immunol 2(10):971–978. https://doi.org/10.1038/ni710
Shimamura M, Ohteki T, Beutner U, MacDonald HR (1997) Lack of directed V alpha 14-J alpha 281 rearrangements in NK1+ T cells. Eur J Immunol 27(6):1576–1579. https://doi.org/10.1002/eji.1830270638
Egawa T, Eberl G, Taniuchi I, Benlagha K, Geissmann F, Hennighausen L, Bendelac A, Littman DR (2005) Genetic evidence supporting selection of the Valpha14i NKT cell lineage from double-positive thymocyte precursors. Immunity 22(6):705–716. https://doi.org/10.1016/j.immuni.2005.03.011
Kronenberg M (2014) When less is more: T lymphocyte populations with restricted antigen receptor diversity. J Immunol 193(3):975–976. https://doi.org/10.4049/jimmunol.1401491
Godfrey DI, Stankovic S, Baxter AG (2010) Raising the NKT cell family. Nat Immunol 11(3):197–206. https://doi.org/10.1038/ni.1841
Ohteki T, MacDonald HR (1996) Stringent V beta requirement for the development of NK1.1+ T cell receptor-alpha/beta+ cells in mouse liver. J Exp Med 183(3):1277–1282. https://doi.org/10.1084/jem.183.3.1277
Gumperz JE, Miyake S, Yamamura T, Brenner MB (2002) Functionally distinct subsets of CD1d-restricted natural killer T cells revealed by CD1d tetramer staining. J Exp Med 195(5):625–636
Geissmann F, Cameron TO, Sidobre S, Manlongat N, Kronenberg M, Briskin MJ, Dustin ML, Littman DR (2005) Intravascular immune surveillance by CXCR6+ NKT cells patrolling liver sinusoids. PLoS Biol 3(4):e113. https://doi.org/10.1371/journal.pbio.0030113
Kawano T, Cui J, Koezuka Y, Toura I, Kaneko Y, Motoki K, Ueno H, Nakagawa R, Sato H, Kondo E, Koseki H, Taniguchi M (1997) CD1d-restricted and TCR-mediated activation of valpha14 NKT cells by glycosylceramides. Science 278(5343):1626–1629. https://doi.org/10.1126/science.278.5343.1626
Matsuda JL, Naidenko OV, Gapin L, Nakayama T, Taniguchi M, Wang CR, Koezuka Y, Kronenberg M (2000) Tracking the response of natural killer T cells to a glycolipid antigen using CD1d tetramers. J Exp Med 192(5):741–754. https://doi.org/10.1084/jem.192.5.741
Rossjohn J, Pellicci DG, Patel O, Gapin L, Godfrey DI (2012) Recognition of CD1d-restricted antigens by natural killer T cells. Nat Rev Immunol 12(12):845–857. https://doi.org/10.1038/nri3328
Brennan PJ, Brigl M, Brenner MB (2013) Invariant natural killer T cells: an innate activation scheme linked to diverse effector functions. Nat Rev Immunol 13(2):101–117. https://doi.org/10.1038/nri3369
Leite-de-Moraes MC, Dy M (1997) Natural killer T cells: a potent cytokine-producing cell population. Eur Cytokine Netw 8(3):229–237
Yoshimoto T, Paul WE (1994) CD4pos, NK1.1pos T cells promptly produce interleukin 4 in response to in vivo challenge with anti-CD3. J Exp Med 179(4):1285–1295. https://doi.org/10.1084/jem.179.4.1285
Bendelac A, Matzinger P, Seder RA, Paul WE, Schwartz RH (1992) Activation events during thymic selection. J Exp Med 175(3):731–742. https://doi.org/10.1084/jem.175.3.731
Arase H, Arase N, Nakagawa K, Good RA, Onoé K (1993) NK1.1+ CD4+ CD8- thymocytes with specific lymphokine secretion. Eur J Immunol 23(1):307–310. https://doi.org/10.1002/eji.1830230151
Singh N, Hong S, Scherer DC, Serizawa I, Burdin N, Kronenberg M, Koezuka Y, Van Kaer L (1999) Cutting edge: activation of NK T cells by CD1d and alpha-galactosylceramide directs conventional T cells to the acquisition of a Th2 phenotype. J Immunol 163(5):2373–2377
Brown DR, Fowell DJ, Corry DB, Wynn TA, Moskowitz NH, Cheever AW, Locksley RM, Reiner SL (1996) Beta 2-microglobulin-dependent NK1.1+ T cells are not essential for T helper cell 2 immune responses. J Exp Med 184(4):1295–1304
Korsgren M, Persson CG, Sundler F, Bjerke T, Hansson T, Chambers BJ, Hong S, Van Kaer L, Ljunggren HG, Korsgren O (1999) Natural killer cells determine development of allergen-induced eosinophilic airway inflammation in mice. J Exp Med 189(3):553–562
Zhang Y, Rogers KH, Lewis DB (1996) Beta 2-microglobulin-dependent T cells are dispensable for allergen-induced T helper 2 responses. J Exp Med 184(4):1507–1512
Das J, Eynott P, Jupp R, Bothwell A, Van Kaer L, Shi Y, Das G (2006) Natural killer T cells and CD8+ T cells are dispensable for T cell-dependent allergic airway inflammation. Nat Med 12(12):1345–1346; author reply 1347. https://doi.org/10.1038/nm1206-1345
McKnight CG, Morris SC, Perkins C, Zhu Z, Hildeman DA, Bendelac A, Finkelman FD (2017) NKT cells contribute to basal IL-4 production but are not required to induce experimental asthma. PLoS One 12(11):e0188221. https://doi.org/10.1371/journal.pone.0188221
Lisbonne M, Diem S, de Castro KA, Lefort J, Araujo LM, Hachem P, Fourneau JM, Sidobre S, Kronenberg M, Taniguchi M, Van Endert P, Dy M, Askenase P, Russo M, Vargaftig BB, Herbelin A, Leite-de-Moraes MC (2003) Cutting edge: invariant V alpha 14 NKT cells are required for allergen-induced airway inflammation and hyperreactivity in an experimental asthma model. J Immunol 171(4):1637–1641
Akbari O, Stock P, Meyer E, Kronenberg M, Sidobre S, Nakayama T, Taniguchi M, Grusby MJ, DeKruyff RH, Umetsu DT (2003) Essential role of NKT cells producing IL-4 and IL-13 in the development of allergen-induced airway hyperreactivity. Nat Med 9(5):582–588. https://doi.org/10.1038/nm851
Olszak T, An D, Zeissig S, Vera MP, Richter J, Franke A, Glickman JN, Siebert R, Baron RM, Kasper DL, Blumberg RS (2012) Microbial exposure during early life has persistent effects on natural killer T cell function. Science 336(6080):489–493. https://doi.org/10.1126/science.1219328
An D, Oh SF, Olszak T, Neves JF, Avci FY, Erturk-Hasdemir D, Lu X, Zeissig S, Blumberg RS, Kasper DL (2014) Sphingolipids from a symbiotic microbe regulate homeostasis of host intestinal natural killer T cells. Cell 156(1–2):123–133. https://doi.org/10.1016/j.cell.2013.11.042
Remot A, Descamps D, Noordine ML, Boukadiri A, Mathieu E, Robert V, Riffault S, Lambrecht B, Langella P, Hammad H, Thomas M (2017) Bacteria isolated from lung modulate asthma susceptibility in mice. ISME J 11(5):1061–1074. https://doi.org/10.1038/ismej.2016.181
Chandra S, Zhao M, Budelsky A, de Mingo PA, Day J, Fu Z, Siegel L, Smith D, Kronenberg M (2015) A new mouse strain for the analysis of invariant NKT cell function. Nat Immunol 16(8):799–800. https://doi.org/10.1038/ni.3203
Tumes D, Hirahara K, Papadopoulos M, Shinoda K, Onodera A, Kumagai J, Yip KH, Pant H, Kokubo K, Kiuchi M, Aoki A, Obata-Ninomiya K, Tokoyoda K, Endo Y, Kimura MY, Nakayama T (2019) Ezh2 controls development of natural killer T cells, which cause spontaneous asthma-like pathology. J Allergy Clin Immunol 144(2):549-560.e510. https://doi.org/10.1016/j.jaci.2019.02.024
Michel ML, Keller AC, Paget C, Fujio M, Trottein F, Savage PB, Wong CH, Schneider E, Dy M, Leite-de-Moraes MC (2007) Identification of an IL-17-producing NK1.1(neg) iNKT cell population involved in airway neutrophilia. J Exp Med 204(5):995–1001. https://doi.org/10.1084/jem.20061551
Pichavant M, Goya S, Meyer EH, Johnston RA, Kim HY, Matangkasombut P, Zhu M, Iwakura Y, Savage PB, DeKruyff RH, Shore SA, Umetsu DT (2008) Ozone exposure in a mouse model induces airway hyperreactivity that requires the presence of natural killer T cells and IL-17. J Exp Med 205(2):385–393. https://doi.org/10.1084/jem.20071507
Koh YI, Shim JU (2010) Association between sputum natural killer T cells and eosinophilic airway inflammation in human asthma. Int Arch Allergy Immunol 153(3):239–248. https://doi.org/10.1159/000314364
Shim JU, Koh YI (2014) Increased Th2-like invariant natural killer T cells in peripheral blood from patients with asthma. Allergy Asthma Immunol Res 6(5):444–448. https://doi.org/10.4168/aair.2014.6.5.444
Lezmi G, Abou Taam R, Dietrich C, Chatenoud L, de Blic J, Leite-de-Moraes M (2018) Circulating IL-17-producing mucosal-associated invariant T cells (MAIT) are associated with symptoms in children with asthma. Clin Immunol 188:7–11. https://doi.org/10.1016/j.clim.2017.11.009
Chandra S, Wingender G, Greenbaum JA, Khurana A, Gholami AM, Ganesan AP, Rosenbach M, Jaffee K, Gern JE, Wood R, O'Connor G, Sandel M, Kattan M, Bacharier L, Togias A, Horner AA, Kronenberg M (2018) Development of asthma in inner-city children: possible roles of MAIT cells and variation in the home environment. J Immunol 200(6):1995–2003. https://doi.org/10.4049/jimmunol.1701525
Akbari O, Faul JL, Hoyte EG, Berry GJ, Wahlström J, Kronenberg M, DeKruyff RH, Umetsu DT (2006) CD4+ invariant T-cell-receptor+ natural killer T cells in bronchial asthma. N Engl J Med 354(11):1117–1129. https://doi.org/10.1056/NEJMoa053614
Vijayanand P, Seumois G, Pickard C, Powell RM, Angco G, Sammut D, Gadola SD, Friedmann PS, Djukanovic R (2007) Invariant natural killer T cells in asthma and chronic obstructive pulmonary disease. N Engl J Med 356(14):1410–1422. https://doi.org/10.1056/NEJMoa064691
Thomas SY, Lilly CM, Luster AD (2006) Invariant natural killer T cells in bronchial asthma. N Engl J Med 354(24):2613–2616; author reply 2613-2616. https://doi.org/10.1056/NEJMc066189
Matangkasombut P, Marigowda G, Ervine A, Idris L, Pichavant M, Kim HY, Yasumi T, Wilson SB, DeKruyff RH, Faul JL, Israel E, Akbari O, Umetsu DT (2009) Natural killer T cells in the lungs of patients with asthma. J Allergy Clin Immunol 123(5):1181–1185. https://doi.org/10.1016/j.jaci.2009.02.013
Pham-Thi N, de Blic J, Le Bourgeois M, Dy M, Scheinmann P, Leite-de-Moraes MC (2006) Enhanced frequency of immunoregulatory invariant natural killer T cells in the airways of children with asthma. J Allergy Clin Immunol 117(1):217–218. https://doi.org/10.1016/j.jaci.2005.09.052
Kjer-Nielsen L, Patel O, Corbett AJ, Le Nours J, Meehan B, Liu L, Bhati M, Chen Z, Kostenko L, Reantragoon R, Williamson NA, Purcell AW, Dudek NL, McConville MJ, O'Hair RA, Khairallah GN, Godfrey DI, Fairlie DP, Rossjohn J, McCluskey J (2012) MR1 presents microbial vitamin B metabolites to MAIT cells. Nature 491(7426):717–723. https://doi.org/10.1038/nature11605
Kjer-Nielsen L, Corbett AJ, Chen Z, Liu L, Mak JY, Godfrey DI, Rossjohn J, Fairlie DP, McCluskey J, Eckle SB (2018) An overview on the identification of MAIT cell antigens. Immunol Cell Biol 96(6):573–587. https://doi.org/10.1111/imcb.12057
Koay HF, Gherardin NA, Enders A, Loh L, Mackay LK, Almeida CF, Russ BE, Nold-Petry CA, Nold MF, Bedoui S, Chen Z, Corbett AJ, Eckle SB, Meehan B, d'Udekem Y, Konstantinov IE, Lappas M, Liu L, Goodnow CC, Fairlie DP, Rossjohn J, Chong MM, Kedzierska K, Berzins SP, Belz GT, McCluskey J, Uldrich AP, Godfrey DI, Pellicci DG (2016) A three-stage intrathymic development pathway for the mucosal-associated invariant T cell lineage. Nat Immunol 17(11):1300–1311. https://doi.org/10.1038/ni.3565
Lee OJ, Cho YN, Kee SJ, Kim MJ, Jin HM, Lee SJ, Park KJ, Kim TJ, Lee SS, Kwon YS, Kim N, Shin MG, Shin JH, Suh SP, Ryang DW, Park YW (2014) Circulating mucosal-associated invariant T cell levels and their cytokine levels in healthy adults. Exp Gerontol 49:47–54. https://doi.org/10.1016/j.exger.2013.11.003
Lepore M, Kalinichenko A, Colone A, Paleja B, Singhal A, Tschumi A, Lee B, Poidinger M, Zolezzi F, Quagliata L, Sander P, Newell E, Bertoletti A, Terracciano L, De Libero G, Mori L (2014) Parallel T-cell cloning and deep sequencing of human MAIT cells reveal stable oligoclonal TCRbeta repertoire. Nat Commun 5:3866. https://doi.org/10.1038/ncomms4866
Kurioka A, Jahun AS, Hannaway RF, Walker LJ, Fergusson JR, Sverremark-Ekstrom E, Corbett AJ, Ussher JE, Willberg CB, Klenerman P (2017) Shared and distinct phenotypes and functions of human CD161++ Valpha7.2+ T cell subsets. Front Immunol 8:1031. https://doi.org/10.3389/fimmu.2017.01031
Rahimpour A, Koay HF, Enders A, Clanchy R, Eckle SB, Meehan B, Chen Z, Whittle B, Liu L, Fairlie DP, Goodnow CC, McCluskey J, Rossjohn J, Uldrich AP, Pellicci DG, Godfrey DI (2015) Identification of phenotypically and functionally heterogeneous mouse mucosal-associated invariant T cells using MR1 tetramers. J Exp Med 212(7):1095–1108. https://doi.org/10.1084/jem.20142110
Lantz O, Legoux F (2019) MAIT cells: programmed in the thymus to mediate immunity within tissues. Curr Opin Immunol 58:75–82. https://doi.org/10.1016/j.coi.2019.04.016
Legoux F, Gilet J, Procopio E, Echasserieau K, Bernardeau K, Lantz O (2019) Molecular mechanisms of lineage decisions in metabolite-specific T cells. Nat Immunol 20(9):1244–1255. https://doi.org/10.1038/s41590-019-0465-3
Legoux F, Bellet D, Daviaud C, El Morr Y, Darbois A, Niort K, Procopio E, Salou M, Gilet J, Ryffel B, Balvay A, Foussier A, Sarkis M, El Marjou A, Schmidt F, Rabot S, Lantz O (2019) Microbial metabolites control the thymic development of mucosal-associated invariant T cells. Science. https://doi.org/10.1126/science.aaw2719
Martin E, Treiner E, Duban L, Guerri L, Laude H, Toly C, Premel V, Devys A, Moura IC, Tilloy F, Cherif S, Vera G, Latour S, Soudais C, Lantz O (2009) Stepwise development of MAIT cells in mouse and human. PLoS Biol 7(3):e54. https://doi.org/10.1371/journal.pbio.1000054
Li C, Lu Z, Bi K, Wang K, Xu Y, Guo P, Chen Y, Zhou P, Wei Z, Jiang H, Cao Y (2019) CD4. Reprod Biol Endocrinol 17(1):78. https://doi.org/10.1186/s12958-019-0524-5
Dusseaux M, Martin E, Serriari N, Peguillet I, Premel V, Louis D, Milder M, Le Bourhis L, Soudais C, Treiner E, Lantz O (2011) Human MAIT cells are xenobiotic-resistant, tissue-targeted, CD161hi IL-17-secreting T cells. Blood 117(4):1250–1259. https://doi.org/10.1182/blood-2010-08-303339
Chen Z, Wang H, D'Souza C, Sun S, Kostenko L, Eckle SB, Meehan BS, Jackson DC, Strugnell RA, Cao H, Wang N, Fairlie DP, Liu L, Godfrey DI, Rossjohn J, McCluskey J, Corbett AJ (2017) Mucosal-associated invariant T-cell activation and accumulation after in vivo infection depends on microbial riboflavin synthesis and co-stimulatory signals. Mucosal Immunol 10(1):58–68. https://doi.org/10.1038/mi.2016.39
Ussher JE, Bilton M, Attwod E, Shadwell J, Richardson R, de Lara C, Mettke E, Kurioka A, Hansen TH, Klenerman P, Willberg CB (2014) CD161++ CD8+ T cells, including the MAIT cell subset, are specifically activated by IL-12+IL-18 in a TCR-independent manner. Eur J Immunol 44(1):195–203. https://doi.org/10.1002/eji.201343509
van Wilgenburg B, Scherwitzl I, Hutchinson EC, Leng T, Kurioka A, Kulicke C, de Lara C, Cole S, Vasanawathana S, Limpitikul W, Malasit P, Young D, Denney L, Moore MD, Fabris P, Giordani MT, Oo YH, Laidlaw SM, Dustin LB, Ho LP, Thompson FM, Ramamurthy N, Mongkolsapaya J, Willberg CB, Screaton GR, Klenerman P, consortium S-H (2016) MAIT cells are activated during human viral infections. Nat Commun 7:11653. https://doi.org/10.1038/ncomms11653
Sattler A, Dang-Heine C, Reinke P, Babel N (2015) IL-15 dependent induction of IL-18 secretion as a feedback mechanism controlling human MAIT-cell effector functions. Eur J Immunol 45(8):2286–2298. https://doi.org/10.1002/eji.201445313
Salou M, Franciszkiewicz K, Lantz O (2017) MAIT cells in infectious diseases. Curr Opin Immunol 48:7–14. https://doi.org/10.1016/j.coi.2017.07.009
Sakala IG, Kjer-Nielsen L, Eickhoff CS, Wang X, Blazevic A, Liu L, Fairlie DP, Rossjohn J, McCluskey J, Fremont DH, Hansen TH, Hoft DF (2015) Functional heterogeneity and antimycobacterial effects of mouse mucosal-associated invariant T cells specific for riboflavin metabolites. J Immunol 195(2):587–601. https://doi.org/10.4049/jimmunol.1402545
Wong EB, Gold MC, Meermeier EW, Xulu BZ, Khuzwayo S, Sullivan ZA, Mahyari E, Rogers Z, Kløverpris H, Sharma PK, Worley AH, Lalloo U, Baijnath P, Ambaram A, Naidoo L, Suleman M, Madansein R, McLaren JE, Ladell K, Miners KL, Price DA, Behar SM, Nielsen M, Kasprowicz VO, Leslie A, Bishai WR, Ndung'u T, Lewinsohn DM (2019) TRAV1-2 + CD8 + T-cells including oligoconal expansions of MAIT cells are enriched in the airways in human tuberculosis. Commun Biol 2:203. https://doi.org/10.1038/s42003-019-0442-2
Malka-Ruimy C, Ben Youssef G, Lambert M, Tourret M, Ghazarian L, Faye A, Caillat-Zucman S, Houdouin V (2019) Mucosal-associated invariant T cell levels are reduced in the peripheral blood and lungs of children with active pulmonary tuberculosis. Front Immunol 10:206. https://doi.org/10.3389/fimmu.2019.00206
Hinks TS, Zhou X, Staples KJ, Dimitrov BD, Manta A, Petrossian T, Lum PY, Smith CG, Ward JA, Howarth PH, Walls AF, Gadola SD, Djukanovic R (2015) Innate and adaptive T cells in asthmatic patients: relationship to severity and disease mechanisms. J Allergy Clin Immunol 136(2):323–333. https://doi.org/10.1016/j.jaci.2015.01.014
Lezmi G, Abou-Taam R, Garcelon N, Dietrich C, Machavoine F, Delacourt C, Adel-Patient K, Leite-de-Moraes M (2019) Evidence for a MAIT-17-high phenotype in children with severe asthma. J Allergy Clin Immunol. https://doi.org/10.1016/j.jaci.2019.08.003
Funding
This work was supported by the grant ANR-18-CE14-0011-01 SevAsthma-children, Paris, France.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Ethical Approval and Informed Consent
No approvals or informed consents were obtained, as this manuscript does not contain primary research data.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Victor, J.R., Lezmi, G. & Leite-de-Moraes, M. New Insights into Asthma Inflammation: Focus on iNKT, MAIT, and γδT Cells. Clinic Rev Allerg Immunol 59, 371–381 (2020). https://doi.org/10.1007/s12016-020-08784-8
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12016-020-08784-8