Abstract
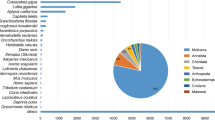
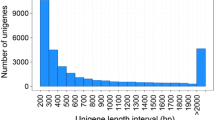
Transcriptome assemblies for the stomach, midgut, and fecal mucosa of Tachypleus tridentatus and Carcinoscorpius rotundicauda are reported for the first time in this study. Genome assembly resulted in 373069 unigenes with an N50 of 1314, while transcriptome assembly resulted in 736378 unigenes and an N50 of 2121. The annotated unigenes showed the highest similarity to fishes, amphibians, and reptiles, with most unigenes closely related to metabolism, translation, biogenesis, signal transduction, energy production, immune response, and secretion. Unigenes (4768) were mapped to KEGG pathways, and the most enriched pathways were involved in translation, environmental information signal processing, metabolism, endocrine system, immune system, nervous system, and varying metabolism. Totally 65889, 2001, and 7162 differentially expressed genes (DEGs) were identified in the stomach, midgut, and fecal mucosa between the two horseshoe crab species, respectively. Compared with C. rotundicauda, 99.95% of the DEGs in the T. tridentatus stomach were down-regulated, while 99.97% of the DEGs of T. tridentatus fecal mucosa and 55.42% of the DEGs of the T. tridentatus midgut were up-regulated. Most midgut DEGs were involved in hydrolase activity, protein metabolism, and cell cytoplasm, while most stomach and fecal mucosa DEGs were involved in catalytic activity and primary metabolic processes. Most stomach DEGs were assigned to the cellular component in cellular macromolecular complexes, while most fecal mucosa DEGs were assigned to the cellular component in cell and cell parts. These results will benefit the investigation of the molecular mechanisms of the digestive tract related to the feeding habits and environmental traces of horseshoe crabs.
Similar content being viewed by others
References
Carmichael, R. H., Gaines, E., Sheller, Z., Tong, A., Clapp, A., and Valiela, I., 2009. Diet composition of juvenile horseshoe crabs: Implications for growth and survival of natural and cultured stocks. In: Biology and Conservation of Horseshoe Crabs. Tanacredi, J. T., et al., eds., Springer, New York, 521–534, DOI: https://doi.org/10.1007/978-0-387-89959-6_33.
Chatterji, A., Vijayakumar, R., and Parulekar, A. H., 1988. Growth and morphometric characteristic in the horseshoe crab, Carcinoscorpius rotundicauda (Latreille) from Canning (West Bengal), India. Pakistan Journal of Scientific and Industrial Research, 31: 352–353.
Chen, C. P., Yang, M. C., Fan, L. F., Qiu, G., Liao, Y. Y., and Hsieh, H. L., 2015. Co-occurrence of juvenile horseshoe crabs Tachypleus tridentatus and Carcinoscorpius rotundicauda in an estuarine bay, southwestern China. Aquatic Biology, 24(2): 117–126, DOI: https://doi.org/10.3354/ab00641.
Chen, X. L., Zhu, W. L., Yang, C. L., Luo, B., Li, Q. Z., Peng, J. X., et al., 2016. Morphological and histological study on the digestive tract of Carcinoscorpius rotundicauda and Tachypleus tridentatus. Journal of Hydroecology, 37(5): 92–100, DOI: https://doi.org/10.15928/j.1674-3075.2016.05.014 (in Chinese with English abstract).
Fan, L. F., Chen, C. P., Yang, M. C., Qiu, G., Liao, Y. Y., and Hsieh, H. L., 2017. Ontogenetic changes in dietary carbon sources and trophic position of two co-occurring horseshoe crab species in southwestern China. Aquatic Biology, 26: 15–26, DOI: https://doi.org/10.3354/ab00670.
Gaines, E. F., Carmichael, R. H., Grady, S. P., and Valiela, I., 2002. Stable isotopic evidence for changing nutritional sources of juvenile horseshoe crabs. The Biological Bulletin, 203(2): 228–230, DOI: https://doi.org/10.2307/1543412.
Grabherr, M. G., Haas, B. J., Yassour, M., Levin, J. Z., Thompson, D. A., Amit, I., et al., 2011. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nature Biotechnology, 29: 644–652, DOI: https://doi.org/10.1038/nbt.1883.
Hong, S. G., 2011. Biology of Horseshoe Crabs, Tachypleus tridentatus. Xiamen University Press, Xiamen, 342pp (in Chinese).
Laurie, K., Chen, C. P., Cheung, S. G., Do, V., Hsieh, H., John, A., et al., 2019. Tachypleus tridentatus (errata version published in 2019). The IUCN Red List of Threatened Species 2019: e.T21309A149768986.
Liao, Y. Y., Xu, P. W., Kwan, K. Y., Ma, Z. Y., Fang, H. Y., Xu, J. Y., et al., 2019. Draft genomic and transcriptome resources for marine chelicerate Tachypleus tridentatus. Scientific Data, 6: 190029, DOI: https://doi.org/10.1038/sdata.2019.29.
Novitsky, T. J., 2015. Biomedical implications for managing the Limulus polyphemus harvest along the northeast coast of the United States, In: Changing Global Perspectives on Horseshoe Crab Biology, Conservation and Management. Carmichael, R. H., et al., eds., Springer, New York, 483–500, DOI: https://doi.org/10.1007/978-3-319-19542-1.
Smith, D. R., Brockmann, H. J., Beekey, M. A., King, T. L., Millard, M. J., and Zaldívar-Rae, J., 2016. Conservation status of the American horseshoe crab, (Limulus polyphemus): A regional assessment. Reviews in Fish Biology and Fisheries, 27(1): 135–175, DOI: https://doi.org/10.1007/s11160-016-9461-y.
Tanacredi, J. T., Botton, M. L., and Smith, D. R., 2009. Biology and Conservation of Horseshoe Crabs. Springer, New York, 692pp, DOI: https://doi.org/10.1007/978-0-387-89959-6.
Van Roy, P., Orr, P. J., Botting, J. P., Muir, L. A., Vinther, J., Lefebvre, B., et al., 2010. Ordovician faunas of Burgess Shale type. Nature, 465(7295): 215–218, DOI: https://doi.org/10.1038/nature09038.
Wang, C. C., Kwan, K. Y., Shin, P. K. S., Cheung, S. G., Itaya, S., Iwasaki, Y., et al., 2020. Future of Asian horseshoe crab conservation under explicit baseline gaps: A global perspective. Global Ecology and Conservation, 24: e01373, DOI: https://doi.org/10.1016/j.gecco.2020.e01373.
Wang, L., Yang, X., Zhou, S., Lyu, T., Shi, L., Dong, Y., et al., 2021. Comparative transcriptome analysis revealed omnivorous adaptation of the small intestine of Melinae. Scientific Reports, 11(1): 19162, DOI: https://doi.org/10.1038/s41598-021-98561-0.
Xie, M. J., Zhong, J. X., Xie, X. Y., Li, J. W., and Zhu, C. B., 2018. Histology and mucous cell distribution in digestive tract of Carcinoscorpius rotundicaud. Chinese Journal of Zoology, 53(2): 270–277, DOI: https://doi.org/10.13859/j.cjz.201802013.
Young, M. D., Wakefield, M. J., Smyth, G. K., and Oshlack, A., 2010. Gene ontology analysis for RNA-seq: Accounting for selection bias. Genome Biology, 11(2): R14, DOI: https://doi.org/10.1186/gb-2010-11-2-r14.
Zhang, L., Xu, B., Wu, T., Yang, Y., Fan, L., Wen, M., et al., 2017. Transcriptomic profiling of two Pak Choi varieties with contrasting anthocyanin contents provides an insight into structural and regulatory genes in anthocyanin biosynthetic pathway. BMC Genomics, 18(1): 288, DOI: https://doi.org/10.1186/s12864-017-3677-7.
Acknowledgements
This work was supported by the Scientific Research Project of Huaqiao University (No. 605-50X18005), the Guangxi BaGui Youth Scholars Program, and the Guangxi Recruitment Program of 100 Global Expert.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Li, Y., Kwan, K.Y., Li, S. et al. Transcriptome Analysis of the Digestive Tract of Tachypleus tridentatus and Carcinoscorpius rotundicauda. J. Ocean Univ. China 21, 591–600 (2022). https://doi.org/10.1007/s11802-022-5321-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11802-022-5321-7