Abstract
We model the process of cell fate determination of the flower Arabidopsis-thaliana employing a system of reaction–diffusion equations governed by a potential field. This potential field mimics the flower’s epigenetic landscape as defined by Waddington. It is derived from the underlying genetic regulatory network (GRN), which is based on detailed experimental data obtained during cell fate determination in the early stages of development of the flower. The system of equations has a variational structure, and we use minimax techniques (in particular the Mountain Pass Lemma) to show that the minimal energy solution of our functional is, in fact, the one that traverses the epigenetic landscape (the potential field) in the spatial order that corresponds to the correct architecture of the flower, that is, following the observed geometrical features of the meristem. This approach can generally be applied to systems with similar structures to establish a genotype to phenotype correspondence. From a broader perspective, this problem is related to phase transition models with a multiwell vector potential, and the results and methods presented here can potentially be applied in this case.





Similar content being viewed by others
Notes
This situation is analogous to what happens in phase transition models for a one-dimensional multiwell potential in which the spatial order of the different phases is determined by the corresponding order of the minima of the wells (see Alama and Bronsard (1997))
The genes considered are: FUL (Fruitfull), FT (Terminal flower), AP1 (Apetala 1), EMF1 (Embryonic flower 1), LFY (Leafy), AP2 (Apetala 2), WUS (Wuschel), AG (Agamous), TFL1 (Terminal flower 1), P1 (Piscilata 1), SEP (Sepallata), AP3 (Apetala 3), and UFO (Unusual flower organ) Alvarez-Buylla et al. (2010).
We do not take into account two additional attractors that are irrelevant in this work because they correspond to flower organs that are already considered (petals and stamens), and that have very few initial conditions that reach them.
SVD is a dimensional reduction technique related to the more known principal component analysis (PCA). PCA is usually explained via an eigen decomposition of the covariance matrix; however, it can also be performed via singular value decomposition (SVD) of the data matrix
The radial symmetry of the solutions can be proved analytically, using Gidas-Ni-Nirenberg’s theorem if we manage to construct an epigenetic landscape (potential field) that satisfies Lipschitz continuity at every point. This is the subject of subsequent work.
References
Alama S, Gui C, Bronsard L (1997) Stationary layered solutions in \(\mathbf{R^2}\) for an allen-cahn system with multiple well potential. Calc Var Partial Differ Equ 5(4):359–390
Allen S, Cahn J (1979) A microscopic theory for antiphase boundary motion and its application to antiphase domain coarsening. Acta Metal 27(6):1085–1095
Alvarez-Buylla E, Chaos A, Aldana M, Benítez M, Cortes-Poza Y, Espinosa-Soto C, Hartasánchez D, Lotto B, Malkin D, Escalera-Santos G, Padilla-Longoria P (2008) Floral morphogenesis: stochastic explorations of a gene network epigenetic landscape. PLoS One 3(11):e3626 e3626 e3626
Alvarez-Buylla E, Benítez M, Corvera-Poiré A, Chaos A, Folter S, Gamboa de Buen A, Garay-Arroyo A, García-Ponce B, Jaimes-Miranda F, Pérez-Ruiz R, Piñeyro-Nelson A, Sánchez-Corrales Y (2010) Flower development. Arabidopsis Book 8:2–57
Alvarez-Buylla E, Cortes-Poza Y, Padilla-Longoria P (2018) Spatial dynamics on flower organ formation. J Theor Biol 454:30–40
Bronsard L, Reitich F (1993) On 3-phase boundary motion and the singular limit of a vector valued ginzburg-landau equation. Arch Ration Mech Ana 124(4):355–379
Feldmann KA, Goff SA (2014) Chapter four - the first plant genome sequence–arabidopsis thaliana, Genomes of Herbaceous Land Plants (Andrew H. Paterson, ed.), Advances in Botanical Research, vol. 69, Academic Press, pp 91–117
Fife P (2000) Models for phase separation and their mathematics. Electron J Differ Equ 2000(48):1–26
Mendoza L, Alvarez-Buylla E (1998) Dynamics of the genetic regulatory network for arabidopsis thaliana flower morphogenesis. Theor Biol 193(2):307–319
Mullins WW (1999) Fundamental contributions to the continuum theory of evolving phase interfaces in solids ch. Two-Dimensional Motion of Idealized Grain Boundaries, Springer, Berinl, pp 70–74
Mullins W (1988) On idealiezed two dimensional motion grain growth. Scripta Metall 22(9):1441–1444
Somller J (1994) Shock waves and reaction-diffusion equations, 2nd edn. Springer-Verlag, Berlin
Turing AM (1952) The chemical basis of morphogenesis. Philos Trans Royal Soc Lond B 237(641):37–72
Van Loan CF, Golub GH (2013) Matrix computations, 4 ed., Johns Hopkins Studies in the Mathematical Sciences
Vergara-Silva F et al (2003) Inside-out flowers characteristic of lacandonia schismatica evolved at least before its divergence from a closely related taxon, triuris brevistylis. Int J Plant Sci 164(3):345–357
Waddington CH (1942) The epigenotype. Endeavour 1:18–20
Widom B, Rowlinson JS (2003) Molecular theory of capillarity, reprint. Dover, United States
Widom B, Widom H (1991) Model for line tension in three phase equilibruim. Phys A 173(1–2):72–110
Woodruff D (1976) The solid-liquid interface. Cambridge University Press, Cambridge
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendix
Appendix
1.1 Coordinate Reduction
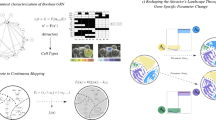
We want to interpolate the discrete dynamical system to obtain a continuous one. We shall recall that each one of the components of the vectors (Sect. 2) corresponds to a specific gene of the organ that can be in one of two different states: (0 or 1). Since there are thirteen genes in the network, when doing an interpolation, we would obtain a system in a 13-dimensional space that would be very difficult and costly to solve. For this reason, we reduce the dimensionality of the system using singular value decomposition. Through a least square adjustment, we find the bidimensional plane that best fits the four points (i.e., the one that minimizes the square of the sum of the distances of each one of the fixed points to it), and we then project the 13-dimensional points to this plane. Let \(e_1\) and \(e_2\) be two orthonormal vectors in \(\mathbb {R}^{13}\) and \(\varPi =\langle e_1,e_2 \rangle \) the plane generated by these vectors. We want to find \(e_1\) and \(e_2\) such that the sum of the distances of each vector (fixed point) \(q_1, q_2, q_3, q_4\) to the plane \(\varPi \) is the least one. We have to minimize the quantity
where \(S:=S(e_1,e_2)\), and \(d^2(q_i,\varPi )\) is the square of the distance of the vector \(q_i\) to the plane \(\varPi \). That is,
where \(Pq_i\) is the projection of the vector \(q_i\) to the plane \(\varPi \),
and \((\cdot )\) denotes the scalar product between the corresponding vectors.
To minimize S, we use singular value decomposition (SVD), which is an excellent tool when working with sets of equations of matrices that are singular or numerically nearly singular.
We start by adjusting a plane to the set of vectors \(\{q_1,q_2,q_3,q_4\}\). Let Q be the matrix of size \(m\times n\) \((m=13, n=4)\), formed by placing the vectors \(q_i\) as columns, i.e.,
Using SVD, we can decompose the matrix Q into three factors and find its singular values. Namely, we express the matrix Q as the product of three matrices: U, an orthogonal (by columns) matrix of size \(m\times n\); D, a diagonal matrix of size \(n\times n\), whose entries are greater or equal to zero; and finally, V, a transpose matrix of an orthogonal one, of size \(n\times n\). That is,
Since both U and V have orthogonal columns, then \(U^TU=V^TV=I\) and \(V\cdot V^T=I\); therefore, D will be the diagonal matrix given by:
where \(w_i\) will be the singular values of Q and \(w_i\ge 0\ \forall i\).
Having obtained the matrices U, V, and D, we will take \(e_1\) and \(e_2\) as the first and second columns of \(V^T\) (which are mutually orthogonal), respectively. The plane formed by these orthogonal vectors is the plane that minimizes the sum of the distances of each vector \(q_i\) to it.
Now we can compute the projection \(Pq_i\) of each vector \(q_i\) to the plane \(\varPi =\varPi <e_1,e_2>\),
If we take \(e_1\) and \(e_2\) as the vectors that generate, respectively, the horizontal and vertical axis of a bidimensional coordinate system, we can compute both coordinates of each vector \(q_i\) with respect to \(\{e_1,e_2\}\), using the following equation
where x and y denote the horizontal and vertical axis, respectively.
We find that a basis that generates the plane we need is given by
and
By projecting our original vectors into this plane, we obtain the following four points in \(\mathbb R^2\), each corresponding to a flower organ (see Table 2).
Note that if we wish to stay in the positive octant, it suffices to do a translation in such a way that the four initial conditions \((u_1,v_1),(u_2,v_2),(u_3,v_3)\), and \((u_4,v_4)\) are all positive.
Rights and permissions
About this article
Cite this article
Cortés-Poza, Y., Padilla-Longoria, P. A Variational Approach to Morphogenesis. Bull Math Biol 84, 33 (2022). https://doi.org/10.1007/s11538-022-00993-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11538-022-00993-w