Abstract
Prediction of the stage of cancer plays an important role in planning the course of treatment and has been largely reliant on imaging tools which do not capture molecular events that cause cancer progression. Gene-expression data–based analyses are able to identify these events, allowing RNA-sequence and microarray cancer data to be used for cancer analyses. Breast cancer is the most common cancer worldwide, and is classified into four stages — stages 1, 2, 3, and 4 [2]. While machine learning models have previously been explored to perform stage classification with limited success, multi-class stage classification has not had significant progress. There is a need for improved multi-class classification models, such as by investigating deep learning models. Gene-expression-based cancer data is characterised by the small size of available datasets, class imbalance, and high dimensionality. Class balancing methods must be applied to the dataset. Since all the genes are not necessary for stage prediction, retaining only the necessary genes can improve classification accuracy. The breast cancer samples are to be classified into 4 classes of stages 1 to 4. Invasive ductal carcinoma breast cancer samples are obtained from The Cancer Genome Atlas (TCGA) and Molecular Taxonomy of Breast Cancer International Consortium (METABRIC) datasets and combined. Two class balancing techniques are explored, synthetic minority oversampling technique (SMOTE) and SMOTE followed by random undersampling. A hybrid feature selection pipeline is proposed, with three pipelines explored involving combinations of filter and embedded feature selection methods: Pipeline 1 — minimum-redundancy maximum-relevancy (mRMR) and correlation feature selection (CFS), Pipeline 2 — mRMR, mutual information (MI) and CFS, and Pipeline 3 — mRMR and support vector machine–recursive feature elimination (SVM-RFE). The classification is done using deep learning models, namely deep neural network, convolutional neural network, recurrent neural network, a modified deep neural network, and an AutoKeras generated model. Classification performance post class-balancing and various feature selection techniques show marked improvement over classification prior to feature selection. The best multiclass classification was found to be by a deep neural network post SMOTE and random undersampling, and feature selection using mRMR and recursive feature elimination, with a Cohen-Kappa score of 0.303 and a classification accuracy of 53.1%. For binary classification into early and late-stage cancer, the best performance is obtained by a modified deep neural network (DNN) post SMOTE and random undersampling, and feature selection using mRMR and recursive feature elimination, with an accuracy of 81.0% and a Cohen-Kappa score (CKS) of 0.280. This pipeline also showed improved multiclass classification performance on neuroblastoma cancer data, with a best area under the receiver operating characteristic (auROC) curve score of 0.872, as compared to 0.71 obtained in previous work, an improvement of 22.81%. The results and analysis reveal that feature selection techniques play a vital role in gene-expression data-based classification, and the proposed hybrid feature selection pipeline improves classification performance. Multi-class classification is possible using deep learning models, though further improvement particularly in late-stage classification is necessary and should be explored further.
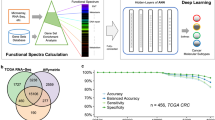
Graphical Abstract


















Similar content being viewed by others
References
Abadi M, Agarwal A, Barham P, Brevdo E, Chen Z, Citro C, Zheng X (2015) TensorFlow: Large-scale machine learning on heterogeneous systems. Retrieved July 31, 2023, from https://www.tensorflow.org/
Ahmed O, Brifcani A (2019, April) Gene expression classification based on deep learning. In 2019 4th Scientific International Conference Najaf (SICN). IEEE, pp 145–149
American Cancer Society (2021, June 28) Stages of breast cancer: Understand breast cancer staging. Retrieved October 25, 2021, from https://www.cancer.org/cancer/breast-cancer/understanding-a-breast-cancer-diagnosis/stages-of-breast-cancer.html
Barrett T, Wilhite SE, Ledoux P, Evangelista C, Kim IF, Tomashevsky M, Marshall KA, Phillippy KH, Sherman PM, Holko M, Yefanov A, Lee H, Zhang N, Robertson CL, Serova N, Davis S, Soboleva A (2013) NCBI GEO: archive for functional genomics data sets–update. Nucleic Acids Res 41:D991–D995
Castillo D, Gálvez JM, Herrera LJ, Román BS, Rojas F, Rojas I (2017) Integration of RNA-Seq data with heterogeneous microarray data for breast cancer profiling. BMC Bioinformatics 18(1):506
cBioPortal for Cancer Genomics (2016) Breast cancer (METABRIC, Nature 2012 & Nat Commune 2016). Retrieved May 25, 2022, from http://www.cbioportal.org/study/summary?id=brca/_metabric
Daoud M, Mayo M (2019) A survey of neural network-based cancer prediction models from microarray data. Artif Intell Med 97:204–214
Dertat A (2017, October 9) Applied deep learning — part 1: Artificial neural networks. Medium. Retrieved October 25, 2021, from https://towardsdatascience.com/applied-deep-learning-part-1-artificial-neural-networks-d7834f67a4f6
Ding C, Peng H (2005) Minimum redundancy feature selection from microarray gene expression data. J Bioinform Comput Biol 3(02):185–205
Fathi H, AlSalman H, Gumaei A, Manhrawy II, Hussien AG, El-Kafrawy P (2021) An efficient cancer classification model using microarray and high-dimensional data. Comput Intell Neurosci 2021
Goodfellow I, Bengio Y, Courville A (2016) Deep learning. Retrieved October 25, 2021, from https://www.deeplearningbook.org
Google Developers (2020, Feb 11) Classification: Precision and recall | Machine learning crash course. https://developers.google.com/machine-learning/crash-course/classification/precision-and-recall
Google Developers (n.d.) Classification: ROC curve and AUC. Retrieved May 25, 2022, from https://developers.google.com/machine-learning/crash-course/classification/roc-and-auc
Google (n.d.) Google Colab. Google Colaboratory. Retrieved May 25, 2022, from https://research.google.com/colaboratory/faq.html
Gosain A, Sardana S (2017) Handling class imbalance problem using oversampling techniques: A review. 2017 International Conference on Advances in Computing, Communications and Informatics (ICACCI)
Griffith M, Walker J, Spies N, Ainscough B, Griffith O (2015) Informatics for RNA sequencing: a Web resource for analysis on the cloud. Plos Comput Biol 11(8):e1004393
Hambali MA, Oladele TO, Adewole KS (2020) Microarray cancer feature selection: Review, challenges and research directions. Int J Cogn Comput Eng 1:78–97
IBM Cloud Education (2020) What is deep learning? IBM. Retrieved October 25, 2021, from https://www.ibm.com/cloud/learn/deep-learning
Jin H, Chollet F, Song Q, Hu X (2023) AutoKeras: an AutoML library for deep learning. J Mach Learn Res 6:1–6
Liang H, Zhou G, Lv L et al (2021) KRAS expression is a prognostic indicator and associated with immune infiltration in breast cancer. Breast Cancer 28:379–386. https://doi.org/10.1007/s12282-020-01170-4
Lin Z, Ou-Yang L (2023) Inferring gene regulatory networks from single-cell gene expression data via deep multi-view contrastive learning. Brief Bioinforma 24(1):bbac586. https://doi.org/10.1093/bib/bbac586
Mignone P, Pio G, D’Elia D, Ceci M (2020) Exploiting transfer learning for the reconstruction of the human gene regulatory network. Bioinformatics 36(5):1553–1561. https://doi.org/10.1093/bioinformatics/btz781
Park A, Nam S (2019) Deep learning for stage prediction in neuroblastoma using gene expression data. Genom Inform 17(3)
Pedregosa F, Varoquaux G, Gramfort A, Michel V, Thirion B, Grisel O, Duchesnay É (2011) Scikit-learn: Machine learning in python. J Mach Learn Res 12:2825–2830
Pereira B, Chin SF, Rueda O et al (2016) The somatic mutation profiles of 2,433 breast cancers refine their genomic and transcriptomic landscapes. Nat Commun 7:11479. https://doi.org/10.1038/ncomms11479
Rajbhandari P, Lopez G, Capdevila C, Salvatori B et al (2018May) Cross-cohort analysis identifies a TEAD4-MYCN positive feedback loop as the core regulatory element of high-risk neuroblastoma. Cancer Discov 8(5):582–599
Roy S, Kumar R, Mittal V, Gupta D (2020) Classification models for invasive ductal carcinoma progression, based on gene expression data-trained supervised machine learning. Sci Rep 10(1):1–15
Scitable by Nature Education (2014) Gene Expression Is Analyzed by Tracking RNA. Retrieved May 25, 2022, from https://www.nature.com/scitable/topicpage/gene-expression-is-analyzed-by-tracking-rna-6525038/
Sun L, Kong X, Xu J, Zhai R, Zhang S (2019) A hybrid gene selection method based on ReliefF and ant colony optimization algorithm for tumor classification. Sci Rep 9(1):1–14
Suzuki E, Sugimoto M, Kawaguchi K et al (2019) Gene expression profile of peripheral blood mononuclear cells may contribute to the identification and immunological classification of breast cancer patients. Breast Cancer 26:282–289. https://doi.org/10.1007/s12282-018-0920-2
The Cancer Genome Atlas Program (n.d.) National Cancer Institute. Retrieved May 25, 2022, from https://www.cancer.gov/about-nci/organization/ccg/research/structural-genomics/tcga
UICC (2022) UICC and the TNM classification of malignant tumours. UICC. Retrieved May 25, 2022, from https://www.uicc.org/who-we-are/about-uicc/uicc-and-tnm-classification-malignant-tumours
Urbanowicz RJ, Meeker M, La Cava W, Olson RS, Moore JH (2018) Relief-based feature selection: introduction and review. J Biomed Inform 85:189–203
Viera AJ, Garrett JM (2005) Understanding interobserver agreement: the kappa statistic. Fam Med 37(5):360–363
World Health Organization (2021) Breast cancer. World Health Organization. Retrieved October 25, 2021, from https://www.who.int/news-room/fact-sheets/detail/breast-cancer
Yao F, Zhang C, Du W, Liu C, Xu Y (2015) Identification of gene-expression signatures and protein markers for breast cancer grading and staging. Plos One 10(9):e0138213
Yuan F, Lu L, Zou Q (2020) Analysis of gene expression profiles of lung cancer subtypes with machine learning algorithms. Biochimica et Biophysica Acta (BBA)-Mol Basis Dis 1866(8):165822
Yang ZJ, Yu Y, Chi JR et al (2018) The combined pN stage and breast cancer subtypes in breast cancer: a better discriminator of outcome can be used to refine the 8th AJCC staging manual. Breast Cancer 25:315–324. https://doi.org/10.1007/s12282-018-0833-0
Zhong L, Meng Q, Chen Y, Du L, Wu P (2021) A laminar augmented cascading flexible neural forest model for classification of cancer subtypes based on gene expression data. BMC Bioinformatics 22(1):1–17. https://doi.org/10.1186/s12859-021-04391-2
Author information
Authors and Affiliations
Contributions
Akash Kishore: literature survey, collected data set, data preparation and implementation. Lokeswari Y Venkataramana: evaluating the implementation, updating the manuscript, and reviewing the work. D Venkata Vara Prasad: reviewed the paper, suggestions on using different variations of dataset. Akshaya Mohan: literature survey, implementation, writing the manuscript. Bhavya Jha: literature survey, implementation, prepared figures and tables.
Corresponding author
Ethics declarations
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors. All the authors have agreed to publish this manuscript. The manuscript is not submitted to any other journal or not under consideration of any journal.
Informed consent
Informed consent is not necessary as this article does not involve human or animal participants.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kishore, A., Venkataramana, L., Prasad, D.V. et al. Enhancing the prediction of IDC breast cancer staging from gene expression profiles using hybrid feature selection methods and deep learning architecture. Med Biol Eng Comput 61, 2895–2919 (2023). https://doi.org/10.1007/s11517-023-02892-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11517-023-02892-1