Abstract
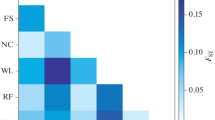
China is rich in chicken genetic resources, and many indigenous breeds can be found throughout the country. Due to poor productive ability, some of them are threatened by the commercial varieties from domestic and foreign breeding companies. In a large-scale investigation into the current status of Chinese poultry genetic resources, 78 indigenous chicken breeds were surveyed and their blood samples collected. The genomes of these chickens were screened using microsatellite analysis. A total of 2740 individuals were genotyped for 27 microsatellite markers on 13 chromosomes. The number of alleles of the 27 markers ranged from 6 to 51 per locus with a mean of 18.74. Heterozygosity (H) values of the 78 chicken breeds were all more than 0.5. The average H value (0.622) and polymorphism information content (PIC, 0.573) of these breeds suggested that the Chinese indigenous chickens possessed more genetic diversity than that reported in many other countries. The fixation coefficients of subpopulations within the total population (F ST) for the 27 loci varied from 0.065 (LEI0166) to 0.209 (MCW0078), with a mean of 0.106. For all detected microsatellite loci, only one (LEI0194) deviated from Hardy-Weinberg equilibrium (HWE) across all the populations. As genetic drift or non-random mating can occur in small populations, breeds kept on conservation farms such as Langshan chicken generally had lower H values, while those kept on large populations within conservation regions possessed higher polymorphisms. The high genetic diversity in Chinese indigenous breeds is in agreement with great phenotypic variation of these breeds. Using Nei’s genetic distance and the Neighbor-Joining method, the indigenous Chinese chickens were classified into six categories that were generally consistent with their geographic distributions. The molecular information of genetic diversity will play an important role in conservation, supervision, and utilization of the chicken resources.
Similar content being viewed by others
References
West B, Zhou B X. Did chickens go north? New evidence for domestication. J Arch Sci, 1988, 14: 515–533
Fulton J E, Delany M E. Poultry genetic resources-operation rescue needed. Science, 2003, 300: 1667–1668
Buchanan F C, Adams L J, Littlejohn R P, Maddox J F, Crawford A M. Determination of evolutionary relationships among sheep breeds using microsatellites. Genomics, 1994, 22: 397–403
MacHugh E, Lofftus R T, Barley D G, Sharp P M, Cunningham P. Microsatellite DNA variation within and among European cattle breeds. Proc Roy Soc Biol Sci, 1994, 256: 25–31
Romanov M N, Weigends S. Genetic diversity in chicken populations based on microsatellite markers. In: Dekkers J C M, Lamont S, Rothschild M F, eds. Proc. of the Conference “From Jay Lush to Genomics: Visions for Animal Breeding and Genetics”. Ames: Iowa State University, 1999. 174
Crooijmans R P M A, Groen A F, van Kampen A J A, van der Beek S, Van der Poel J J, Groenen M A M. Microsatellite polymorphism in commercial broiler and layers lines estimated using pooled blood samples. Poult Sci, 1996, 72: 334–348
Hillel J, Korol A, Kirzner V, Freidlin P, Weigend S, Barre-Dirie A, Groenen M A M, Crooijmans R P M A, Tixier-Boichard M, Vignal A, Wimmers K, Burke T, Thomson P A, Maki-Tanila A, Elo K, Zhivotovsky L A, Feldman M W. Biodiversity of chickens based on DNA pools: First results of the EC funded project AVIANDIV. In: Preisinger R, ed. Proceedings of the Poultry Genetic Symposium. Mariensee, Germany, Cuxhaven: Lohmann Tierzucht, 1999. 22–29
Hillel J, Groenen M A M, Tixier-Boichard M, Korol A B, David L, Kirzhner V M, Burke T, Barre-Dirie A, Crooijmans R P M A, Elo K, Feldman M W, Freidlin P J, Mäki-Tanila A, Oortwijn M, Thomson P, Vignal A, Wimmers K, Weigend S. Biodiversity of 52 chicken populations assessed by microsatellite typing of DNA pools. Genet Sel Evol, 2003, 35: 533–557
Weigend S, Hillel J, Groenen M A M, Tixier-Boichard M, Korol A, Kirzner V, Freidlin P, Crooijmans R P M A, Vignal A, Wimmers K, Ponsuksili S, Thomson P A, Burke T, Maki-Tanila A, Elo K, Barre-Dirie A, Zhivotovsky L A, Feldman M W. Assessment of biodiversity in a wide range of chicken breeds by genotyping DNA pools for microsatellite loci. In Proc. of the 27th International Conference on Animal Genetics. Minneapolis, USA, 2000. 83
Weigend S, Romanov M N. Current strategies for the assessment and evaluation of genetic diversity in chicken resources. World’s Poult Sci J, 2001, 57: 275–288
Romanov M N, Weigend S. Using RAPD markers for assessment of genetic diversity in chickens. Arch Geflugelkd, 2001, 65: 1–4
Zhang X, Leung F C, Chan D K, Chen Y, Wu C. Comparative analysis of allozyme, random amplified polymorphic DNA, and microsatellite polymorphism on Chinese native chickens. Poult Sci, 2002, 81: 1093–1098
Zhang X, Leung F C, Chan D K, Yang G, Wu C. Genetic diversity of Chinese native chicken breeds based on protein polymorphism, randomly amplified polymorphic DNA, and microsatellite polymorphism. Poult Sci, 2002b, 81: 1463–1472
Rosenberg N A, Burke T, Elo K, Feldman M W, Freidlin P J, Groenen M A M, Hillel J, Mäki-Tanila A, Tixier-Boichard M, Vignal A, Wimmers K, Weigend S. Empirical evaluation of genetic clustering methods using multilocus genotypes from 20 chicken breeds. Genetics, 2001, 159: 699–713
Qu L J, Li X Y, Wu G, Yang N. Efficient and sensitive method of DNA silver staining in polyacrylamide gels. Electrophoresis, 2005, 26: 99–101
Marshall T. User’s Manual for Cervus. University of Edinburgh, UK. 1998
Botstein D, White R L, Skolnick M, Davis R W. Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet, 1980, 32: 314–331
Nei M. Genetic distance between populations. Am Nat, 1972, 106: 283–293
Nei M. Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics, 1978, 89: 583–590
Du Z Q, Qu L J, Li X Y, Hu X X, Huang Y H, Li N, Yang N. Genetic Diversity in Tibetan Chicken. Hereditas, 2004, 26: 167–171
Ponsuksili S, Wimmers K, Schmoll F, Horst P, Schellander K. Comparison of multilocus DNA fingerprints and microsatellites in an estimate of genetic distance in chickens. Hered, 1999, 90: 656–659
Vanhala T, Tuiskula-Havisto M, Elo K, Vilkki J, Maki-Tanila A. Evaluation of genetic variability and genetic distances between eight chicken lines using microsatellite markers. Poult Sci, 1998, 77: 783–790
Zhou H, Lamont S J. Genetic characterization of biodiversity in highly inbred chicken lines by microsatellite markers. Anim Genet, 1999, 30: 256–264
Shriver M D, Jin L, Boerwinkle E, Deka R, Ferrell R E, Chakraborty R. A novel measure of genetic distance for highly polymorphic tandem repeat loci. Mol Biol Evol, 1995, 12: 914–920
Chu J Y, Huang W, Kuang S Q, Wang J M, Xu J J, Chu Z T, Yang Z Q, Lin K Q, Li P, Wu M, Geng Z C, Tan C C, Du R F, Jin L. Genetic relationship of populations in China. Proc Natl Acad Sci USA, 1998, 95: 11763–11768
Wu D C, Yang Z. Game chickens. In: Chinese Game Chickens. Huhhot: Yuanfang Publisher, 1993. 4–7
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Qu, L., Li, X., Xu, G. et al. Evaluation of genetic diversity in Chinese indigenous chicken breeds using microsatellite markers. SCI CHINA SER C 49, 332–341 (2006). https://doi.org/10.1007/s11427-006-2001-6
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s11427-006-2001-6