Abstract
The morphological and microscopy were combined with DNA-barcoding, together with rapid TLC for the characterization of Piper betle (PB), P. nigrum (PN), P. retrofractum (PR), P. sarmentosum (PS), and P. wallichii (PW), five medicinal Piper plants announced in the Thai Herbal Pharmacopoeia (THP). The authentic plants collected from various locations and voucher Piper products bought from commercial sites in Thailand were studied. The reproductive parts of authentic plants were subjected to ensure their morphological characters. Using sequencing analysis and genetic divergence for analyzing discriminatory performance, ITS2 was selected from eight candidate DNA markers to authenticate the origin of Piper crude drugs together with microscopic and TLC profiles for examining their characters, admixtures, adulterants, and substituents. PB and PR exhibited unique characters of the species, with no admixture, adulteration, and substitution. PN showed no variable characters of morphology and genetics. However, the microscopy could illustrate some commercial products of PN sold in Thailand have been adulterated with rice starch and roasted rice. In the herbal trade, PS has been sold in the form of mixed leaf, root, and stem more than the isolated part, but there is no variable character of the species. PW has shown more than one character of species explained by microscopic, chemical components, and genetic data. In conclusion, the conventional and molecular pharmacognostic data combined with chemical profile of authentic five Piper plants could be applied to indicate the plant origin and clarify the situations of admixture, adulteration, and substitution of the commercial Piper products launched in Thailand.
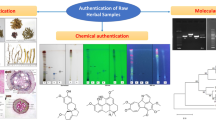
Graphical abstract








Similar content being viewed by others
References
Mabberley D (2008) Mabberley's Plant Book. A Portable Dictionary of Plants, their Classification and Uses. Cambridge University Press, USA
Suwanphakdee C, Simpson DA, Hodkinson TR, Chantaranothai P (2020) A synopsis of Thai Piper (Piperaceae). Thai Forest Bull Bot 48(2):145–183. https://doi.org/10.20531/tfb.2020.48.2.08
Suwanphakdee C, Simpson DA, Hodkinson TR, Chantaranothai P (2016) Taxonomic notes on the genus Piper (Piperaceae). Nord J Bot 34:605–618. https://doi.org/10.1111/njb.01114
Salehi B, Zakaria ZA, Gyawali R, Ibrahim SA, Rajkovic J, Shinwari ZK, Khan T, Sharifi-Rad J, Ozleyen A, Turkdonmez E, Valussi M, Tumer TB, Fidalgo LM, Martorell M, Setzer WN (2019) Piper species: a comprehensive review on their phytochemistry, biological activities and applications. Molecules 24:1364. https://doi.org/10.3390/molecules24071364
Yu N, Gu H, Wei Y, Zhu N, Wang Y, Zhang H, Zhu Y, Zhang X, Ma C, Sun A (2016) Suitable DNA barcoding for identification and supervision of Piper kadsura in Chinese medicine markets. Molecules 21:1221. https://doi.org/10.3390/molecules21091221
Ooi SL, Handerson P, Pak SC (2018) Kava for generalized anxiety disorder: a review of current evidence. J Altern Complement Med 24(8):770–780. https://doi.org/10.1089/acm.2018.0001
Egydio Brandao APM, Yamaguchi LF, Tepe EJ, Salatin A, Kato MJ (2020) Evaluation of DNA markers for molecular identification of three Piper species from Brazilian Atlantic Forest. PLoS ONE 15(10):e0239056. https://doi.org/10.1371/journal.pone.0239056
Bureau of Drug and Narcotics, Department of Medicinal Sciences, Ministry of Public Health (2020) Thai Herbal Pharmacopoeia. Nonthaburi, Thailand
Chokevivat V, Chuthaputti A (2005) The Role of Thai Traditional Medicine in Health Promotion. Bangkok, Thailand
Parmar VS, Jain SC, Gupta S, Talwar S, Rajwanshi VK, Kumar R, Azim A, Malhotra S, Kumar N, Jain R, Sharma NK, Tyagi OD, Lawrie SJ, Errington W, Howarth OW, Olsen CE, Singh SK, Wengel J (1998) Polyphenols and alkaloids from Piper species. Phytochem 49(4):1069–1078. https://doi.org/10.1016/S0031-9422(98)00208-8
Parvathy VA, Swetha VP, Sheeja TE, Leela NK, Chempakam B, Sasikumar B (2014) DNA barcoding to detect chili adulteration in traded black pepper powder. Food Biotech 28(1):25–40. https://doi.org/10.1080/08905436.2013.870078
Sarifudin A, Sholichah E, Setiaboma W, Ekafitri R, Afifah N, Ratnawati L, Pudiyanto EA, Achyadi NS (2021) Adulterated powdered white pepper products by tapioca flour sold in Indonesian’s online marker investigated by simple FTIR analytical method. J Food Nutr Res 9:297–303. https://doi.org/10.12691/jfnr-9-6-5
Dhanya K, Syamkumar S, Sasikumar B (2009) Development and application of SCAR marker for the detection of papaya seed adulteration in traded black pepper. Food Biotechnol 23:37–106. https://doi.org/10.1080/08905430902873007
Khan S, Mirza KJ, Anwar F, Abdin MZ (2010) Development of RAPD markers of authentication of Piper nigrum (L.). Environ We Int J Sci Tech 5:47–56
Menghani E, Jain SC, Jain R (2010) Scientific validation of Piper nigrum by HPLC and anti-oxidative assay markers. Asian J Biotech 2(2):133–138. https://doi.org/10.3923/ajbkr.2010.133.138
Paradkar MM, Singhal RS, Kulkarni PR (2001) A new TLC method to detect the presence of ground papaya seed in ground black pepper. J Sci Food Agric 81:1322–1325. https://doi.org/10.1002/jsfa.946
Parvathy VA, Swetha VP, Sheeja TE, Sasikumar B (2018) A two locus barcode for discriminating Piper nigrum from its related adulterant species. IJBT 17:346–350
Dhanya K, Sasikumar B (2010) Molecular marker based adulteration detection in trades food and agricultural commodities of plant origin with special reference to spices. Curr Trends Biotechnol Pharm 4:454–489
Urumarudappa SKJ, Tungphatthong C, Sukrong S (2019) Mitigating the impact of admixtures in Thai herbal products. Front Pharmacol 10:1–13. https://doi.org/10.3389/fphar.2019.01205
Tremlova B (2001) Evidence of spice black pepper adulteration. Czech J Food Sci 19:235–239
Rana CS, Kimothi GP, Sasibhushan V, Rai RK, Narayan SB, Sastry JLN (2020) Microscopical identification of two Piper spp., Piper longum and Piper retrofractum. J Res Educ Indian Med 00:00–00. https://doi.org/10.5455/JREIM.82-1568366104
Dong W, Xu C, Li C, Sun J, Zuo Y, Shi S, Cheng T, Guo J, Zhou S (2015) ycf1, the most promising plastid DNA barcode of land plants. Sci Rep 5:8348. https://doi.org/10.1186/s43141-020-00057-3
Stanford AM, Harden R, Park CR (2000) Phylogeny and biogeography of Juglans (Juglandaceae) based on matK and ITS sequence data. Am J Bot 87:872–882. https://doi.org/10.2307/2656895
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: PCR protocols: a guide to methods and application. Academic Press, New York
Kuzmina ML, Johnson KL, Barron HR, Hebert PDN (2012) Identification of the vascular plants of Churchill, Manitoba, using a DNA barcode library. BMC Ecol 12:1–11. https://doi.org/10.1186/1472-6785-12-25
Hamilton MB (1999) Four primer pairs for the amplification of chloroplast intergenic regions with intraspecific variation. Mol Ecol 8:513–525
Taberlet P, Gielly L, Pautou G, Bouvet J (1991) Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Mol Biol 17:1105–1109. https://doi.org/10.1007/BF00037152
Fay FM, Bayer C, Alverson WS, Bruijin AY, Chase MW (1998) Plastid rbcL sequence data indicate a close affinity between Diegodendron and Bixa. Taxon 47:43–50. https://doi.org/10.2307/1224017
Hollingsworth PM, Forrest LL, Spouge JL, Hajibabaei M (2009) A DNA barcode for land plants. Proc Nat Acad Sci USA 106:12794–12797. https://doi.org/10.1073/pnas.0905845106
Thompson JD, Higgins DG, Gibson TJ, Clustal W (1994) Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acid Res 22(22):4673–4680. https://doi.org/10.1093/nar/22.22.4673
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Window 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Kimura MA (1980) Simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16(2):111–120. https://doi.org/10.1007/BF01731581
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35(6):1547–1549. https://doi.org/10.1093/molbev/msy096
Somanabandhu A (1986) A Guide to the Identification and Authentication of some Thai Medicinal Plants, In: Specification of Thai Medicinal Plants Volume 1. Aksornsampan Press, Bangkok
Ondora OM, Gikonyo NK, Nyambaka HN, Thoithi GN (2020) Macroscopic and microscopic features of diagnostic value for Warburgia ugandensis Sprague leaf and stem-bark herbal materials. J Pharmacognosy Phytother 12(2):36–43. https://doi.org/10.5897/JPP2019.0569
Jaramillo MA, Callejas R, Davidson C, Smith JF, Stevens AC, Tape EJ (2008) A phylogeny of the tropical genus Piper using ITS and the chloroplast in psbJ-petA. Syst Bot 33(4):647–660. https://doi.org/10.1600/036364408786500244
Suwanphakdee C, Simpson DA, Hodkinson TR, Chantaranothai P (2018) Typification of Piper species (Piperaceae) in Southeast Asia, especially Thailand. Kew Bull 73:33. https://doi.org/10.1007/s12225-018-9761-6
Dapar MLG, Demayo CG, Meve U, Liede-Schumann S, Alejandro GJD (2020) Molecular confirmation, constituents and cytotoxicity evaluation of two medicinal Piper species used by the Manobo tribe of Agusan del Sur, Philippines. Phytochem Lett 36:24–31. https://doi.org/10.1016/j.phytol.2020.01.017
Suwanphakdee C, Chantaranothai P (2011) A new species and three taxonomic changes in Piper (Piperaceae) from Thailand. Blumea 56:235–239. https://doi.org/10.3767/000651911X607338
Thebpatiphat S, Pengprecha S, Ternai B (1988) Some constituents of the stems of Piper interruptum Opiz. J Sci Soc 14:225–231. https://doi.org/10.2306/scienceasia1513-1874.1988.14.225
Sireeratawong S, Iharat A, Lerdvuthisopon N, Piyabhan P, Khonsung P, Boonraeng S, Jaijoy K (2012) Anti-inflammatory, analgesic, and antipyretic activities of the ethanol extract of Piper interruptum Opiz. and Piper chaba Linn. ISRN Pharmacol. https://doi.org/10.5402/2012/480265
Temrangsee P, Itharat A, Sattaponpan C, Pipatrattanaseree W (2019) Inhibitory effect on alpha-glucosidase activity of Benchakul, Soros Benjakul and their plant components. TMJ 19(4):645–653
Thanatakitti P, Sathornwiriyapong S, Charupant K, Wangboonsakul C, Picheansunthorn C (2010) Specification of Sakhan. J Thai Trad Alt Med 8(1):47–60
Sudmoon R, Tanee T, Wongpanich V, Bletter N, Chaveerach A (2012) Ethnobotany and species specific molecular markers of some medicinal sakhan (Piper, Piperaceae). J Med Plant Res 6(7):1168–1175. https://doi.org/10.5897/JMPR11.807
Committee of Flora of China (1999) Piper. Flora of China 4:110–129
Gardner RO (2006) Piper (Piperaceae) in the Philippines Islands: the climbing species. Blumea 51:569–586. https://doi.org/10.3767/000651906X622139
Gardner RO (2010) Piper (Piperaceae) in the Solomon Islands: the climbing species. Blumea 55:4–13. https://doi.org/10.3767/000651910X499114
Gardner RO (2013) Piper (Piperaceae) in New Guinea: the climbing species. Blumea 57:275–294. https://doi.org/10.3767/000651903X686051
Director of National Parks, Australian Government. Piper interruptum. Australian National Botanic Garden Centre for Australian National Biodiversity Research. https://www.anbg.gov.au/cpbr/cd keys/RFK7/key/RFK7/Media/Html/entities/Piper_ interruptum.htm Accessed 29 May 2021
Lecomte PH (1910) Piperaceae. In: Lecomte PH (ed) Fl. Indo-Chine. 2(5):78
The plant list (2021) http://www.theplantlist.org/ Accessed 23 May 2021
Persson C (2001) Phylogenetic relationships in Polygalaceae based on plastid DNA sequences from the trnL-F region. Taxon 50:763–779. https://doi.org/10.2307/1223706
Hodkinson TR, Chase MW, Lledó DM, Salamin N, Renvoize SA (2002) Phylogenetics of Miscanthus, Saccharum and related genera (Saccharinae, Andropogoneae, Poaceae) based on DNA sequences from ITS nuclear ribosomal DNA and plastid trnL intron and trnL-F intergenic spacers. J Plant Res 115:381–392. https://doi.org/10.1007/s10265-002-0049-3
Hodkinson TR, Chase MW, Takahashi C, Leitch IJ, Bennett MD, Renvoize SA (2002) The use of DNA sequencing (ITS and trnL-F), AFLP, and fluorescent in situ hybridization to study allopolyploid Miscanthus (Poaceae). Am J Bot 89(2):279–286. https://doi.org/10.3732/ajb.89.2.279
Pirie MD, Vargas MPB, Botermans M, Bakker FT, Chatrou LW (2007) Ancient paralogy in the cpDNA trnL-F region in Annonaceae: implications for plant molecular systematics. Am J Bot 94(6):1003–1016. https://doi.org/10.3732/ajb.94.6.1003
Neubig KM, Whitten WM, Carlsward BS, Blanco MA, Endara L, Williams NH, Moore M (2009) Phylogenetic utility of ycf1 in orchids: a plastid gene more variable than matK. Plant Syst Evol 277:75–84. https://doi.org/10.1007/s00606-008-0105-0
Dastpak A, Osaloo SK, Maassoumi AA (2018) Safar KN. Molecular phylogeny of Astragalus sect. Ammodendron (Fabaceae) inferred from chloroplast ycf1 gene. Ann Bot Fennici 55:75–82. https://doi.org/10.5735/085.055.0108
Amar MH (2020) ycf1-ndhF genes, the most promising plastid genomic barcode, sheds light on phylogeny at low taxonomic levels in Prunus persica. J Genet Eng Biotechnol 18:42. https://doi.org/10.1186/s43141-020-00057-3
Downie SR, Katz-Downie DS, Watson MF (2000) A phylogeny of the flowering plant family Apiaceae based on chloroplast DNA rpl16 and rpoC1 intron sequences: toward a suprageneric classification of subfamily Apioideae. Am J Bot 87(2):273–292. https://doi.org/10.2307/2656915
Rydberg A (2010) DNA barcoding as a tool for the identification of unknown plant material. Biological Education Centre and Department of Systematic Biology, Uppsala University
Naim DM, Mahboob S (2020) Molecular identification of herbal species belonging to genus Piper within family Piperaceae from Northern Peninsular Malaysia. J King Saud Univ Sci 32:1417–1426. https://doi.org/10.1016/j.jksus.2019.11.036
Egydio Brandão APM, Yamaguchi LF, Tepe EJ, Salatino A, Kato MJ (2020) Evaluation of DNA markers for molecular identification of three Piper species from Brazilian Atlantic Rainforest. PLoS ONE 15(10):e0239056. https://doi.org/10.1371/journal.pone.0239056
Sass C, Little DP, Stevenson DW, Specht CD (2007) DNA Barcoding in the Cycadales: testing the potential of proposed barcoding markers for species identification of cycads. PLoS ONE 2(11):e1154. https://doi.org/10.1371/journal.pone.0001154
Chen S, Yao H, Han J, Liu C, Song J, Shi L, Zhu Y, Ma X, Gao T, Pang X, Luo K, Li Y, Li X, Jia X, Lin Y, Leon C (2010) Validation of the ITS2 region as a novel DNA barcode for identifying medicinal plant species. PLoS ONE 5(1):e8613. https://doi.org/10.1371/journal.pone.0008613
Gao T, Yao H, Song J, Liu C, Zhu Y, Ma X, Pang X, Xu H, Chen S (2010) Identification of medicinal plants in the family Fabaceae using a potential DNA barcode ITS2. J Ethnopharmacol 130:116–121. https://doi.org/10.1016/j.jep.2010.04.026
Pang X, Song J, Zhu Y, Xu H, Huang L, Chen S (2011) Applying plant DNA barcodes for Rosaceae species identification. Cladistics 27:165–170. https://doi.org/10.1111/j.1096-0031.2010.00328.x
Pang X, Shi L, Song J, Chen X, Chen S (2013) Use of the potential DNA ITS2 to identify herbal materials. J Nat Med 67:571–575. https://doi.org/10.1007/s11418-012-0715-2
Han J, Zhu Y, Chen X, Liao B, Yao H, Jingyuan S, Chen S, Meng F (2013) The short ITS2 sequence serves as an efficient taxonomic sequence tag in comparison with the full-length ITS. Bio Med Res Int. https://doi.org/10.1155/2013/741476
Chen S, Pang X, Song J, Shi L, Yao H, Han J, Leon C (2014) A renaissance in herbal medicine identification: from morphology to DNA. Biotechnol Adv 32:1237–1244. https://doi.org/10.1016/j.biotechadv.2014.07.004
Parveen I, Gafner S, Techen N, Murch SJ, Khan IA (2016) DNA barcoding for the identification of botanicals in herbal medicine and dietary supplements: strengths and limitation. Planta Med 82:1225–1235. https://doi.org/10.1055/s-0042-111208
Gehrke B, Bräuchler C, Romoleroux K, Lundberg M, Heubl G, Erikson T (2008) Molecular phylogenetics of Alchemilla, Aphanes and Lachemilla (Rosaceae) inferred from plastid and nuclear intron and spacer DNA sequences, with comments on generic classification. Mol Phylogenetics Evol 47:1030–1044. https://doi.org/10.1016/j.ympev.2008.03.004
Lui Z, Zeng X, Yang D, Chu G, Yuan Z, Chen S (2012) Applying DNA barcodes for identification of plant species in the family Araliaceae. Gene 499:76–80. https://doi.org/10.1016/j.gene.2012.02.016
Acknowledgements
We are grateful to Graduate Studies, Faculty of Pharmacy, Chiang Mai University for financial and equipment supports and thankful for technician of Laboratory of Pharmacognosy, Faculty of Pharmacy, Chiang Mai University to provide research facility.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Pinya, T., Intharuksa, A., Yanaso, S. et al. Conventional and molecular pharmacognostic characters integrated with chemical profiles of five Piper plants in the Thai herbal pharmacopoeia and their admixture/adulteration/substitution situations in Thailand. J Nat Med 76, 605–620 (2022). https://doi.org/10.1007/s11418-022-01607-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11418-022-01607-4