Abstract
Introduction
A plasmid named pDNS10 was detected from an atrazine-degrading strain Arthrobacter sp. DNS10 which has been isolated previously in our laboratory.
Materials and methods
In this paper, a special plasmid-detecting method and drop assays experiments were mainly used to achieve research goals.
Results and discussion
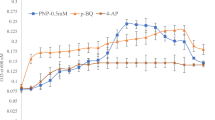
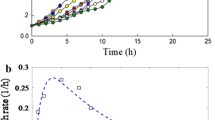
pDNS10 exhibited an excellent stability because it also could be detected even when the strain DNS10 has been subcultured under nonselective conditions for eight times. Over a 48-h incubation period, the OD600 of samples inoculated with strain DNS10 and strain DNS10-ST (both of them contained pDNS10) were 0.31 ± 0.042 and 0.305 ± 0.034, respectively ,whereas the OD600 of samples inoculated strain without pDNS10 (strain DNS10-PE) was only 0.138 ± 0.018. No atrazine was detected in the inoculated strain DNS10 and strain DNS10-ST samples at this period. Contrarily, the atrazine-degrading rate of strain DNS10-PE was only 5.23 ± 0.71%. Furthermore, both the two types of strains containing pDNS10 confirmed the presence of known degrading genes such as trzN, atzB, and atzC. It suggests that pDNS10 is an atrazine catabolic plasmid. In drop assays experiments, the wild-type strain DNS10 cells were chemotactically attracted to atrazine, whereas strain DNS10-PE showed no chemotaxis to atrazine and hydroxyatrazine. There was some relationship between atrazine degradation and the chemotactic response towards atrazine in strain DNS10.
Conclusions
The biochemical characteristics of pDNS10 and the chemotaxis characteristics of strain DNS10 could help us in better understanding of the mechanism of atrazine degradation by strain DNS10.




Similar content being viewed by others
References
Assaf NA, Turco RF (1994) Influence of carbon and nitrogen application on the mineralization of atrazine and its metabolites in soil. Pest Sci 41:41–47
Barbash JE, Thelin GP, Kolpin DW, Gilliom RJ (2001) Major herbicides in ground water: results from the national water-quality assessment. J Environ Qual 30:831–845
Beasley FC (2004) Characterization of diversity, chromate resistance, and aromatic hydrocarbon degradation among Arthrobacter isolates from mixed waste soil. MS Thesis. Purdue University. West Lafayette
Cheney MA, Shin JY, Crowley DE, Alvey S, Malengreau N, Sposito G (1998) Atrazine dealkylation on a manganese oxide surface. Colloid Surf 137:267–273
Crocker FH, Fredrickson JK, White DC, Ringelberg DB, Balkwill DL (2000) Phylogenetic and physiological diversity of Arthrobacter strains isolated from unconsolidated subsurface sediments. Microbiology 146:1295–1310
de Souza ML, Seffernick J, Martinez B, Sadowsky MJ, Wackett LP (1998) The atrazine catabolism genes atzABC are widespread and highly conserved. J Bacteriol 180:1951–1954
Dejonghe W, Goris J, Fantroussi SE, Höfte M, Vos PD, Verstraete W, Top EM (2000) Effect of dissemination of 2,4-dichlorophenoxyacetic acid (2,4-D) degradation plasmids on 2,4-D degradation and on bacterial community structure in two different soil horizons. Appl Environ Microbiol 66:3297–3304
Dennis JJ, Zylstra GJ (2004) Complete sequence and genetic organization of pDTG1, the 83 kilobase naphthalene degradation plasmid from Pseudomonas putida strain NCIB 9816-4. J Mol Biol 341:753–768
Fredrickson JK, Zachara JM, Balkwill DL, Kennedy D, Li SW, Kostandarithes HM, Daly MJ, Romine MF, Brockman FJ (2004) Geomicrobiology of high-level nuclear waste-contaminated vadose sediments at the Hanford site, Washington state. Appl Environ Microbiol 70:4230–4241
Getenga Z, Dörfler U, Iwobi A, Schmid M, Schroll R (2009) Atrazine and terbuthylazine mineralization by an Arthrobacter sp. isolated from a sugarcane-cultivated soil in Kenya. Chemosphere 77:534–539
Gogarten JP, Townsend JP (2005) Horizontal gene transfer, genome innovation and evolution. Nat Rev Microbiol 3:679–6873
Grimm AC, Harwood CS (1999) NahY, a catabolic plasmid-encoded receptor required for chemotaxis of Pseudomonas putida to the aromatic hydrocarbon naphthalene. J Bacteriol 181:3310–3316
Harwood CS, Nichols NN, Kim MK, Ditty JL, Parales RE (1994) Identification of the pcaRKF gene cluster from Pseudomonas putida: involvement in chemotaxis, biodegradation, and transport of 4-hydroxybenzoate. J Bacteriol 176:6479–6488
Hayes T (2002) Hermaphroditic, demasculinized frogs after exposure to the herbicide, atrazine, at low ecologically relevant doses. Proc Nat Acad Sci 99:5476–5480
Iwaki H, Muraki T, Ishihara S, Hasegawa Y, Rankin KN, Sulea T, Boyd J, Lau PC (2007) Characterization of a pseudomonad 2-nitrobenzoate nitroreductase and its catabolic pathway-associated 2-hydroxylaminobenzoate mutase and a chemoreceptor involved in 2-nitrobenzoate chemotaxis. J Bacteriol 189:3502–3514
Kado CI, Liu ST (1981) Rapid procedure for detection and isolation of large and small plasmids. J Bacteriol 145:1365–1373
Li Q, Li Y, Zhu X, Cai B (2008) Isolation and characterization of atrazine-degrading Arthrobacter sp. AD26 and use of this strain in bioremediation of contaminated soil. J Environ Sci 20:1226–1230
Liu X, Parales RE (2009) Bacterial chemotaxis to atrazine and related s-triazines. Appl Environ Microbiol 75:5481–5488
Mandelbaum RT, Allan DL, Wackett LP (1995) Isolation and characterization of a Pseudomonas sp. that mineralizes the s-triazine herbicide atrazine. Appl Environ Microbiol 61:1451–1457
Mulbry WW, Zhu H, Nour SM, Topp E (2002) The triazine hydrolase gene trzN from Nocardioides sp. strain C190: cloning and construction of gene-specific primers. FEMS Microbiol Lett 206:75–79
Pandey G, Jain RK (2002) Bacterial chemotaxis toward environmental pollutants: role in bioremediation. Appl Environ Microbiol 68:5789–5795
Parales RE, Harwood CS (2002) Bacterial chemotaxis to pollutants and plant-derived aromatic molecules. Curr Opin Microbiol 5:266–273
Rhine ED, Fuhrmann JJ, Radosevich M (2003) Microbial community responses to atrazine exposure and nutrient availability: linking degradation capacity to community structure. Microb Ecol 46:145–160
Rousseaux S, Soulas G, Hartmann A (2002) Plasmid localisation of atrazine-degrading genes in newly described Chelatobacter and Arthrobacter strains. FEMS Microbiol Ecol 41:69–75
Sajjaphan K, Shapir N, Wackett LP, Palmer M, Blackmon B, Tomkins J, Sadowsky MJ (2004) Arthrobacter aurescens TC1 atrazine catabolism genes trzN, atzB, and atzC are linked on a 160-kilobase region and are functional in Escherichia coli. Appl Environ Microb 70:4402–4407
Samanta SK, Jain RK (2000) Evidence for plasmid-mediated chemotaxis of Pseudomonas putida towards naphthalene and salicylate. Can J Microbiol 46:1–6
Smith D, Alvey S, Crowley DE (2005) Cooperative catabolic pathways within an atrazine-degrading enrichment culture isolated from soil. FEMS Microbiol Ecol 53:265–273
Strong LC, Rosendahl C, Johnson G, Sadowsky MJ, Wackett LP (2002) Arthrobacter aurescens TC1 metabolizes diverse s-triazine ring compounds. Appl Environ Microbiol 68:5973–5980
Vaishampayan PA, Kanekar PP, Dhakephalkar PK (2007) Isolation and characterization of Arthrobacter sp. strain MCM B-436, an atrazine-degrading bacterium, from rhizospheric soil. Int Biodeter Biodegr 60:273–278
Wackett LP, Sadowsky MJ, Martinez B, Shapir N (2002) Biodegradation of atrazine and related s-triazine compounds: from enzymes to field studies. Appl Microbiol Biot 58:39–45
Wang JH, Zhu LS, Liu AJ, Ma TT, Wang Q, Xie H, Wang J, Jiang T, Zhao RS (2010) Isolation and characterization of an Arthrobacter sp. strain HB-5 that transforms atrazine. Environ Geochem Hlth 33:259–266
Yang C, Li Y, Zhang K, Wang X, Ma CQ, Tang HZ, Xu P (2010) Atrazine degradation by a simple consortium of Klebsiella sp. A1 and Comamonas sp. A2 in nitrogen enriched medium. Biodegradation 21:97–105
Zhang Y, Jiang Z, Cao B, Hu M, Wang ZG, Dong XN (2011) Metabolic ability and gene characteristics of Arthrobacter sp. strain DNS10, the sole atrazine-degrading strain in a consortium isolated from black soil. Int Biodeter Biodegr 65:1140–1144
Acknowledgment
This research was supported by the National Natural Science Foundation of China (30970525), Program for New Century Excellent Talents in Heilongjiang Provincial University (1155-NCET-006), Science Foundation for Distinguished Young Scholars of Heilongjiang Province (JC201006), and New Century Excellent Talents in University (NCET-10-0145), Chang Jiang Scholar Candidates Program for Provincial Universities in Heilongjiang (CSCP).
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Philippe Garrigues
Co-first author: Zhao Jiang
Rights and permissions
About this article
Cite this article
Zhang, Y., Jiang, Z., Cao, B. et al. Chemotaxis to atrazine and detection of a xenobiotic catabolic plasmid in Arthrobacter sp. DNS10. Environ Sci Pollut Res 19, 2951–2958 (2012). https://doi.org/10.1007/s11356-012-0805-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-012-0805-4