Abstract
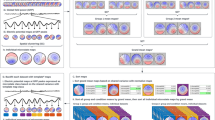
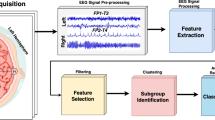
Significant heterogeneity in network structures reflecting individuals’ dynamic processes can exist within subgroups of people (e.g., diagnostic category, gender). This makes it difficult to make inferences regarding these predefined subgroups. For this reason, researchers sometimes wish to identify subsets of individuals who have similarities in their dynamic processes regardless of any predefined category. This requires unsupervised classification of individuals based on similarities in their dynamic processes, or equivalently, in this case, similarities in their network structures of edges. The present paper tests a recently developed algorithm, S-GIMME, that takes into account heterogeneity across individuals with the aim of providing subgroup membership and precise information about the specific network structures that differentiate subgroups. The algorithm has previously provided robust and accurate classification when evaluated with large-scale simulation studies but has not yet been validated on empirical data. Here, we investigate S-GIMME’s ability to differentiate, in a purely data-driven manner, between brain states explicitly induced through different tasks in a new fMRI dataset. The results provide new evidence that the algorithm was able to resolve, in an unsupervised data-driven manner, the differences between different active brain states in empirical fMRI data to segregate individuals and arrive at subgroup-specific network structures of edges. The ability to arrive at subgroups that correspond to empirically designed fMRI task conditions, with no biasing or priors, suggests this data-driven approach can be a powerful addition to existing methods for unsupervised classification of individuals based on their dynamic processes.








Similar content being viewed by others
References
Aghabozorgi, S., Shirkhorshidi, A. S., & Wah, T. Y. (2015). Time-series clustering: A decade review. Information Systems, 53, 16–38.
Arizmendi, C., Gates, K., Fredrickson, B., & Wright, A. (2021). Specifying exogeneity and bilinear effects in data-driven model searches. Behavior Research Methods, 53(3), 1276–1288.
Bellman, R. (1966). Dynamic programming. Science, 153(3731), 34–37.
Beltz, A. M., & Gates, K. M. (2017). Network mapping with gimme. Multivariate Behavioral Research, 52(6), 789–804.
Beltz, A. M., Gates, K. M., Engels, A. S., Molenaar, P. C., Pulido, C., Turrisi, R., Berenbaum, S. A., Gilmore, R. O., & Wilson, S. J. (2013). Changes in alcohol-related brain networks across the first year of college: A prospective pilot study using fMRI effective connectivity mapping. Addictive Behaviors, 38(4), 2052–2059.
Beltz, A. M., & Molenaar, P. C. (2016). Dealing with multiple solutions in structural vector autoregressive models. Multivariate Behavioral Research, 51(2–3), 357–373.
Brett, M., Anton, J.-L., Valabregue, R., Poline, J.-B., et al. (2002). Region of interest analysis using an spm toolbox. In 8th international conference on functional mapping of the human brain, vol. 16, page 497. Sendai.
Bringmann, L. F., Pe, M. L., Vissers, N., Ceulemans, E., Borsboom, D., Vanpaemel, W., Tuerlinckx, F., & Kuppens, P. (2016). Assessing temporal emotion dynamics using networks. Assessment, 23(4), 425–435.
Brodersen, K. H., Deserno, L., Schlagenhauf, F., Lin, Z., Penny, W. D., Buhmann, J. M., & Stephan, K. E. (2014). Dissecting psychiatric spectrum disorders by generative embedding. NeuroImage: Clinical, 4, 98–111.
Brown, T. A. (2006). Confirmatory factor analysis for applied research.
Bullmore, E., & Sporns, O. (2009). Complex brain networks: Graph theoretical analysis of structural and functional systems. Nature Reviews Neuroscience, 10(3), 186–198.
Caliński, T., & Harabasz, J. (1974). A dendrite method for cluster analysis. Communications in Statistics-theory and Methods, 3(1), 1–27.
Dajani, D. R., Burrows, C. A., Nebel, M. B., Mostofsky, S. H., Gates, K. M., & Uddin, L. Q. (2019). Parsing heterogeneity in autism spectrum disorder and attention-deficit/hyperactivity disorder with individual connectome mapping. Brain Connectivity, 9(9), 673–691.
Dickie, E. W., Ameis, S. H., Shahab, S., Calarco, N., Smith, D. E., Miranda, D., Viviano, J. D., & Voineskos, A. N. (2018). Personalized intrinsic network topography mapping and functional connectivity deficits in autism spectrum disorder. Biological Psychiatry, 84(4), 278–286.
Dubois, J., Galdi, P., Paul, L. K., & Adolphs, R. (2018). A distributed brain network predicts general intelligence from resting-state human neuroimaging data. Philosophical Transactions of the Royal Society B: Biological Sciences, 373(1756), 20170284.
Duda, R. O., Hart, P. E., et al. (1973). Pattern classification and scene analysis (Vol. 3). New York: Wiley.
Duffy, K. A., Fisher, Z. F., Arizmendi, C. A., Molenaar, P. C., Hopfinger, J., Cohen, J. R., Beltz, A. M., Lindquist, M. A., Hallquist, M. N., & Gates, K. M. (2021). Detecting task-dependent functional connectivity in group iterative multiple model estimation with person-specific hemodynamic response functions. Brain Connectivity, 11(6), 418–429.
Easson, A. K., Fatima, Z., & McIntosh, A. R. (2019). Functional connectivity-based subtypes of individuals with and without autism spectrum disorder. Network Neuroscience, 3(2), 344–362.
Enders, C. K. (2001). The impact of nonnormality on full information maximum-likelihood estimation for structural equation models with missing data. Psychological Methods, 6(4), 352.
Epskamp, S., van Borkulo, C. D., van der Veen, D. C., Servaas, M. N., Isvoranu, A.-M., Riese, H., & Cramer, A. O. (2018). Personalized network modeling in psychopathology: The importance of contemporaneous and temporal connections. Clinical Psychological Science, 6(3), 416–427.
Ernst, A. F., Timmerman, M. E., Jeronimus, B. F., & Albers, C. J. (2021). Insight into individual differences in emotion dynamics with clustering. Assessment, 28(4), 1186–1206.
Finn, E. S. & Constable, R. T. (2022). Individual variation in functional brain connectivity: Implications for personalized approaches to psychiatric disease. Dialogues in Clinical Neuroscience.
Fisher, A. J., & Boswell, J. F. (2016). Enhancing the personalization of psychotherapy with dynamic assessment and modeling. Assessment, 23(4), 496–506.
Friston, K. J., Holmes, A. P., Worsley, K. J., Poline, J.-P., Frith, C. D., & Frackowiak, R. S. (1994). Statistical parametric maps in functional imaging: A general linear approach. Human Brain Mapping, 2(4), 189–210.
Gates, K. M., Fisher, Z. F., & Bollen, K. A. (2020). Latent variable gimme using model implied instrumental variables (MIIVs). Psychological Methods, 25(2), 227.
Gates, K. M., Henry, T., Steinley, D., & Fair, D. A. (2016). A monte carlo evaluation of weighted community detection algorithms. Frontiers in Neuroinformatics, 10, 45.
Gates, K. M., Lane, S. T., Varangis, E., Giovanello, K., & Guiskewicz, K. (2017). Unsupervised classification during time-series model building. Multivariate Behavioral Research, 52(2), 129–148.
Gates, K. M., & Molenaar, P. C. (2012). Group search algorithm recovers effective connectivity maps for individuals in homogeneous and heterogeneous samples. NeuroImage, 63(1), 310–319.
Gates, K. M., Molenaar, P. C., Iyer, S. P., Nigg, J. T., & Fair, D. A. (2014). Organizing heterogeneous samples using community detection of gimme-derived resting state functional networks. PLoS ONE, 9(3), e91322.
Golino, H. F., & Epskamp, S. (2017). Exploratory graph analysis: A new approach for estimating the number of dimensions in psychological research. PLoS ONE, 12(6), e0174035.
Gumus, M., Mack, M. L., Green, R., Khodadadi, M., Wennberg, R. A., Crawley, A., Colella, B., Tarazi, A., Mikulis, D., & Tator, C. H., et al. (2022). Brain connectivity changes in postconcussion syndrome as the neural substrate of a heterogeneous syndrome. Brain Connectivity.
Heller, R., Stanley, D., Yekutieli, D., Rubin, N., & Benjamini, Y. (2006). Cluster-based analysis of FMRI data. NeuroImage, 33(2), 599–608.
Hennig, C. (2020). fpc: Flexible Procedures for clustering. R package version 2.2-9.
Henry, T. R., Feczko, E., Cordova, M., Earl, E., Williams, S., Nigg, J. T., Fair, D. A., & Gates, K. M. (2019). Comparing directed functional connectivity between groups with confirmatory subgrouping gimme. NeuroImage, 188, 642–653.
Henry, T. R., Feczko, E., Cordova, M., Earl, E., Williams, S., Nigg, J. T., Fair, D. A., & Gates, K. M. (2019). Comparing directed functional connectivity between groups with confirmatory subgrouping gimme. NeuroImage, 188, 642–653.
Hurlburt, R. T., Alderson-Day, B., Fernyhough, C., & Kühn, S. (2015). What goes on in the resting-state? A qualitative glimpse into resting-state experience in the scanner. Frontiers in Psychology, 6, 1535.
Kanwisher, N., McDermott, J., & Chun, M. M. (1997). The fusiform face area: A module in human extrastriate cortex specialized for face perception. Journal of Neuroscience, 17(11), 4302–4311.
Lane, S., Gates, K., Fisher, Z., and Molenaar, P. (2021). Package ‘gimme’.
Lane, S. T., & Gates, K. M. (2017). Automated selection of robust individual-level structural equation models for time series data. Structural Equation Modeling: A Multidisciplinary Journal, 24(5), 768–782.
Lane, S. T., Gates, K. M., Pike, H. K., Beltz, A. M., & Wright, A. G. (2019). Uncovering general, shared, and unique temporal patterns in ambulatory assessment data. Psychological Methods, 24(1), 54.
Laumann, T. O., Gordon, E. M., Adeyemo, B., Snyder, A. Z., Joo, S. J., Chen, M.-Y., Gilmore, A. W., McDermott, K. B., Nelson, S. M., Dosenbach, N. U., et al. (2015). Functional system and areal organization of a highly sampled individual human brain. Neuron, 87(3), 657–670.
Liao, T. W. (2005). Clustering of time series data: A survey. Pattern Recognition, 38(11), 1857–1874.
Luo, L., Fisher, Z. F., Arizmendi, C., Molenaar, P., Beltz, A., & Gates, K. M. (2022). Estimating both directed and undirected contemporaneous relations in time series data using hybrid-group iterative multiple model estimation. Psychological Methods.
Lütkepohl, H. (2005). New introduction to multiple time series analysis. New York: Springer.
MacQueen, J. (1967). Classification and analysis of multivariate observations. In 5th Berkeley Symposium on Mathematical Statistics and Probability, pp. 281–297.
Mathôt, S., Schreij, D., & Theeuwes, J. (2012). Opensesame: An open-source, graphical experiment builder for the social sciences. Behavior Research Methods, 44(2), 314–324.
McLachlan, G., & Chang, S. (2004). Mixture modelling for cluster analysis. Statistical Methods in Medical Research, 13(5), 347–361.
Miller, M. B., & Van Horn, J. D. (2007). Individual variability in brain activations associated with episodic retrieval: A role for large-scale databases. International Journal of Psychophysiology, 63(2), 205–213.
Miranda, L., Paul, R., Pütz, B., Koutsouleris, N., & Müller-Myhsok, B. (2021). Systematic review of functional MRI applications for psychiatric disease subtyping. Frontiers in Psychiatry, 12.
Molenaar, P. C. (2017). Equivalent dynamic models. Multivariate Behavioral Research, 52(2), 242–258.
Newman, M. E. (2004). Fast algorithm for detecting community structure in networks. Physical Review E, 69(6), 066133.
Nichols, T. T., Gates, K. M., Molenaar, P. C., & Wilson, S. J. (2014). Greater bold activity but more efficient connectivity is associated with better cognitive performance within a sample of nicotine-deprived smokers. Addiction Biology, 19(5), 931–940.
Olszowy, W., Aston, J., Rua, C., & Williams, G. B. (2019). Accurate autocorrelation modeling substantially improves fMRI reliability. Nature Communications, 10(1), 1–11.
Pons, P. & Latapy, M. (2005). Computing communities in large networks using random walks. In International symposium on computer and information sciences, pp. 284–293. Springer.
Power, J. D., Cohen, A. L., Nelson, S. M., Wig, G. S., Barnes, K. A., Church, J. A., Vogel, A. C., Laumann, T. O., Miezin, F. M., Schlaggar, B. L., et al. (2011). Functional network organization of the human brain. Neuron, 72(4), 665–678.
Price, R. B., Gates, K., Kraynak, T. E., Thase, M. E., & Siegle, G. J. (2017). Data-driven subgroups in depression derived from directed functional connectivity paths at rest. Neuropsychopharmacology, 42(13), 2623–2632.
Price, R. B., Lane, S., Gates, K., Kraynak, T. E., Horner, M. S., Thase, M. E., & Siegle, G. J. (2017). Parsing heterogeneity in the brain connectivity of depressed and healthy adults during positive mood. Biological Psychiatry, 81(4), 347–357.
Rosenberg, M. D., Finn, E. S., Scheinost, D., Papademetris, X., Shen, X., Constable, R. T., & Chun, M. M. (2016). A neuromarker of sustained attention from whole-brain functional connectivity. Nature Neuroscience, 19(1), 165–171.
Rubinov, M., & Sporns, O. (2010). Complex network measures of brain connectivity: Uses and interpretations. NeuroImage, 52(3), 1059–1069.
Saris, W. E., Satorra, A., & Sörbom, D. (1987). The detection and correction of specification errors in structural equation models. Sociological Methodology, pp. 105–129.
Scherf, K. S., Behrmann, M., Humphreys, K., & Luna, B. (2007). Visual category-selectivity for faces, places and objects emerges along different developmental trajectories. Developmental Science, 10(4), F15–F30.
Schwarz, G. (1978). Estimating the dimension of a model. The Annals of Statistics, pp. 461–464.
Shirer, W. R., Ryali, S., Rykhlevskaia, E., Menon, V., & Greicius, M. D. (2012). Decoding subject-driven cognitive states with whole-brain connectivity patterns. Cerebral Cortex, 22(1), 158–165.
Sörbom, D. (1989). Model modification. Psychometrika, 54(3), 371–384.
Sporns, O. (2016). Networks of the brain. MIT Press.
Tokuda, T., Yoshimoto, J., Shimizu, Y., Okada, G., Takamura, M., Okamoto, Y., Yamawaki, S., & Doya, K. (2018). Identification of depression subtypes and relevant brain regions using a data-driven approach. Scientific Reports, 8(1), 1–13.
Tottenham, N., Tanaka, J. W., Leon, A. C., McCarry, T., Nurse, M., Hare, T. A., Marcus, D. J., Westerlund, A., Casey, B. J., & Nelson, C. (2009). The nimstim set of facial expressions: Judgments from untrained research participants. Psychiatry Research, 168(3), 242–249.
Volkmar, F. R., Lord, C., Bailey, A., Schultz, R. T., & Klin, A. (2004). Autism and pervasive developmental disorders. Journal of Child Psychology and Psychiatry, 45(1), 135–170.
Wang, Y., Tang, S., Zhang, L., Bu, X., Lu, L., Li, H., Gao, Y., Hu, X., Kuang, W., Jia, Z., et al. (2021). Data-driven clustering differentiates subtypes of major depressive disorder with distinct brain connectivity and symptom features. The British Journal of Psychiatry, 219(5), 606–613.
Ward, J. H., Jr., & Hook, M. E. (1963). Application of an hierarchical grouping procedure to a problem of grouping profiles. Educational and Psychological Measurement, 23(1), 69–81.
Weigard, A., Lane, S., Gates, K., & Beltz, A. (2021). The influence of autoregressive relation strength and search strategy on directionality recovery in group iterative multiple model estimation. Psychological Methods.
Wright, A. G., Gates, K. M., Arizmendi, C., Lane, S. T., Woods, W. C., & Edershile, E. A. (2019). Focusing personality assessment on the person: Modeling general, shared, and person specific processes in personality and psychopathology. Psychological Assessment, 31(4), 502.
Xu, P., Huang, R., Wang, J., Van Dam, N. T., Xie, T., Dong, Z., Chen, C., Gu, R., Zang, Y.-F., He, Y., et al. (2014). Different topological organization of human brain functional networks with eyes open versus eyes closed. NeuroImage, 90, 246–255.
Yang, Z., Xu, Y., Xu, T., Hoy, C. W., Handwerker, D. A., Chen, G., Northoff, G., Zuo, X.-N., & Bandettini, P. A. (2014). Brain network informed subject community detection in early-onset schizophrenia. Scientific Reports, 4(1), 1–12.
Yang, Z., Xu, Y., Xu, T., Hoy, C. W., Handwerker, D. A., Chen, G., Northoff, G., Zuo, X.-N., & Bandettini, P. A. (2014). Brain network informed subject community detection in early-onset schizophrenia. Scientific Reports, 4(1), 1–12.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
We acknowledge the primary funding source for this project: National Institute of Health-National Institute of Biomedical Imaging and Bioengineering (R01-EB02290).
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fisher, Z.F., Parsons, J., Gates, K.M. et al. Blind Subgrouping of Task-based fMRI. Psychometrika 88, 434–455 (2023). https://doi.org/10.1007/s11336-023-09907-8
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11336-023-09907-8