Abstract
Introduction
Myasthenia gravis (MG) and rheumatoid arthritis (RA) are examples of antibody-mediated chronic, progressive autoimmune diseases. Phenotypically dissimilar, MG and RA share common immunological features. However, the immunometabolomic features common to humoral autoimmune diseases remain largely unexplored.
Objectives
The aim of this study was to reveal and illustrate the metabolomic profile overlap found between these two diseases and describe the immunometabolomic significance.
Methods
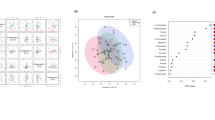
Metabolic analyses using acid- and dansyl-labelled was performed on serum from adult patients with seropositive MG (n = 46), RA (n = 23) and healthy controls (n = 49) presenting to the University of Alberta Hospital specialty clinics. Chemical isotope labelling liquid chromatography mass spectrometry (CIL LC–MS) methods were utilized to assess the serum metabolome in patients; 12C/13C-dansyl chloride (DnsCl) was used to label amine/phenol metabolites and 12C/13C-p-dimethylaminophenacyl bromide (DmPA) was used for carboxylic acids. Metabolites matching our criteria for significance were selected if they were present in both groups. Multivariate statistical analysis [including principal component analysis (PCA) and partial least squares discriminant analysis (PLS-DA)] and biochemical pathway analysis was then conducted to gain understanding of the principal pathways involved in antibody-mediated pathogenesis.
Results
We found 20 metabolites dysregulated in both MG and RA when compared to healthy controls. Most prominently, observed changes were related to pathways associated with phenylalanine metabolism, tyrosine metabolism, ubiquinone and other terpenoid-quinone biosynthesis, and pyruvate metabolism.
Conclusion
From these results it is evident that many metabolites are common to humoral disease and exhibit significant immunometabolomic properties. This observation may lead to an enhanced understanding of the metabolic underpinnings common to antibody-mediated autoimmune disease. Further, contextualizing these findings within a larger clinical and systems biology context could provide new insights into the pathogenesis and management of these diseases.




Similar content being viewed by others
References
Akaogi, J., Barker, T., Kuroda, Y., Nacionales, D. C., Yamasaki, Y., Stevens, B. R., et al. (2006). Role of non-protein amino acid l-canavanine in autoimmunity. Autoimmunity Reviews, 5(6), 429–435. https://doi.org/10.1016/j.autrev.2005.12.004.
Allen, R. H., Stabler, S. P., Savage, D. G., & Lindenbaum, J. (1993). Metabolic abnormalities in cobalamin (vitamin B12) and folate deficiency. FASEB Journal: Official Publication of the Federation of American Societies for Experimental Biology, 7(14), 1344–1353. https://doi.org/10.1096/fasebj.7.14.7901104.
Amersfoort, J., & Kuiper, J. (2017). T cell metabolism in metabolic disease-associated autoimmunity. Immunobiology, 222(10), 925–936. https://doi.org/10.1016/j.imbio.2017.03.001.
Amorini, A. M., Nociti, V., Petzold, A., Gasperini, C., Quartuccio, E., Lazzarino, G., et al. (2014). Serum lactate as a novel potential biomarker in multiple sclerosis. Biochimica et Biophysica Acta—Molecular Basis of Disease, 1842(7), 1137–1143. https://doi.org/10.1016/j.bbadis.2014.04.005.
Araki, K., Turner, A. P., Shaffer, V. O., Gangappa, S., Keller, S. A., Bachmann, M. F., et al. (2009). mTOR regulates memory CD8 T-cell differentiation. Nature, 460(7251), 108–112. https://doi.org/10.1038/nature08155.
Astudillo, A. M., Balgoma, D., Balboa, M. A., & Balsinde, J. (2012). Dynamics of arachidonic acid mobilization by inflammatory cells. Biochimica et Biophysica Acta (BBA)—Molecular and Cell Biology of Lipids, 1821(2), 249–256. https://doi.org/10.1016/j.bbalip.2011.11.006.
Bass, A. J., Dabney, A., & Robinson, D. (2015). qvalue: Q-value estimation for false discovery rate control. R package version 2.10.0. Retrieved from http://github.com/jdstorey/qvalue.
Bengtsson, A. A., Trygg, J., Wuttge, D. M., Sturfelt, G., Theander, E., Donten, M., et al. (2016). Metabolic profiling of systemic lupus erythematosus and comparison with primary Sjögren’s syndrome and systemic sclerosis. PLoS ONE, 11(7), e0159384. https://doi.org/10.1371/journal.pone.0159384.
Bhargava, P., & Calabresi, P. A. (2016). Metabolomics in multiple sclerosis. Multiple Sclerosis, 22(4), 451–460. https://doi.org/10.1177/1352458515622827.
Blackmore, D., Siddiqi, Z., Li, L., Wang, N., & Maksymowych, W. (2019). Beyond the antibodies: Serum metabolomic profiling of myasthenia gravis. Metabolomics. https://doi.org/10.1007/s11306-019-1571-9.
Bonuccelli, G., Tsirigos, A., Whitaker-Menezes, D., Pavlides, S., Pestell, R. G., Chiavarina, B., et al. (2010). Ketones and lactate “fuel” tumor growth and metastasis. Cell Cycle, 9(17), 3506–3514. https://doi.org/10.4161/cc.9.17.12731.
Buck, M. D., Sowell, R. T., Kaech, S. M., & Pearce, E. L. (2017). Metabolic Instruction of Immunity. Cell, 169(4), 570–586. https://doi.org/10.1016/j.cell.2017.04.004.
Chiang, J. Y. L. (2013). Bile acid metabolism and signaling. Comprehensive Physiology, 3(3), 1191–1212. https://doi.org/10.1002/cphy.c120023.
Choi, S. Y. C., Collins, C. C., Gout, P. W., & Wang, Y. (2013). Cancer-generated lactic acid: A regulatory, immunosuppressive metabolite? Journal of Pathology, 230(4), 350–355. https://doi.org/10.1002/path.4218.
Cummings, B. S., McHowat, J., & Schnellmann, R. G. (2000). Phospholipase A2s in cell injury and death. Journal of Pharmacology and Experimental Therapeutics, 294(3), 793–799.
de Candia, P., De Rosa, V., Gigantino, V., Botti, G., Ceriello, A., & Matarese, G. (2017). Immunometabolism of human autoimmune diseases: from metabolites to extracellular vesicles. FEBS Letters, 591(19), 3119–3134. https://doi.org/10.1002/1873-3468.12733.
De Rosa, V., Galgani, M., Porcellini, A., Colamatteo, A., Santopaolo, M., Zuchegna, C., et al. (2015). Glycolysis controls the induction of human regulatory T cells by modulating the expression of FOXP3 exon 2 splicing variants. Nature Immunology, 16(11), 1174–1184. https://doi.org/10.1038/ni.3269.
Dimeloe, S., Burgener, A.-V., Grählert, J., & Hess, C. (2017). T-cell metabolism governing activation, proliferation and differentiation; a modular view. Immunology, 150(1), 35–44. https://doi.org/10.1111/imm.12655.
Ernster, L., & Dallner, G. (1995). Biochemical, physiological and medical aspects of ubiquinone function. Biochimica et Biophysica Acta, 1271(1), 195–204.
Foster, D. W. (2012). Malonyl-CoA: The regulator of fatty acid synthesis and oxidation. The Journal of Clinical Investigation, 122(6), 1958–1959. https://doi.org/10.1172/jci63967.
Franken, C., Lambrechts, N., Govarts, E., Koppen, G., Den Hond, E., Ooms, D., et al. (2017). Phthalate-induced oxidative stress and association with asthma-related airway inflammation in adolescents. International Journal of Hygiene and Environmental Health, 220(2), 468–477. https://doi.org/10.1016/j.ijheh.2017.01.006.
Frolkis, A., Knox, C., Lim, E., Jewison, T., Law, V., Hau, D. D., et al. (2010). SMPDB: The Small Molecule Pathway Database. Nucleic Acids Research, 38(Database issue), D480-7.
Gentleman, R. C., Carey, V. J., Bates, D. M., Bolstad, B., Dettling, M., Dudoit, S., et al. (2004). Bioconductor: open software development for computational biology and bioinformatics. Genome Biology, 5(10), R80. https://doi.org/10.1186/gb-2004-5-10-r80.
Grist, J. T., Jarvis, L. B., Georgieva, Z., Thompson, S., Kaur Sandhu, H., Burling, K., et al. (2018). Extracellular lactate: A novel measure of T cell proliferation. Journal of immunology (Baltimore, Md. : 1950), 200(3), 1220–1226. https://doi.org/10.4049/jimmunol.1700886.
Guzik, T. J., Korbut, R., & Adamek-Guzik, T. (2003). Nitric oxide and superoxide in inflammation and immune regulation. Journal of Physiology and Pharmacology: An Official Journal of the Polish Physiological Society, 54(4), 469–487.
Haas, R., Smith, J., Rocher-Ros, V., Nadkarni, S., Montero-Melendez, T., D’Acquisto, F., et al. (2015). Lactate regulates metabolic and pro-inflammatory circuits in control of T cell migration and effector functions. PLoS Biology, 13(7), e1002202. https://doi.org/10.1371/journal.pbio.1002202.
Hart, I. K. (2000). Acquired neuromyotonia: A new autoantibody-mediated neuronal potassium channelopathy. American Journal of the Medical Sciences, 319(4), 209–216.
Hirschhaeuser, F., Sattler, U. G. A., & Mueller-Klieser, W. (2011). Lactate: A metabolic key player in cancer. Cancer Research, 71(22), 6921–6925. https://doi.org/10.1158/0008-5472.can-11-1457.
Hutchinson, D. (1999). Classification criteria: The 1987 American Rheumatism Association revised criteria for the classification of rheumatoid arthritis. CPD Rheumatology, 1(1), 13–14.
Jaigirdar, S. A., Benson, R. A., Elmesmari, A., Kurowska-Stolarska, M. S., McInnes, I. B., Garside, P., et al. (2017). Sphingosine-1-phosphate promotes the persistence of activated CD4 T cells in inflamed sites. Frontiers in Immunology. https://doi.org/10.3389/fimmu.2017.01627.
Jenkins, R. C., & Weetman, A. P. (2002). Disease associations with autoimmune thyroid disease. Thyroid, 12(11), 977–988. https://doi.org/10.1089/105072502320908312.
Kaikkonen, J., Nyyssönen, K., Tuomainen, T.-P., Ristonmaa, U., & Salonen, J. T. (1999). Determinants of plasma coenzyme Q10 in humans. FEBS Letters, 443(2), 163–166. https://doi.org/10.1016/s0014-5793(98)01712-8.
Kang, J., Zhu, L., Lu, J., & Zhang, X. (2015). Application of metabolomics in autoimmune diseases: Insight into biomarkers and pathology. Journal of Neuroimmunology, 279, 25–32. https://doi.org/10.1016/j.jneuroim.2015.01.001.
Kim, Y. S. (2002). Malonate metabolism: Biochemistry, molecular biology, physiology, and industrial application. Journal of Biochemistry and Molecular Biology, 35(5), 443–451.
Kim, S., Hwang, J., Kim, J., Ahn, J. K., Cha, H.-S., & Kim, K. H. (2017). Metabolite profiles of synovial fluid change with the radiographic severity of knee osteoarthritis. Joint Bone Spine, 84(5), 605–610. https://doi.org/10.1016/j.jbspin.2016.05.018.
Knebel, L. A., Zanatta, Â., Tonin, A. M., Grings, M., Alvorcem, L. D. M., Wajner, M., et al. (2012). 2-Methylbutyrylglycine induces lipid oxidative damage and decreases the antioxidant defenses in rat brain. Brain Research, 1478, 74–82. https://doi.org/10.1016/j.brainres.2012.08.039.
Kolins, J., & Gilroy, J. (1972). Serum enzyme levels in patients with myasthenia gravis after aerobic and ischaemic exercise. Journal of Neurology, Neurosurgery and Psychiatry, 35(1), 34–40.
Kominsky, D. J., Campbell, E. L., & Colgan, S. P. (2010). Metabolic shifts in immunity and inflammation. Journal of immunology (Baltimore, Md. : 1950), 184(8), 4062–4068. https://doi.org/10.4049/jimmunol.0903002.
Lafay, S., & Gil-Izquierdo, A. (2008). Bioavailability of phenolic acids. Phytochemistry Reviews, 7(2), 301–311. https://doi.org/10.1007/s11101-007-9077-x.
Lee, Y. S., Wollam, J., & Olefsky, J. M. (2018). An integrated view of immunometabolism. Cell, 172(1–2), 22–40. https://doi.org/10.1016/j.cell.2017.12.025.
Li, J., Xie, X.-W., Zhou, H., Wang, B., Zhang, M.-J., & Tang, F.-Y. (2017). Metabolic profiling reveals new serum biomarkers of lupus nephritis. Lupus, 26(11), 1166–1173. https://doi.org/10.1177/0961203317694256.
Lindahl, A., Forshed, J., & Nordström, A. (2016). Overlap in serum metabolic profiles between non-related diseases: Implications for LC-MS metabolomics biomarker discovery. Biochemical and Biophysical Research Communications, 478(3), 1472–1477. https://doi.org/10.1016/j.bbrc.2016.08.155.
Loftus, R. M., & Finlay, D. K. (2016). Immunometabolism: Cellular metabolism turns immune regulator. The Journal of Biological Chemistry, 291(1), 1–10. https://doi.org/10.1074/jbc.r115.693903.
Lone, A. M., & Taskén, K. (2013). Proinflammatory and immunoregulatory roles of eicosanoids in T cells. Frontiers in Immunology, 4, 130. https://doi.org/10.3389/fimmu.2013.00130.
Lu, Y., Wang, C., Chen, Z., Zhao, H., Chen, J., Liu, X., et al. (2012). Serum metabolomics for the diagnosis and classification of myasthenia gravis. Metabolomics, 8(4), 704–713. https://doi.org/10.1007/s11306-011-0364-6.
Maceyka, M., Harikumar, K. B., Milstien, S., & Spiegel, S. (2012). Sphingosine-1-phosphate signaling and its role in disease. Trends in Cell Biology, 22(1), 50–60. https://doi.org/10.1016/j.tcb.2011.09.003.
Maceyka, M., & Spiegel, S. (2014). Sphingolipid metabolites in inflammatory disease. Nature, 510(7503), 58–67. https://doi.org/10.1038/nature13475.
Manley, K., Han, W., Zelin, G., & Lawrence, D. A. (2018). Crosstalk between the immune, endocrine, and nervous systems in immunotoxicology. Current Opinion in Toxicology, 10, 37–45. https://doi.org/10.1016/j.cotox.2017.12.003.
Masubuchi, N., Sugihara, M., Sugita, T., Amano, K., Nakano, M., & Matsuura, T. (2016). Oxidative stress markers, secondary bile acids and sulfated bile acids classify the clinical liver injury type: Promising diagnostic biomarkers for cholestasis. Chemico-Biological Interactions, 255, 83–91. https://doi.org/10.1016/j.cbi.2015.08.016.
Matsuki, T., Horai, R., Sudo, K., & Iwakura, Y. (2003). IL-1 plays an important role in lipid metabolism by regulating insulin levels under physiological conditions. The Journal of Experimental Medicine, 198(6), 877–888. https://doi.org/10.1084/jem.20030299.
Mehta, M. M., & Chandel, N. S. (2015). Targeting metabolism for lupus therapy. Science Translational Medicine. https://doi.org/10.1126/scitranslmed.aaa6731.
Michalek, R. D., Gerriets, V. A., Jacobs, S. R., Macintyre, A. N., MacIver, N. J., Mason, E. F., et al. (2011). Cutting edge: Distinct glycolytic and lipid oxidative metabolic programs are essential for effector and regulatory CD4 + T cell subsets. The Journal of Immunology, 186(6), 3299–3303. https://doi.org/10.4049/jimmunol.1003613.
Moco, S., Candela, M., Chuang, E., Draper, C., Cominetti, O., Montoliu, I., et al. (2014). Systems biology approaches for inflammatory bowel disease: Emphasis on gut microbial metabolism. Inflammatory Bowel Diseases, 20(11), 2104–2114. https://doi.org/10.1097/mib.0000000000000116.
Narang, A. S., & Boddu, S. H. S. (Eds.). (2015). Excipient applications in formulation design and drug delivery. Cham: Springer International Publishing. https://doi.org/10.1007/978-3-319-20206-8.
Nguyen, A., & Bouscarel, B. (2008). Bile acids and signal transduction: Role in glucose homeostasis. Cellular Signalling, 20(12), 2180–2197. https://doi.org/10.1016/j.cellsig.2008.06.014.
O’Neill, L. A. J., Kishton, R. J., & Rathmell, J. (2016). A guide to immunometabolism for immunologists. Nature Reviews Immunology, 16(9), 553–565. https://doi.org/10.1038/nri.2016.70.
Olenchock, B. A., Rathmell, J. C., & Vander Heiden, M. G. (2017). Biochemical underpinnings of immune cell metabolic phenotypes. Immunity, 46(5), 703–713. https://doi.org/10.1016/j.immuni.2017.04.013.
Pacheco, R., Contreras, F., & Zouali, M. (2014). The dopaminergic system in autoimmune diseases. Frontiers in Immunology. https://doi.org/10.3389/fimmu.2014.00117.
Pan, Y., Tian, T., Park, C. O., Lofftus, S. Y., Mei, S., Liu, X., et al. (2017). Survival of tissue-resident memory T cells requires exogenous lipid uptake and metabolism. Nature, 543(7644), 252–256. https://doi.org/10.1038/nature21379.
Pearce, E. L., & Pearce, E. J. (2013). Metabolic pathways in immune cell activation and quiescence. Immunity, 38(4), 633–643. https://doi.org/10.1016/j.immuni.2013.04.005.
Pearce, E. L., Poffenberger, M. C., Chang, C.-H., & Jones, R. G. (2013). Fueling immunity: Insights into metabolism and lymphocyte function. Science, 342(6155), 1242454. https://doi.org/10.1126/science.1242454.
Pelz, A., Schaffert, H., Diallo, R., Hiepe, F., Meisel, A., & Kohler, S. (2018). S1P receptor antagonists fingolimod and siponimod do not improve the outcome of experimental autoimmune myasthenia gravis mice after disease onset. European Journal of Immunology, 48(3), 498–508. https://doi.org/10.1002/eji.201747187.
Poffenberger, M. C., & Jones, R. G. (2014). Amino acids fuel T cell-mediated inflammation. Immunity, 40(5), 635–637. https://doi.org/10.1016/j.immuni.2014.04.017.
Regenold, W. T., Phatak, P., Makley, M. J., Stone, R. D., & Kling, M. A. (2008). Cerebrospinal fluid evidence of increased extra-mitochondrial glucose metabolism implicates mitochondrial dysfunction in multiple sclerosis disease progression. Journal of the Neurological Sciences, 275(1–2), 106–112. https://doi.org/10.1016/j.jns.2008.07.032.
Rodríguez-Carrio, J., Alperi-López, M., López, P., Ballina-García, F. J., & Suárez, A. (2016). Non-esterified fatty acids profiling in rheumatoid arthritis: associations with clinical features and Th1 response. PLoS ONE, 11(8), e0159573. https://doi.org/10.1371/journal.pone.0159573.
Sabelli, H. C., Fawcett, J., Gusovsky, F., Javaid, J., Edwards, J., & Jeffriess, H. (1983). Urinary phenyl acetate: A diagnostic test for depression? Science, 220(4602), 1187–1188.
Schaeffler, A., Gross, P., Buettner, R., Bollheimer, C., Buechler, C., Neumeier, M., et al. (2009). Fatty acid-induced induction of Toll-like receptor-4/nuclear factor-κB pathway in adipocytes links nutritional signalling with innate immunity. Immunology, 126(2), 233–245. https://doi.org/10.1111/j.1365-2567.2008.02892.x.
Schieber, M., & Chandel, N. S. (2014). ROS function in redox signaling and oxidative stress. Current Biology: CB, 24(10), R453–R462. https://doi.org/10.1016/j.cub.2014.03.034.
Schönfeld, P., & Reiser, G. (2013). Why does brain metabolism not favor burning of fatty acids to provide energy? Reflections on disadvantages of the use of free fatty acids as fuel for brain. Journal of Cerebral Blood Flow and Metabolism: Official Journal of the International Society of Cerebral Blood Flow and Metabolism, 33(10), 1493–1499. https://doi.org/10.1038/jcbfm.2013.128.
Schönfeld, P., & Wojtczak, L. (2016). Short- and medium-chain fatty acids in energy metabolism: The cellular perspective. Journal of Lipid Research, 57(6), 943–954. https://doi.org/10.1194/jlr.r067629.
Seeger, K. (2009). Metabolic changes in autoimmune diseases. Current Drug Discovery Technologies, 6(4), 256–261.
Seki, S. M., & Gaultier, A. (2017). Exploring non-metabolic functions of glycolytic enzymes in immunity. Frontiers in Immunology, 8, 1549. https://doi.org/10.3389/fimmu.2017.01549.
Sengupta, M., Cheema, A., Kaminski, H. J., Kusner, L. L., Gribbestad, I., et al. (2014). Serum metabolomic response of myasthenia gravis patients to chronic prednisone treatment. PLoS ONE, 9(7), e102635. https://doi.org/10.1371/journal.pone.0102635.
Shi, H., Kokoeva, M. V., Inouye, K., Tzameli, I., Yin, H., & Flier, J. S. (2006). TLR4 links innate immunity and fatty acid–induced insulin resistance. Journal of Clinical Investigation, 116(11), 3015–3025. https://doi.org/10.1172/jci28898.
Shin, T. H., Kim, H.-A., Jung, J.-Y., Baek, W.-Y., Lee, H.-S., Park, H. J., et al. (2018). Analysis of the free fatty acid metabolome in the plasma of patients with systemic lupus erythematosus and fever. Metabolomics, 14(1), 14. https://doi.org/10.1007/s11306-017-1308-6.
Siemieniuk, E., & Skrzydlewska, E. (2005). Coenzyme Q10: its biosynthesis and biological significance in animal organisms and in humans. Postepy Higieny I Medycyny Doswiadczalnej (Online), 59, 150–159.
Sotnikova, T. D., Beaulieu, J.-M., Espinoza, S., Masri, B., Zhang, X., Salahpour, A., et al. (2010). The dopamine metabolite 3-methoxytyramine is a neuromodulator. PLoS ONE, 5(10), e13452. https://doi.org/10.1371/journal.pone.0013452.
Staels, B., & Fonseca, V. A. (2009). Bile acids and metabolic regulation: mechanisms and clinical responses to bile acid sequestration. Diabetes Care, 32(Suppl 2), S237–S245. https://doi.org/10.2337/dc09-s355.
Staley, C., Weingarden, A. R., Khoruts, A., & Sadowsky, M. J. (2017). Interaction of gut microbiota with bile acid metabolism and its influence on disease states. Applied Microbiology and Biotechnology, 101(1), 47–64. https://doi.org/10.1007/s00253-016-8006-6.
Storey, J. D. (2003). The positive false discovery rate: A Bayesian interpretation and the q -value. The Annals of Statistics, 31(6), 2013–2035. https://doi.org/10.1214/aos/1074290335.
van der Windt, G. J. W., Everts, B., Chang, C.-H., Curtis, J. D., Freitas, T. C., Amiel, E., et al. (2012). Mitochondrial respiratory capacity is a critical regulator of CD8 + T cell memory development. Immunity, 36(1), 68–78. https://doi.org/10.1016/j.immuni.2011.12.007.
van der Windt, G. J. W., O’Sullivan, D., Everts, B., Huang, S. C.-C., Buck, M. D., Curtis, J. D., et al. (2013). CD8 memory T cells have a bioenergetic advantage that underlies their rapid recall ability. Proceedings of the National Academy of Sciences, 110(35), 14336–14341. https://doi.org/10.1073/pnas.1221740110.
Vashi, P., Edwin, P., Popiel, B., Lammersfeld, C., & Gupta, D. (2016). Methylmalonic acid and homocysteine as indicators of vitamin B-12 deficiency in cancer. PLoS ONE, 11(1), e0147843. https://doi.org/10.1371/journal.pone.0147843.
Villoslada, P., Alonso, C., Agirrezabal, I., Kotelnikova, E., Zubizarreta, I., Pulido-Valdeolivas, I., et al. (2017). Metabolomic signatures associated with disease severity in multiple sclerosis. Neurology: Neuroimmunology and NeuroInflammation. https://doi.org/10.1212/nxi.0000000000000321.
Wakil, S. J., & Abu-Elheiga, L. A. (2009). Fatty acid metabolism: target for metabolic syndrome. Journal of Lipid Research, 50(Supplement), S138–S143. https://doi.org/10.1194/jlr.r800079-jlr200.
Walenta, S., Schroeder, T., & Mueller-Klieser, W. (2004). Lactate in solid malignant tumors: Potential basis of a metabolic classification in clinical oncology. Current Medicinal Chemistry, 11(16), 2195–2204. https://doi.org/10.2174/0929867043364711.
Westerblad, H., Allen, D. G., & Lännergren, J. (2002). Muscle fatigue: Lactic acid or inorganic phosphate the major cause? Physiology, 17(1), 17–21. https://doi.org/10.1152/physiologyonline.2002.17.1.17.
Win-Shwe, T.-T., Yanagisawa, R., Koike, E., Nitta, H., & Takano, H. (2013). Expression levels of neuroimmune biomarkers in hypothalamus of allergic mice after phthalate exposure. Journal of Applied Toxicology, 33(10), 1070–1078. https://doi.org/10.1002/jat.2835.
Wu, G. (2013). Amino acids. Boca Raton: CRC Press. https://doi.org/10.1201/b14661.
Wu, T., Xie, C., Han, J., Ye, Y., Weiel, J., Li, Q., et al. (2012). Metabolic disturbances associated with systemic lupus erythematosus. PLoS ONE. https://doi.org/10.1371/journal.pone.0037210.
Xia, J., Mandal, R., Sinelnikov, I. V., Broadhurst, D., & Wishart, D. S. (2012). MetaboAnalyst 2.0–a comprehensive server for metabolomic data analysis. Nucleic Acids Research, 40(W1), W127–W133. https://doi.org/10.1093/nar/gks374.
Yan, B., Huang, J., Zhang, C., Hu, X., Gao, M., Shi, A., et al. (2016). Serum metabolomic profiling in patients with systemic lupus erythematosus by GC/MS. Modern Rheumatology, 26(6), 914–922. https://doi.org/10.3109/14397595.2016.1158895.
Yang, Z., Matteson, E. L., Goronzy, J. J., & Weyand, C. M. (2015). T-cell metabolism in autoimmune disease. Arthritis Research & Therapy, 17(1), 29. https://doi.org/10.1186/s13075-015-0542-4.
Yegambaram, M., Manivannan, B., Beach, T. G., & Halden, R. U. (2015). Role of environmental contaminants in the etiology of Alzheimer’s disease: A review. Current Alzheimer Research, 12(2), 116–146. https://doi.org/10.2174/1567205012666150204121719.
Yin, Y., Choi, S.-C., Xu, Z., Perry, D. J., Seay, H., Croker, B. P., et al. (2015). Normalization of CD4+ T cell metabolism reverses lupus. Science Translational Medicine. https://doi.org/10.1126/scitranslmed.aaa0835.
Young, S. P., Kapoor, S. R., Viant, M. R., Byrne, J. J., Filer, A., Buckley, C. D., et al. (2013). The impact of inflammation on metabolomic profiles in patients with arthritis. Arthritis and Rheumatism, 65(8), 2015–2023. https://doi.org/10.1002/art.38021.
Yuan, Z.-Y., Gao, S.-G., Mu, J.-W., Xue, Q., Mao, Y.-S., Wang, D.-L., et al. (2016). Prognostic value of preoperative serum lactate dehydrogenase in thymic carcinoma. Journal of Thoracic Disease, 8(9), 2464. https://doi.org/10.21037/jtd.2016.08.56.
Zhao, C., Fernandes, M. J., Turgeon, M., Tancrède, S., Di Battista, J., Poubelle, P. E., et al. (2008). Specific and overlapping sphingosine-1-phosphate receptor functions in human synoviocytes: Impact of TNF-α. Journal of Lipid Research, 49(11), 2323–2337. https://doi.org/10.1194/jlr.m800143-jlr200.
Zhao, S., Luo, X., & Li, L. (2016). Chemical isotope labeling LC-MS for high coverage and quantitative profiling of the hydroxyl submetabolome in metabolomics. Analytical Chemistry, 88(21), 10617–10623. https://doi.org/10.1021/acs.analchem.6b02967.
Zhao, B., Moochhala, S. M., Lu, J., Tan, D., & Hoon Lai, M. (2006). Determination of pyridostigmine bromide and its metabolites in biological samples. The Journal of Pharmacy and Pharmaceutical Sciences (www.cspsCanada.org), 9(1), 71–81.
Zhou, J., Chen, J., Hu, C., Xie, Z., Li, H., Wei, S., et al. (2016). Exploration of the serum metabolite signature in patients with rheumatoid arthritis using gas chromatography–mass spectrometry. Journal of Pharmaceutical and Biomedical Analysis, 127, 60–67. https://doi.org/10.1016/j.jpba.2016.02.004.
Zhou, R., Tseng, C.-L., Huan, T., & Li, L. (2014). IsoMS: Automated processing of LC-MS data generated by a chemical isotope labeling metabolomics platform. Analytical Chemistry, 86(10), 4675–4679. https://doi.org/10.1021/ac5009089.
Zhu, C., Fuchs, C. D., Halilbasic, E., & Trauner, M. (2016). Bile acids in regulation of inflammation and immunity: Friend or foe? Clinical and Experimental Rheumatology, 34(4 Suppl 98), 25–31.
Author information
Authors and Affiliations
Contributions
D.B. developed the concept and designed the experiment, acquired all blood samples but those from rheumatoid arthritis patients, conducted all databasing, bioinformatics analyses and metabolite-database matching. Further, D.B. prepared all charts, images and tables and wrote the paper. L.L. developed the chemical labelling process and several chemometric analysis tools used in this study as well as the chemical identification libraries used in the positive identification of observed metabolites. L.L. provided oversight in the choice of chemical and statistical techniques used to characterize the metabolome. L.L also offered conceptual advice, supervised project analysis and edited the manuscript. N.W. performed the sample preparation, chemical labelling and mass spectrometric analysis. W.M provided the rheumatoid arthritis samples for the disease control cohort. E.Y. contributed to study planning and design, procurement of rheumatoid samples and providing periodic guidance. Z.S. developed the concept and provided project oversight as the clinical expert in myasthenia gravis and enrolled study patients. Z.S. also offered conceptual advice, supervised project analysis and edited the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interests.
Ethical approval
This work received ethics approval from the University of Alberta Research Ethics Board and is in compliance with the ethical standards of this institutional research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Blackmore, D., Li, L., Wang, N. et al. Metabolomic profile overlap in prototypical autoimmune humoral disease: a comparison of myasthenia gravis and rheumatoid arthritis. Metabolomics 16, 10 (2020). https://doi.org/10.1007/s11306-019-1625-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-019-1625-z