Abstract
Introduction
Microorganisms catabolize carbon-containing compounds in their environment during growth, releasing a subset of metabolic byproducts as volatile compounds. However, the relationship between growth media and the production of volatile compounds has been largely unexplored to-date.
Objectives
To assess the core and media-specific components of the Klebsiella pneumoniae volatile metabolome via growth in four in vitro culture media.
Methods
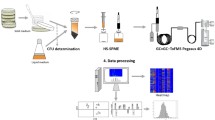
Headspace volatiles produced by cultures of K. pneumoniae after growth to stationary phase in four rich media (brain heart infusion broth, lysogeny broth, Mueller-Hinton broth, and tryptic soy broth) were analyzed using comprehensive two-dimensional gas chromatography time-of-flight mass spectrometry (GC×GC-TOFMS). Differences in the composition of headspace volatiles as a function of growth media were assessed using hierarchical clustering analysis (HCA) and principal component analysis (PCA).
Results
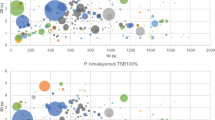
A total of 365 volatile compounds were associated with the growth of K. pneumoniae across all media, of which 36 (10%) were common to all growth media, and 148 (41%) were specific to a single medium. In addition, utilizing all K. pneumoniae-associated volatile compounds, strains clustered as a function of growth media, demonstrating the importance of media in determining the metabolic profile of this organism.
Conclusion
K. pneumoniae produces a core suite of volatile compounds across all growth media studied, although the volatile metabolic signature of this organism is fundamentally media-dependent.




Similar content being viewed by others
References
Bean, H. D., Dimandja, J. M., & Hill, J. E. (2012). Bacterial volatile discovery using solid phase microextraction and comprehensive two-dimensional gas chromatography-time-of-flight mass spectrometry. Journal of chromatography. B, Analytical Technologies in the Biomedical and Life Sciences, 901, 41–46.
Benjamini, Y., & Hochberg, Y. (1995). Controlling the false discovery rate - a practical and powerful approach to multiple testing. Journal of the Royal Statistical Society. Series B, Statistical Methodology, 57(1), 289–300.
Boots, A. W., Smolinska, A., Van Berkel, J. J., Fijten, R. R., Stobberingh, E. E., Boumans, M. L., Moonen, E. J., Wouters, E. F., Dallinga, J. W., & Van Schooten, F. J. (2014). Identification of microorganisms based on headspace analysis of volatile organic compounds by gas chromatography-mass spectrometry. Journal of Breath Research, 8(2), 027106.
Borer, A., Saidel-Odes, L., Riesenberg, K., Eskira, S., Peled, N., Nativ, R., Schlaeffer, F., & Sherf, M. (2009). Attributable mortality rate for carbapenem-resistant Klebsiella pneumoniae bacteremia. Infection Control and Hospital Epidemiology: The Official Journal of the Society of Hospital Epidemiologists of America, 30(10), 972–976.
Bos, L. D., Sterk, P. J., & Schultz, M. J. (2013). Volatile metabolites of pathogens: a systematic review. PLoS Pathogens, 9(5), e1003311.
Bruins, M., Bos, A., Petit, P. L., Eadie, K., Rog, A., Bos, R., Van Ramshorst, G. H., & Van Belkum, A. (2009). Device-independent, real-time identification of bacterial pathogens with a metal oxide-based olfactory sensor. European Journal of Clinical Microbiology & Infectious Diseases: Official Publication of the European Society of Clinical Microbiology, 28(7), 775–780.
Diancourt, L., Passet, V., Verhoef, J., Grimont, P. A., & Brisse, S. (2005). Multilocus sequence typing of Klebsiella pneumoniae nosocomial isolates. Journal of Clinical Microbiology, 43(8), 4178–4182.
Dieterle, F., Ross, A., Schlotterbeck, G., & Senn, H. (2006). Probabilistic quotient normalization as robust method to account for dilution of complex biological mixtures. Application in 1 H NMR metabonomics. Analytical Chemistry, 78(13), 4281–4290.
Elgaali, H., Hamilton-Kemp, T. R., Newman, M. C., Collins, R. W., Yu, K., & Archbold, D. D. (2002). Comparison of long-chain alcohols and other volatile compounds emitted from food-borne and related Gram positive and Gram negative bacteria. Journal of Basic Microbiology, 42(6), 373–380.
Gao, J., Zou, Y., Wang, Y., Wang, F., Lang, L., Wang, P., Zhou, Y., & Ying, K. (2016). Breath analysis for noninvasively differentiating Acinetobacter baumannii ventilator-associated pneumonia from its respiratory tract colonization of ventilated patients. Journal of Breath Research, 10(2), 027102.
Gisbert, J. P., & Pajares, J. M. (2004). Review article: 13 C-urea breath test in the diagnosis of Helicobacter pylori infection – a critical review. Alimentary Pharmacology and Therapeutics, 20(10), 1001–1017.
Hirsch, E. B., Chang, K. T., Zucchi, P. C., Francoeur, D. N., Ledesma, K. R., Tam, V. H., & Lasco, T. M. (2014). An evaluation of multiple phenotypic screening methods for Klebsiella pneumoniae carbapenemase (KPC)-producing Enterobacteriaceae. Journal of Infection and Chemotherapy: Official Journal of the Japan Society of Chemotherapy, 20(3), 224–227.
Hotelling, H. (1933). Analysis of a complex of statistical variables into principal components. Journal of Educational Psychology, 24, 417–441.
Julak, J., Prochazkova-Francisci, E., Stranska, E., & Rosova, V. (2003). Evaluation of exudates by solid phase microextraction-gas chromatography. Journal of microbiological methods, 52(1), 115–122.
Julak, J., Stranska, E., Prochazkova-Francisci, E., & Rosova, V. (2000). Blood cultures evaluation by gas chromatography of volatile fatty acids. Medical Science Monitor: International Medical Journal of Experimental and Clinical Research, 6(3), 605–610.
Junger, M., Vautz, W., Kuhns, M., Hofmann, L., Ulbricht, S., Baumbach, J. I., Quintel, M., & Perl, T. (2012). Ion mobility spectrometry for microbial volatile organic compounds: a new identification tool for human pathogenic bacteria. Applied Microbiology and Biotechnology, 93(6), 2603–2614.
Kiviranta, H., Tuomainen, A., Reiman, M., Laitinen, S., Liesivuori, J., & Nevalainen, A. (1998). Qualitative identification of volatile metabolites from two fungi and three bacteria species cultivated on two media. Central European Journal of Public Health, 6(4), 296–299.
Koo, S., Thomas, H. R., Daniels, S. D., Lynch, R. C., Fortier, S. M., Shea, M. M., Rearden, P., Comolli, J. C., Baden, L. R., & Marty, F. M. (2014). A breath fungal secondary metabolite signature to diagnose invasive aspergillosis. Clinical Infectious Diseases: an Official Publication of The Infectious Diseases Society of America, 59(12), 1733–1740.
Kovats, E. (1958). Gas-Chromatographische Charakterisierung Organischer Verbindungen 0.1. Retentionsindices Aliphatischer Halogenide, Alkohole, Aldehyde Und Ketone. Helvetica Chimica Acta, 41(7), 1915–1932.
Lechner, M., Fille, M., Hausdorfer, J., Dierich, M. P., & Rieder, J. (2005). Diagnosis of bacteria in vitro by mass spectrometric fingerprinting:a pilot study. Current Microbiology, 51(4), 267–269.
Lee, C. J., Demilo, A. B., Moreno, D. S., & Martinez, A. J. (1995). Analysis of the volatile components of a bacterial fermentation that is attractive to the mexican fruit-fly, Anastrepha ludens. Journal of Agricultural and Food Chemistry, 43(5), 1348–1351.
Liu, M., Durfee, T., Cabrera, J. E., Zhao, K., Jin, D. J., & Blattner, F. R. (2005). Global transcriptional programs reveal a carbon source foraging strategy by Escherichia coli. The Journal of Biological Chemistry, 280(16), 15921–15927.
Mann, H. B., & Whitney, D. R. (1947). On a Test of Whether One of 2 Random Variables Is Stochastically Larger Than the Other. The Annals of Mathematical Statistics, 18(1), 50–60.
Podschun, R., & Ullmann, U. (1998). Klebsiella spp. as nosocomial pathogens: epidemiology, taxonomy, typing methods, and pathogenicity factors. Clinical Microbiology Reviews, 11(4), 589–603.
Rees, C. A., Franchina, F. A., Nordick, K. V., Kim, P. J., & Hill, J. E. (2016a). Expanding the Klebsiella pneumoniae volatile metabolome using advanced analytical instrumentation for the detection of novel metabolites. Journal of Applied Microbiology. doi:10.1111/jam.13372.
Rees, C. A., Smolinska, A., & Hill, J. E. (2016b). The volatile metabolome of Klebsiella pneumoniae in human blood. Journal of Breath Research, 10(2), 027101.
Robacker, D. C., & Bartelt, R. J. (1997). Chemicals attractive to Mexican fruit fly from Klebsiella pneumoniae and Citrobacter freundii cultures sampled by solid-phase microextraction. Journal of Chemical Ecology, 23(12), 2897–2915.
Schulz, S., & Dickschat, J. S. (2007). Bacterial volatiles: the smell of small organisms. Natural Product Reports, 24(4), 814–842.
Scott-Thomas, A. J., Syhre, M., Pattemore, P. K., Epton, M., Laing, R., Pearson, J., & Chambers, S. T. (2010). 2-Aminoacetophenone as a potential breath biomarker for Pseudomonas aeruginosa in the cystic fibrosis lung. BMC Pulmonary Medicine, 10, 56.
Sethi, S., Nanda, R., & Chakraborty, T. (2013). Clinical application of volatile organic compound analysis for detecting infectious diseases. Clinical Microbiology Reviews, 26(3), 462–475.
Shnayderman, M., Mansfield, B., Yip, P., Clark, H. A., Krebs, M. D., Cohen, S. J., Zeskind, J. E., Ryan, E. T., Dorkin, H. L., Callahan, M. V., Stair, T. O., Gelfand, J. A., Gill, C. J., Hitt, B., & Davis, C. E. (2005). Species-specific bacteria identification using differential mobility spectrometry and bioinformatics pattern recognition. Analytical Chemistry, 77(18), 5930–5937.
Storer, M. K., Hibbard-Melles, K., Davis, B., & Scotter, J. (2011). Detection of volatile compounds produced by microbial growth in urine by selected ion flow tube mass spectrometry (SIFT-MS). Journal of Microbiological Methods, 87(1), 111–113.
Syhre, M., & Chambers, S. T. (2008). The scent of Mycobacterium tuberculosis. Tuberculosis, 88(4), 317–323.
Syhre, M., Manning, L., Phuanukoonnon, S., Harino, P., & Chambers, S. T. (2009). The scent of Mycobacterium tuberculosis–part II breath. Tuberculosis, 89(4), 263–266.
Tait, E., Perry, J. D., Stanforth, S. P., & Dean, J. R. (2014). Identification of volatile organic compounds produced by bacteria using HS-SPME-GC-MS. Journal of Chromatographic Science, 52(4), 363–373.
Zechman, J. M., Aldinger, S., & Labows, J. N. Jr. (1986). Characterization of pathogenic bacteria by automated headspace concentration-gas chromatography. Journal of Chromatography, 377, 49–57.
Zhu, J., Bean, H. D., Jimenez-Diaz, J., & Hill, J. E. (2013a). Secondary electrospray ionization-mass spectrometry (SESI-MS) breathprinting of multiple bacterial lung pathogens, a mouse model study. Journal of Applied Physiology (Bethesda, Md. : 1985), 114(11), 1544–1549.
Zhu, J., Bean, H. D., Wargo, M. J., Leclair, L. W., & Hill, J. E. (2013b). Detecting bacterial lung infections: in vivo evaluation of in vitro volatile fingerprints. Journal of Breath Research, 7(1), 016003.
Acknowledgements
This work was supported by National Institutes of Health grant R21 AI121076 to JE Hill and Lennart KA Lundblad. Dartmouth College holds an Institutional Program Unifying Population and Laboratory Based Sciences award from the Burroughs Wellcome Fund, and CA Rees was supported by this grant (Grant#1014106). KV Nordick was supported by the Paul K. Richter and Evalyn E. Cook Richter Memorial Fund awarded through Dartmouth College. AE Lewis was supported by the Presidential Scholarship awarded through Dartmouth College. We thank Mavra Nasir and Theodore R Mellors for their aid in editing this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors report no potential conficts of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Rees, C.A., Nordick, K.V., Franchina, F.A. et al. Volatile metabolic diversity of Klebsiella pneumoniae in nutrient-replete conditions. Metabolomics 13, 18 (2017). https://doi.org/10.1007/s11306-016-1161-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-016-1161-z