Abstract
Introduction
Malignant astrocytomas are composed of heterogeneous cell populations. Compared to grade IV glioblastoma, low-grade astrocytomas have more differentiated cells and are associated with a better prognosis. Therefore, inducing cellular differentiation to alter the behaviour of high-grade astrocytomas may serve as a therapeutic strategy. The nuclear factor one (NFI) transcription factors are essential for normal astrocytic differentiation. Here, we investigate whether family members NFIA and NFIB act as effectors of cellular differentiation in glioblastoma.
Methods
We analysed expression of NFIA and NFIB in mRNA expression data of high-grade astrocytoma and with immunofluorescence co-staining. Furthermore, we induced NFI expression in patient-derived subcutaneous glioblastoma xenografts via in vivo electroporation.
Results
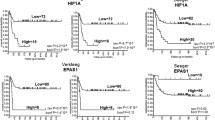
The expression of NFIA and NFIB is reduced in glioblastoma as compared to lower grade astrocytomas. At a cellular level, their expression is associated with differentiated and mature astrocyte-like tumour cells. In vivo analyses consistently demonstrate that expression of either NFIA or NFIB is sufficient to promote tumour cell differentiation in glioblastoma xenografts.
Conclusion
Our findings indicate that both NFIA and NFIB may have an endogenous pro-differentiative function in astrocytomas, similar to their role in normal astrocyte differentiation. Overall, our study establishes a basis for further investigation of targeting NFI-mediated differentiation as a potential differentiation therapy.





Similar content being viewed by others
References
Aum DJ, Kim DH, Beaumont TL, Leuthardt EC, Dunn GP, Kim AH (2014) Molecular and cellular heterogeneity: the hallmark of glioblastoma. Neurosurg Focus 37(6):E11. https://doi.org/10.3171/2014.9.FOCUS14521
Louis DN, Ohgaki H, Wiestler OD, Cavenee WK, Burger PC, Jouvet A, Scheithauer BW, Kleihues P (2007) The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol 114(2):97–109. https://doi.org/10.1007/s00401-007-0243-4
Patel AP, Tirosh I, Trombetta JJ, Shalek AK, Gillespie SM, Wakimoto H, Cahill DP, Nahed BV, Curry WT, Martuza RL, Louis DN, Rozenblatt-Rosen O, Suva ML, Regev A, Bernstein BE (2014) Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344(6190):1396–1401. https://doi.org/10.1126/science.1254257
Huang ME, Ye YC, Chen SR, Chai JR, Lu JX, Zhoa L, Gu LJ, Wang ZY (1988) Use of all-trans retinoic acid in the treatment of acute promyelocytic leukemia. Blood 72(2):567–572
Nowak D, Stewart D, Koeffler HP (2009) Differentiation therapy of leukemia: 3 decades of development. Blood 113(16):3655–3665. https://doi.org/10.1182/blood-2009-01-198911
Campos B, Wan F, Farhadi M, Ernst A, Zeppernick F, Tagscherer KE, Ahmadi R, Lohr J, Dictus C, Gdynia G, Combs SE, Goidts V, Helmke BM, Eckstein V, Roth W, Beckhove P, Lichter P, Unterberg A, Radlwimmer B, Herold-Mende C (2010) Differentiation therapy exerts antitumor effects on stem-like glioma cells. Clin Cancer Res 16(10):2715–2728. https://doi.org/10.1158/1078-0432.CCR-09-1800
Choschzick I, Hirseland E, Cramer H, Schultz S, Leppert J, Tronnier V, Zechel C (2014) Responsiveness of stem-like human glioma cells to all-trans retinoic acid and requirement of retinoic acid receptor isotypes alpha, beta and gamma. Neuroscience 279:44–64. https://doi.org/10.1016/j.neuroscience.2014.07.078
Caren H, Stricker SH, Bulstrode H, Gagrica S, Johnstone E, Bartlett TE, Feber A, Wilson G, Teschendorff AE, Bertone P, Beck S, Pollard SM (2015) Glioblastoma stem cells respond to differentiation cues but fail to undergo commitment and terminal cell-cycle arrest. Stem Cell Rep 5(5):829–842. https://doi.org/10.1016/j.stemcr.2015.09.014
Faigle R, Liu L, Cundiff P, Funa K, Xia Z (2008) Opposing effects of retinoid signaling on astrogliogenesis in embryonic day 13 and 17 cortical progenitor cells. J Neurochem 106(4):1681–1698. https://doi.org/10.1111/j.1471-4159.2008.05525.x
Hadinger N, Varga BV, Berzsenyi S, Kornyei Z, Madarasz E, Herberth B (2009) Astroglia genesis in vitro: distinct effects of retinoic acid in different phases of neural stem cell differentiation. Int J Dev Neurosci 27(4):365–375. https://doi.org/10.1016/j.ijdevneu.2009.02.004
Piper M, Moldrich RX, Lindwall C, Little E, Barry G, Mason S, Sunn N, Kurniawan ND, Gronostajski RM, Richards LJ (2009) Multiple non-cell-autonomous defects underlie neocortical callosal dysgenesis in Nfib-deficient mice. Neural Dev 4:43. https://doi.org/10.1186/1749-8104-4-43
Shu T, Butz KG, Plachez C, Gronostajski RM, Richards LJ (2003) Abnormal development of forebrain midline glia and commissural projections in Nfia knock-out mice. J Neurosci 23(1):203–212
Gobius I, Morcom L, Suarez R, Bunt J, Bukshpun P, Reardon W, Dobyns WB, Rubenstein JL, Barkovich AJ, Sherr EH, Richards LJ (2016) Astroglial-mediated remodeling of the interhemispheric midline is required for the formation of the corpus callosum. Cell Rep 17(3):735–747. https://doi.org/10.1016/j.celrep.2016.09.033
Namihira M, Kohyama J, Semi K, Sanosaka T, Deneen B, Taga T, Nakashima K (2009) Committed neuronal precursors confer astrocytic potential on residual neural precursor cells. Dev Cell 16(2):245–255. https://doi.org/10.1016/j.devcel.2008.12.014
Piper M, Barry G, Hawkins J, Mason S, Lindwall C, Little E, Sarkar A, Smith AG, Moldrich RX, Boyle GM, Tole S, Gronostajski RM, Bailey TL, Richards LJ (2010) NFIA controls telencephalic progenitor cell differentiation through repression of the Notch effector Hes1. J Neurosci 30(27):9127–9139. https://doi.org/10.1523/JNEUROSCI.6167-09.2010
Bunt J, Osinski JM, Lim JWC, Vidovic D, Ye Y, Zalucki O, O’Connor TR, Harris L, Gronostajski RM, Richards LJ, Piper M (2017) Combined allelic dosage of Nfia and Nfib regulates cortical development. Brain Neurosci Adv 1:1–21. https://doi.org/10.1177/2398212817739433
Plachez C, Lindwall C, Sunn N, Piper M, Moldrich RX, Campbell CE, Osinski JM, Gronostajski RM, Richards LJ (2008) Nuclear factor I gene expression in the developing forebrain. J Comp Neurol 508(3):385–401. https://doi.org/10.1002/cne.21645
Chen KS, Harris L, Lim JWC, Harvey TJ, Piper M, Gronostajski RM, Richards LJ, Bunt J (2017) Differential neuronal and glial expression of nuclear factor I proteins in the cerebral cortex of adult mice. J Comp Neurol 525(11):2465–2483. https://doi.org/10.1002/cne.24206
Caiazzo M, Giannelli S, Valente P, Lignani G, Carissimo A, Sessa A, Colasante G, Bartolomeo R, Massimino L, Ferroni S, Settembre C, Benfenati F, Broccoli V (2015) Direct conversion of fibroblasts into functional astrocytes by defined transcription factors. Stem Cell Rep 4(1):25–36. https://doi.org/10.1016/j.stemcr.2014.12.002
Canals I, Ginisty A, Quist E, Timmerman R, Fritze J, Miskinyte G, Monni E, Hansen MG, Hidalgo I, Bryder D, Bengzon J, Ahlenius H (2018) Rapid and efficient induction of functional astrocytes from human pluripotent stem cells. Nat Methods 15(9):693–696. https://doi.org/10.1038/s41592-018-0103-2
Tchieu J, Calder EL, Guttikonda SR, Gutzwiller EM, Aromolaran KA, Steinbeck JA, Goldstein PA, Studer L (2019) NFIA is a gliogenic switch enabling rapid derivation of functional human astrocytes from pluripotent stem cells. Nat Biotechnol 37(3):267–275. https://doi.org/10.1038/s41587-019-0035-0
Song HR, Gonzalez-Gomez I, Suh GS, Commins DL, Sposto R, Gilles FH, Deneen B, Erdreich-Epstein A (2010) Nuclear factor IA is expressed in astrocytomas and is associated with improved survival. Neuro-oncology 12(2):122–132. https://doi.org/10.1093/neuonc/nop044
Stringer BW, Bunt J, Day BW, Barry G, Jamieson PR, Ensbey KS, Bruce ZC, Goasdoue K, Vidal H, Charmsaz S, Smith FM, Cooper LT, Piper M, Boyd AW, Richards LJ (2016) Nuclear factor one B (NFIB) encodes a subtype-specific tumour suppressor in glioblastoma. Oncotarget 7(20):29306–29320. https://doi.org/10.18632/oncotarget.8720
Brat DJ, Seiferheld WF, Perry A, Hammond EH, Murray KJ, Schulsinger AR, Mehta MP, Curran WJ, Radiation Therapy Oncology Group (2004) Analysis of 1p, 19q, 9p, and 10q as prognostic markers for high-grade astrocytomas using fluorescence in situ hybridization on tissue microarrays from Radiation Therapy Oncology Group trials. Neuro-oncology 6(2):96–103
Rasheed A, Herndon JE, Stenzel TT, Raetz JG, Kendelhardt J, Friedman HS, Friedman AH, Bigner DD, Bigner SH, McLendon RE (2002) Molecular markers of prognosis in astrocytic tumors. Cancer 94(10):2688–2697
Johansson FK, Brodd J, Eklof C, Ferletta M, Hesselager G, Tiger CF, Uhrbom L, Westermark B (2004) Identification of candidate cancer-causing genes in mouse brain tumors by retroviral tagging. Proc Natl Acad Sci USA 101(31):11334–11337. https://doi.org/10.1073/pnas.0402716101
Vyazunova I, Maklakova VI, Berman S, De I, Steffen MD, Hong W, Lincoln H, Morrissy AS, Taylor MD, Akagi K, Brennan CW, Rodriguez FJ, Collier LS (2014) Sleeping Beauty mouse models identify candidate genes involved in gliomagenesis. PLoS ONE 9(11):e113489. https://doi.org/10.1371/journal.pone.0113489
Brun M, Coles JE, Monckton EA, Glubrecht DD, Bisgrove D, Godbout R (2009) Nuclear factor I regulates brain fatty acid-binding protein and glial fibrillary acidic protein gene expression in malignant glioma cell lines. J Mol Biol 391(2):282–300. https://doi.org/10.1016/j.jmb.2009.06.041
Glasgow SM, Zhu W, Stolt CC, Huang TW, Chen F, LoTurco JJ, Neul JL, Wegner M, Mohila C, Deneen B (2014) Mutual antagonism between Sox10 and NFIA regulates diversification of glial lineages and glioma subtypes. Nat Neurosci 17(10):1322–1329. https://doi.org/10.1038/nn.3790
Carlson BL, Pokorny JL, Schroeder MA, Sarkaria JN (2011) Establishment, maintenance and in vitro and in vivo applications of primary human glioblastoma multiforme (GBM) xenograft models for translational biology studies and drug discovery. Curr Protoc Pharmacol.https://doi.org/10.1002/0471141755.ph1416s52
Ponten J, Macintyre EH (1968) Long term culture of normal and neoplastic human glia. Acta Pathol Microbiol Scand 74(4):465–486
Stringer BW, Day BW, D'Souza RCJ, Jamieson PR, Ensbey KS, Bruce ZC, Lim YC, Goasdoue K, Offenhauser C, Akgul S, Allan S, Robertson T, Lucas P, Tollesson G, Campbell S, Winter C, Do H, Dobrovic A, Inglis PL, Jeffree RL, Johns TG, Boyd AW (2019) A reference collection of patient-derived cell line and xenograft models of proneural, classical and mesenchymal glioblastoma. Sci Rep 9(1):4902. https://doi.org/10.1038/s41598-019-41277-z
Bunt J, de Haas TG, Hasselt NE, Zwijnenburg DA, Koster J, Versteeg R, Kool M (2010) Regulation of cell cycle genes and induction of senescence by overexpression of OTX2 in medulloblastoma cell lines. Mol Cancer Res 8(10):1344–1357. https://doi.org/10.1158/1541-7786.MCR-09-0546
Huang da W, Sherman BT, Lempicki RA (2009) Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res 37(1):1–13. https://doi.org/10.1093/nar/gkn923
Chen F, LoTurco J (2012) A method for stable transgenesis of radial glia lineage in rat neocortex by piggyBac mediated transposition. J Neurosci Methods 207(2):172–180. https://doi.org/10.1016/j.jneumeth.2012.03.016
Venteicher AS, Tirosh I, Hebert C, Yizhak K, Neftel C, Filbin MG, Hovestadt V, Escalante LE, Shaw ML, Rodman C, Gillespie SM, Dionne D, Luo CC, Ravichandran H, Mylvaganam R, Mount C, Onozato ML, Nahed BV, Wakimoto H, Curry WT, Iafrate AJ, Rivera MN, Frosch MP, Golub TR, Brastianos PK, Getz G, Patel AP, Monje M, Cahill DP, Rozenblatt-Rosen O, Louis DN, Bernstein BE, Regev A, Suva ML (2017) Decoupling genetics, lineages, and microenvironment in IDH-mutant gliomas by single-cell RNA-seq. Science.https://doi.org/10.1126/science.aai8478
Lee J, Kotliarova S, Kotliarov Y, Li A, Su Q, Donin NM, Pastorino S, Purow BW, Christopher N, Zhang W, Park JK, Fine HA (2006) Tumor stem cells derived from glioblastomas cultured in bFGF and EGF more closely mirror the phenotype and genotype of primary tumors than do serum-cultured cell lines. Cancer Cell 9(5):391–403. https://doi.org/10.1016/j.ccr.2006.03.030
Cahoy JD, Emery B, Kaushal A, Foo LC, Zamanian JL, Christopherson KS, Xing Y, Lubischer JL, Krieg PA, Krupenko SA, Thompson WJ, Barres BA (2008) A transcriptome database for astrocytes, neurons, and oligodendrocytes: a new resource for understanding brain development and function. J Neurosci 28(1):264–278. https://doi.org/10.1523/JNEUROSCI.4178-07.2008
Lein ES, Hawrylycz MJ, Ao N, Ayres M, Bensinger A, Bernard A, Boe AF, Boguski MS, Brockway KS, Byrnes EJ, Chen L, Chen L, Chen TM, Chin MC, Chong J, Crook BE, Czaplinska A, Dang CN, Datta S, Dee NR, Desaki AL, Desta T, Diep E, Dolbeare TA, Donelan MJ, Dong HW, Dougherty JG, Duncan BJ, Ebbert AJ, Eichele G, Estin LK, Faber C, Facer BA, Fields R, Fischer SR, Fliss TP, Frensley C, Gates SN, Glattfelder KJ, Halverson KR, Hart MR, Hohmann JG, Howell MP, Jeung DP, Johnson RA, Karr PT, Kawal R, Kidney JM, Knapik RH, Kuan CL, Lake JH, Laramee AR, Larsen KD, Lau C, Lemon TA, Liang AJ, Liu Y, Luong LT, Michaels J, Morgan JJ, Morgan RJ, Mortrud MT, Mosqueda NF, Ng LL, Ng R, Orta GJ, Overly CC, Pak TH, Parry SE, Pathak SD, Pearson OC, Puchalski RB, Riley ZL, Rockett HR, Rowland SA, Royall JJ, Ruiz MJ, Sarno NR, Schaffnit K, Shapovalova NV, Sivisay T, Slaughterbeck CR, Smith SC, Smith KA, Smith BI, Sodt AJ, Stewart NN, Stumpf KR, Sunkin SM, Sutram M, Tam A, Teemer CD, Thaller C, Thompson CL, Varnam LR, Visel A, Whitlock RM, Wohnoutka PE, Wolkey CK, Wong VY, Wood M, Yaylaoglu MB, Young RC, Youngstrom BL, Yuan XF, Zhang B, Zwingman TA, Jones AR (2007) Genome-wide atlas of gene expression in the adult mouse brain. Nature 445(7124):168–176. https://doi.org/10.1038/nature05453
McKenzie AT, Wang M, Hauberg ME, Fullard JF, Kozlenkov A, Keenan A, Hurd YL, Dracheva S, Casaccia P, Roussos P, Zhang B (2018) Brain cell type specific gene expression and co-expression network architectures. Sci Rep 8(1):8868. https://doi.org/10.1038/s41598-018-27293-5
Glasgow SM, Laug D, Brawley VS, Zhang Z, Corder A, Yin Z, Wong ST, Li XN, Foster AE, Ahmed N, Deneen B (2013) The miR-223/nuclear factor I-A axis regulates glial precursor proliferation and tumorigenesis in the CNS. J Neurosci 33(33):13560–13568. https://doi.org/10.1523/JNEUROSCI.0321-13.2013
Lee JS, Xiao J, Patel P, Schade J, Wang J, Deneen B, Erdreich-Epstein A, Song HR (2014) A novel tumor-promoting role for nuclear factor IA in glioblastomas is mediated through negative regulation of p53, p21, and PAI1. Neuro-oncology 16(2):191–203. https://doi.org/10.1093/neuonc/not167
Chen KS, Lim JWC, Richards LJ, Bunt J (2017) The convergent roles of the nuclear factor I transcription factors in development and cancer. Cancer Lett 410:124–138. https://doi.org/10.1016/j.canlet.2017.09.015
Kang P, Lee HK, Glasgow SM, Finley M, Donti T, Gaber ZB, Graham BH, Foster AE, Novitch BG, Gronostajski RM, Deneen B (2012) Sox9 and NFIA coordinate a transcriptional regulatory cascade during the initiation of gliogenesis. Neuron 74(1):79–94. https://doi.org/10.1016/j.neuron.2012.01.024
Wong YW, Schulze C, Streichert T, Gronostajski RM, Schachner M, Tilling T (2007) Gene expression analysis of nuclear factor I-A deficient mice indicates delayed brain maturation. Genome Biol 8(5):R72. https://doi.org/10.1186/gb-2007-8-5-r72
Houillier C, Mokhtari K, Carpentier C, Criniere E, Marie Y, Rousseau A, Kaloshi G, Dehais C, Laffaire J, Laigle-Donadey F, Hoang-Xuan K, Sanson M, Delattre JY (2010) Chromosome 9p and 10q losses predict unfavorable outcome in low-grade gliomas. Neuro-oncology 12(1):2–6. https://doi.org/10.1093/neuonc/nop002
Genovesi LA, Ng CG, Davis MJ, Remke M, Taylor MD, Adams DJ, Rust AG, Ward JM, Ban KH, Jenkins NA, Copeland NG, Wainwright BJ (2013) Sleeping Beauty mutagenesis in a mouse medulloblastoma model defines networks that discriminate between human molecular subgroups. Proc Natl Acad Sci USA 110(46):E4325–E4334. https://doi.org/10.1073/pnas.1318639110
Wu X, Northcott PA, Dubuc A, Dupuy AJ, Shih DJ, Witt H, Croul S, Bouffet E, Fults DW, Eberhart CG, Garzia L, Van Meter T, Zagzag D, Jabado N, Schwartzentruber J, Majewski J, Scheetz TE, Pfister SM, Korshunov A, Li XN, Scherer SW, Cho YJ, Akagi K, MacDonald TJ, Koster J, McCabe MG, Sarver AL, Collins VP, Weiss WA, Largaespada DA, Collier LS, Taylor MD (2012) Clonal selection drives genetic divergence of metastatic medulloblastoma. Nature 482(7386):529–533. https://doi.org/10.1038/nature10825
Ho CY, Bar E, Giannini C, Marchionni L, Karajannis MA, Zagzag D, Gutmann DH, Eberhart CG, Rodriguez FJ (2013) MicroRNA profiling in pediatric pilocytic astrocytoma reveals biologically relevant targets, including PBX3, NFIB, and METAP2. Neuro-oncology 15(1):69–82. https://doi.org/10.1093/neuonc/nos269
Silber J, Lim DA, Petritsch C, Persson AI, Maunakea AK, Yu M, Vandenberg SR, Ginzinger DG, James CD, Costello JF, Bergers G, Weiss WA, Alvarez-Buylla A, Hodgson JG (2008) miR-124 and miR-137 inhibit proliferation of glioblastoma multiforme cells and induce differentiation of brain tumor stem cells. BMC Med 6:14. https://doi.org/10.1186/1741-7015-6-14
Acknowledgements
We thank the staff of the University of Queensland Biological Resources (UQBR) animal facility and the QBI Advanced Microscopy and Analysis Facility for their expertise and assistance in this project. We thank Rowan Tweedale for critical comments on the manuscript and Alan Ho for expert assistance with statistical analyses. We thank Andrew W. Boyd and Richard M. Gronostajski for their advice on this project. The primary human GBM samples and de-identified data used in this project were sourced from the Wesley Medical Research Tissue Bank with appropriate ethics approval.
Funding
This work was supported by the National Health and Medical Research Council (NHMRC) [GNT1100443, GNT1120615 to LJR]; Tour de Cure [Young Research Grant to JB]; Brain Foundation [research gift to JB]; Ride for Rhonda [research gift to LJR and JB to support CRB]; the University of Malaya [RP049-17HTM to HA]; the University of Queensland (UQ) [International Postgraduate Student Scholarship to KSC, UQ Graduate School Scholarship to ZL, UQ Centennial Scholarship to JWCL]; the Australian Government [Research Training Program Scholarship to JWCL].
Author information
Authors and Affiliations
Contributions
Study concept and design: KSC, LJR, JB. Acquisition of data: KSC, CRB, ZL, JB. Analysis and interpretation of data: KSC, CRB, ZL, JWCL, LJR, JB. GBM cell lines: BWS, BWD. Pathology: RR, KTW, DG, HA. Drafting of the manuscript: KSC, JB, LJR. Revision of the manuscript: KSC, JWCL, ZL, BWS, RR, KTW, DG, HA, BWD, LJR, JB. Administrative and technical support: CRB. Obtained funding: LJR, JB. Study supervision: LJR, JB.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
All procedures performed in studies involving human material were in accordance with the ethical standards of the institutional research committee (University of Queensland Human Ethics Committee) and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. All procedures performed in studies involving animals were in accordance with the Australian Code of Practice for the Care and Use of Animals for Scientific Purposes, and with the approval of the University of Queensland Animal Ethics Committee.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chen, KS., Bridges, C.R., Lynton, Z. et al. Transcription factors NFIA and NFIB induce cellular differentiation in high-grade astrocytoma. J Neurooncol 146, 41–53 (2020). https://doi.org/10.1007/s11060-019-03352-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-019-03352-3