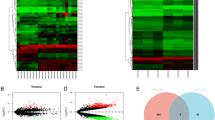
Abstract
Recent trends are moving towards the use of the circulating transcriptome as a potential diagnostic and therapeutic tool for hepatocellular carcinoma (HCC). The aim of this study is to identify circulatory RNA based biomarker panel, in addition to their relationship to the outcome in HCC. First, utilizing bioinformatics tools, we selected an HCC-specific RNA-based biomarker panel that depended on the integration of suppressor of glucose autophagy-associated (SOGA1) gene expression with the chosen panel of epigenetic regulators of this gene [long non-coding RNA antisense for X-inactive-specific transcript (lncRNA-TSIX) and microRNA-548-a-3p (miR-548-a-3p)]. Second, we attempted to validate these biomarkers using the sera of 65 patients with HCC, 34 patients with chronic hepatitis C virus (CHC) infection and 32 healthy volunteers. Finally, the expression levels of the chosen RNA-based biomarker panel were assessed in the serum samples using qRT-PCR assays. The panel of 3 RNA-based biomarkers (lncRNA-TSIX, miR-548-a-3p, and SOGA1) exhibited high sensitivity and specificity in differentiating HCC patients from CHC patients and healthy controls. Among these 3 RNAs, serum lncRNA-TSIX and SOGA1 were independent prognostic factor. The chosen circulatory RNA-based biomarker panel may serve as a diagnostic and prognostic biomarker for HCC.




Similar content being viewed by others
Change history
02 May 2019
The original publication has been updated. The family name of Alaa Habieb contained a typo that has now been corrected.
Abbreviations
- ALT:
-
Alanine transaminase
- AST:
-
Aspartate transaminase
- AFP:
-
Alpha fetoprotein
- CHC:
-
Chronic HCV
- ceRNA:
-
Competing endogenous RNA
- HCC:
-
Hepatocellular carcinoma
- lncRNA-TSIX:
-
Long non-coding RNA antisense for X-inactive-specific transcript
- miRNA, miR:
-
Micro ribonucleic acid
- qRT-PCR:
-
Quantitative reverse-transcriptase polymerase chain reaction
- SOGA1:
-
Suppressor of glucose autophagy-associated
References
American Cancer Society (2018) Cancer facts & figures 2018. American Cancer Society, Atlanta
Zeeneldin A, Salem S, Darwish A, El-Gammal M, Hussein M, Saadeldin M (2015) Untreated hepatocellular carcinoma in Egypt: outcome and prognostic factors. J Hepatocell Carcinoma 2:3–9
Abdelgawad IA, Mossallam GI, Radwan NH, Elzawahry HM, Elhifnawy NM (2013) Can Glypican3 be diagnostic for early hepatocellular carcinoma among Egyptian patients? Asian Pac J Cancer Prev 14(12):7345–7349
Stewart BW, Wild CP (2017) Liver cancer. World Cancer Report 2014. IARC
He Y, Meng XM, Huang C, Wu BM, Zhang L, Lv XW et al (2014) Long noncoding RNAs: novel insights into hepatocelluar carcinoma. Cancer Lett 344(1):20–27
Lamb R, Harrison H, Hulit J, Smith DL, Lisanti MP, Sotgia F (2014) Mitochondria as new therapeutic targets for eradicating cancer stem cells: quantitative proteomics and functional validation via MCT1/2 inhibition. Oncotarget 5(22):11029–11037. https://doi.org/10.18632/oncotarget.2789
Kruse R, Krantz J, Barker N, Coletta RL, Rafikov R, Luo M, Højlund K, Mandarino LJ, Langlais PR (2017) Characterization of the CLASP2 protein interaction network identifies SOGA1 as a microtubule-associated protein. Mol Cell Proteomics 16(10):1718–1735. https://doi.org/10.1074/mcp.RA117.000011
Grace M, Munger K (2017) Proteomic analysis of the gamma human papillomavirus type 197 E6 and E7 associated cellular proteins. Virology 500:71–81
Mattick JS, Makunin IV (2006) Non-coding RNA. Hum Mol Genet 15(Spec No 1):R17–R29
Lei B, Liang XS, Li YW, Zhang JF, Zhang GQ, Pang D (2015) Long non-coding RNA MVIH is associated with poor prognosis and malignant biological behavior in breast cancer. Tumour Biol 37(4):5257–5264
Li JH, Liu S, Zhou H, Qu LH, Yang JH (2014) starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res 42:D92–D97
Jiao F, Hu H, Yuan C et al (2014) Elevated expression level of long noncoding RNA MALAT-1 facilitates cell growth, migration and invasion in pancreatic cancer. Oncol Rep 32:2485–2492
Ren S, Liu Y, Xu W et al (2013) Long noncoding RNA MALAT-1 is a new potential therapeutic target for castration resistant prostate cancer. J Urol 190:2278–2287
Cesana M, Cacchiarelli D, Legnini I, Santini T, Sthandier O, Chinappi M, Tramontano A, Bozzoni I (2011) A long noncoding RNA controls muscle differentiation by functioning as a competing endogenous RNA. Cell 147:358–369
Li H, Yu B, Li J, Su L, Yan M, Zhu Z et al (2014) Overexpression of lncRNA H19 enhances carcinogenesis and metastasis of gastric cancer. Oncotarget. 5:2318–2329
Tsang WP, Ng EK, Ng SS, Jin H, Yu J, Sung JJ et al (2010) Oncofetal H19-derived miR-675 regulates tumor suppressor RB in human colorectal cancer. Carcinogenesis 31:350–358
Reis EM, Nakaya HI, Louro R, Canavez FC, Flatschart AV, Almeida GT et al (2004) Antisense intronic non-coding RNA levels correlate to the degree of tumor differentiation in prostate cancer. Oncogene 23:6684–6692
Wang P, Ning S, Zhang Y, Li R, Ye J, Zhao Z, Zhi H, Wang T, Guo Z, Li X (2015) Identification of lncRNA-associated competing triplets reveals global patterns and prognostic markers for cancer. Nucl Acids Res 43(7):3478–3489
Weakley SM, Wang H, Yao Q, Chen C (2011) Expression and function of a large non-coding RNA gene XIST in human cancer. World J Surg 35(8):1751–1756. https://doi.org/10.1007/s00268-010-0951-0
Ohhata T, Senner CE, Hemberger M, Wutz A (2011) Lineage-specific function of the noncoding Tsix RNA for Xist repression and Xi reactivation in mice. Genes Dev 25(16):1702–1715. https://doi.org/10.1101/gad.16997911
Lee JT, Lu N (1999) Targeted mutagenesis of Tsix leads to nonrandom X inactivation. Cell 99(1):47–57
Wang Y, Zheng F, Gao G, Yan S, Zhang L, Wang L, Cai X, Wang X, Xu D, Wang J (2018) MiR-548a-3p regulates inflammatory response via TLR4/NF-κB signaling pathway in rheumatoid arthritis. J Cell Biochem 120(2):1133–1140
Hu B, Ying X, Wang J, Piriyapongsa J, Jordan IK, Sheng J, Yu F, Zhao P, Li Y, Wang H et al (2015) Identification of a tumor-suppressive human-specific microRNA within the FHIT tumor-suppressor gene. Cancer Res 74(8):2283–2294
Shi Y, Qiu M, Wu Y, Hai L (2015) MiR-548-3p functions as an anti-oncogenic regulator in breast cancer. Biomed Pharmacother 75:111–116. https://doi.org/10.1016/j.biopha.2015.07.027
Murakami Y, Yasuda T, Saigo K et al (2006) Comprehensive analysis of microRNA expression patterns in hepatocellular carcinoma and non-tumorous tissues. Oncogene 25:2537–2545
Zhao X, Zhen Y, Li G, Li D, Zhao Y, Wu Y, Robson S, He L, Xu Y, Miao R, Zhao HT (2012) The role and clinical implications of microRNAs in hepatocellular carcinoma. Sci China Life Sci 55(10):906–919
Farazi TA, Spitzer JI, Morozov P, Tuschl T (2011) miRNAs in human cancer. J Pathol 223:102–115
Ezzat H, Lotfy AM, Alalfy MN, El-Taher SM, Mokhtar A, Mohamed SA, El-Senosy FM (2014) The significance of circulating MicroRNA-122as a non invasive diagnostic marker of liver injury in Egyptian chronic hepatitis C virus infected and cirrhotic patients with and without hepatocellular carcinoma. Clin Med Diagnost. 4(1):1–8
Zhang Y, Cai S, Jia Y et al (2017) Decoding noncoding RNAs: role of microRNAs and long noncoding RNAs in ocular neovascularization. Theranostics 7(12):3155–3167. https://doi.org/10.7150/thno.19646
Child CG, Turcotte JG (1964) Surgery and portal hypertension. Major Probl Clin Surg 1:1–85
Pugh RN, Murray-Lyon IM, Dawson JL et al (1973) Transection of the oesophagus for bleeding oesophageal varices. Br J Surg 60:646–649
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-delta deltaC(T)) method. Methods 25:402–408
Jiang H, Dong Q, Luo X et al (2014) The monoclonal antibody CH12 augments 5-fluorouracil-induced growth suppression of hepatocellular carcinoma xenografts expressing epidermal growth factor receptor variant III. Cancer Lett 342:113–120
Panzitt K, Tschernatsch MM, Guelly C, Moustafa T, Stradner M, Strohmaier HM, Buck C, Denk H, Schroeder R, Trauner M et al (2007) Characterization of HULC, a novel gene with striking up-regulation in hepatocellular carcinoma, as noncoding RNA. Gastroenterology 132:330–342
Cowherd RB, Asmar MM, Alderman JM, Alderman EA, Garland AL, Busby WH et al (2010) Adiponectin lowers glucose production by increasing SOGA. Am J Pathol 177(4):1936–1945
Longnus SL, Wambolt RB, Parsons HL, Brownsey RW, Allard MF (2003) 5-Aminoimidazole-4-carboxamide 1-beta -D-ribofuranoside (AICAR) stimulates myocardial glycogenolysis by allosteric mechanisms. Am J Physiol Regul Integr Comp Physiol 284(4):R936–R944
Camacho RC, Pencek RR, Lacy DB, James FD, Donahue EP, Wasserman DH (2005) Portal venous 5-aminoimidazole-4-carboxamide-1-beta-D-ribofuranoside infusion overcomes hyperinsulinemic suppression of endogenous glucose output. Diabetes 54(2):373–382
Combs Terry P, Marliss Errol B (2014) Adiponectin signaling in the liver. Rev Endocr Metab Disord 15(2):137–147
Combs TP, Snell-Bergeon JK, Maahs DM, Bergman BC, Lamarche M, Iberkleid L, AbdelBaky O, Tisch R, Scherer PE, Marliss EB (2015) Adiponectin-SOGA dissociation in type 1 diabetes. J Clin Endocrinol Metab 100(8):E1065–E1073
Shi Y, Qiu M, Wu Y, Hai L (2015) MiR-548-a-3p-3p functions as an anti-oncogenic regulator in breast cancer. Biomed Pharmacother 75:111–116
Liu WH, Tao KS, You N, Liu ZC, Zhang HT, Dou KF (2011) Differences in the properties and mirna expression profiles between side populations from hepatic cancer cells and normal liver cells. PLoS ONE 6:e23311
Khokhar A, Noorali S, Sheraz M, Mahalingham K, Pace DG, Khanani MR, Bagasra O (2012) Computational analysis to predict functional role of hsa-miR-3065-3p as an antiviral therapeutic agent for treatment of triple infections: HCV, HIV-1, and HBV. Libyan J Med 7:19774
Li Y, Xie J, Xu X, Wang J, Ao F, Wan Y, Zhu Y (2013) MicroRNA-548-a-3p down-regulates host antiviral response via direct targeting of IFN-λ1. Protein Cell 4:130–141
Khokhar A, Noorali S, Sheraz M, Mahalingham K, Pace DG, Khanani MR, Bagasra O (2015) Computational analysis to predict functional role of hsa-miR-3065-3p as an antiviral therapeutic agent for treatment of triple infections: HCV, HIV-1, and HBV. Libyan J Med 7:19774
Gayen S, Maclary E, Buttigieg E, Hinten M, Kalantry S (2015) A primary role for the Tsix lncRNA in maintaining random X-chromosome inactivation. Cell Rep 11(8):1251–1265
Weakley SM, Wang H, Yao Q et al (2011) Expression and function of a large non-coding RNA gene XIST in human cancer. World J Surg 35:1751–1756
Wang Zhongzhi, Jinnin Masatoshi, Nakamura Kayo, Harada Miho, Kudo Hideo, Nakayama Wakana, Inoue Kuniko, Nakashima Taiji, Honda Noritoshi, Fukushima Satoshi, Ihn Hironobu (2016) Long non-coding RNA TSIX is upregulated in scleroderma dermal fibroblasts and controls collagen mRNA stabilization. Exp Dermatol 25(2):131–136
Karreth FA, Pandolfi PP (2013) ceRNA cross-talk in cancer: when ce-bling rivalries go awry. Cancer Discov 3:1113–1121
Abd EL Gwad A, Matboli M, El-Tawdi A, Habib EK, Shehata H, Ibrahim D, Tash F (2018) Role of exosomal competing endogenous RNA in patients with hepatocellular carcinoma. J Cell Biochem 119(10):8600–8610
Zhu HQ, Zhou X, Chang H, Li HG, Liu FF, Ma CQ, Lu J (2015) Aberrant expression of CCAT1 regulated by c-Myc predicts the prognosis of hepatocellular carcinoma. Asian Pac J Cancer Prev 16:5181–5185
Ge Y, Yan X, Jin Y, Yang X, Yu X, Zhou L, Han S, Yuan Q, Yang M (2015) MiRNA-192 [corrected] and miRNA-204 directly suppress lncRNA HOTTIP and interrupt GLS1-mediated glutaminolysis in hepatocellular carcinoma. PLoS Genet 11:e1005726
Acknowledgement
The manuscript was edited for English language usage, grammar, spelling and punctuation by one or more native English-speaking editors at Nature Research Editing Service (CERTIFICATE 8A4A-0338-9D25-FE77-368P). The author acknowledge dr Sara El-Nakeep; Clinical Tropical Medicine Department, Ain Shams University Hospital, for her effort in clinical samples provision.
Funding
This work was supported by the Egyptian Science and Technology Development Fund (RSTDG 2259).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Alaa Habib, Marwa Matboli, Hanaa El-Tayeb, and Farid El-Asmar declares that they have no competing interests.
Ethical approval and informed consent
Written informed consent was obtained from all the participants in this study, which was performed in accordance with the Declaration of Helsinki and was approved by the Ethics Committee of Ain Shams Faculty of Medicine, Egypt (Ethical Approval Number; FN000023408).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original version of this article was revised: The family name of Alaa Habieb contained a typo that has now been corrected.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Habieb, A., Matboli, M., El-Tayeb, H. et al. Potential role of lncRNA-TSIX, miR-548-a-3p, and SOGA1 mRNA in the diagnosis of hepatocellular carcinoma. Mol Biol Rep 46, 4581–4590 (2019). https://doi.org/10.1007/s11033-019-04810-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-019-04810-x