Abstract
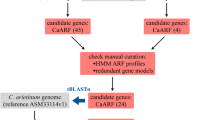
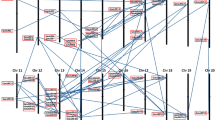
Auxin response factors (ARFs), member of the plant-specific B3 DNA binding superfamily, target specifically to auxin response elements (AuxREs) in promoters of primary auxin-responsive genes and heterodimerize with Aux/IAA proteins in auxin signaling transduction cascade. In previous research, we have isolated and characterized maize Aux/IAA genes in whole-genome scale. Here, we report the comprehensive analysis of ARF genes in maize. A total of 36 ARF genes were identified and validated from the B73 maize genome through an iterative strategy. Thirty-six maize ARF genes are distributed in all maize chromosomes except chromosome 7. Maize ARF genes expansion is mainly due to recent segmental duplications. Maize ARF proteins share one B3 DNA binding domain which consists of seven-stranded β sheets and two short α helixes. Twelve maize ARFs with glutamine-rich middle regions could be as activators in modulating expression of auxin-responsive genes. Eleven maize ARF proteins are lack of homo- and heterodimerization domains. Putative cis-elements involved in phytohormones and light signaling responses, biotic and abiotic stress adaption locate in promoters of maize ARF genes. Expression patterns vary greatly between clades and sister pairs of maize ARF genes. The B3 DNA binding and auxin response factor domains of maize ARF proteins are primarily subjected to negative selection during selective sweep. The mixed selective forces drive the diversification and evolution of genomic regions outside of B3 and ARF domains. Additionally, the dicot-specific proliferation of ARF genes was detected. Comparative genomics analysis indicated that maize, sorghum and rice duplicate chromosomal blocks containing ARF homologs are highly syntenic. This study provides insights into the distribution, phylogeny and evolution of ARF gene family.






Similar content being viewed by others
References
Woodward AW, Bartel B (2005) Auxin: regulation, action, and interaction. Ann Bot 95:707–735. doi:10.1093/aob/mci083
Fleming AJ (2006) Plant signalling: the inexorable rise of auxin. Trends Cell Biol 16:397–402. doi:10.1016/j.tcb.2006.06.005
Abel S, Theologis A (1996) Early genes and auxin action. Plant Physiol 111:9–17. doi:10.1104/pp.111.1.9
Liscum E, Reed JW (2002) Genetics of Aux/IAA and ARF action in plant growth and development. Plant Mol Biol 49:387–400. doi:10.1023/A:1015255030047
Fukaki H, Taniguchi M, Tasaka M (2006) PICKLE is required for SOLITARY ROOT/IAA14-mediated repression of ARF7 and ARF19 activity during Arabidopsis lateral root initiation. Plant J 48:380–389. doi:10.1111/j.1365-313X.2006.02882.x
Ulmasov T, Hagen G, Guilfoyle TJ (1997) ARF1, a transcription factor that binds to auxin response elements. Science 276:1865–1868. doi:10.1126/science.276.5320.1865
Guilfoyle TJ, Hagen G (2001) Auxin response factors. J Plant Growth Reg 20:281–291. doi:10.1007/s003440010026
Tiwari SB, Hagen G, Guilfoyle TJ (2003) The roles of auxin response factor domains in auxin-responsive transcription. Plant Cell 15:533–543. doi:10.1105/tpc.008417
Swaminathan K, Peterson K, Jack T (2008) The plant B3 superfamily. Trends Plant Sci 13:647–655. doi:10.1016/j.tplants.2008.09.00
Salmon J, Ramos J, Callis J (2008) Degradation of the auxin response factor ARF1. Plant J 54:118–128. doi:10.1111/j.1365-313X.2007.03396.x
Shen C, Wang S, Bai Y, Wu Y, Zhang S, Chen M, Guilfoyle TJ, Wu P, Qi Y (2010) Functional analysis of the structural domain of ARF proteins in rice (Oryza sativa L.). J Exp Bot 61:3971–3981. doi:10.1093/jxb/erq208
Schruff MC, Spielman M, Tiwari S, Adams S, Fenby N, Scott RJ (2006) The AUXIN RESPONSE FACTOR 2 gene of Arabidopsis links auxin signalling, cell division, and the size of seeds and other organs. Development 133:251–261. doi:10.1242/dev.02194
Sessions A, Nemhauser JL, McColl A, Roe JL, Feldmann KA, Zambryski PC (1997) ETTIN patterns the Arabidopsis floral meristem and reproductive organs. Development 124:4481–4491
Hardtke CS, Berleth T (1998) The Arabidopsis gene MONOPTEROS encodes a transcription factor mediating embryo axis formation and vascular development. EMBO J 17:1405–1411. doi:10.1093/emboj/17.5.1405
Harper M, Stowe-Evans EL, Luesse DR, Muto H, Tatematsu K, Watahiki MK, Yamamoto K, Liscum E (2000) The NPH4 locus encodes the auxin response factor ARF7, a conditional regulator of differential growth in aerial Arabidopsis tissue. Plant Cell 12:757–770. doi:10.1105/tpc.12.5.757
Goetz M, Vivian-Smith A, Johnson SD, Koltunow AM (2006) AUXIN RESPONSE FACTOR8 is a negative regulator of fruit initiation in Arabidopsis. Plant Cell 18:1873–1886. doi:10.1105/tpc.105.037192
Li J, Dai X, Zhao Y (2006) A role for auxin response factor 19 in auxin and ethylene signaling in Arabidopsis. Plant Physiol 140:899–908. doi:10.1104/pp.105.070987
Okushima Y, Overvoorde PJ, Arima K, Alonso JM, Chan A, Chang C, Ecker JR, Hughes B, Lui A, Nguyen D, Onodera C, Quach H, Smith A, Yu G, Theologis A (2005) Functional genomic analysis of the AUXIN RESPONSE FACTOR gene family members in Arabidopsis thaliana: unique and overlapping functions of ARF7 and ARF19. Plant Cell 17:444–463. doi:10.1105/tpc.104.028316
Wilmoth JC, Wang S, Tiwari SB, Joshi AD, Hagen G, Guilfoyle TJ, Alonso JM, Ecker JR, Reed JW (2005) NPH4/ARF7 and ARF19 promote leaf expansion and auxin-induced lateral root formation. Plant J 43:118–130. doi:10.1111/j.1365-313X.2005.02432.x
Wang JW, Wang LJ, Mao YB, Cai WJ, Xue HW, Chen XY (2005) Control of root cap formation by microRNA-targeted auxin response factors in Arabidopsis. Plant Cell 17:2204–2216. doi:10.1105/tpc.105.033076
Wang D, Pei K, Fu Y, Sun Z, Li S, Liu H, Tang K, Han B, Tao Y (2007) Genome-wide analysis of the auxin response factor (ARF) gene family in rice (Oryza sativa). Gene 394:13–24. doi:10.1016/j.gene.2007.01.006
Kalluri UC, Difazio SP, Brunner AM, Tuskan GA (2007) Genome-wide analysis of Aux/IAA and ARF gene families in Populus trichocarpa. BMC Plant Biol 7:59. doi:10.1186/1471-2229-7-59
Schnable PS, Ware D, Fulton RS, Stein JC, Wei F et al (2009) The B73 maize genome: complexity, diversity, and dynamics. Science 326:1112–1115. doi:10.1126/science.1178534
Meyers BC, Kozik A, Griego A, Kuang H, Michelmore RW (2003) Genome-wide analysis of NBS-LRR-encoding genes in Arabidopsis. Plant Cell 15:809–834. doi:10.1105/tpc.009308
Finn RD, Mistry J, Tate J, Coggill P, Heger A, Pollington JE, Gavin OL, Gunasekaran P, Ceric G, Forslund K, Holm L, Sonnhammer EL, Eddy SR, Bateman A (2010) The Pfam protein families database. Nucleic Acids Res 38:D211–D222. doi:10.1093/nar/gkp985
Eddy SR (2008) A probabilistic model of local sequence alignment that simplifies statistical significance estimation. PLoS Comput Biol 4:e1000069. doi:10.1371/journal.pcbi.1000069
Letunic I, Copley RR, Schmidt S, Ciccarelli FD, Doerks T, Schultz J, Ponting CP, Bork P (2004) SMART 4.0: towards genomic data integration. Nucleic Acids Res 32:D142–D144. doi:10.1093/nar/gkh088
Marchler-Bauer A, Anderson JB, Chitsaz F, Derbyshire MK, DeWeese-Scott C et al (2009) CDD: specific functional annotation with the Conserved Domain Database. Nucleic Acids Res 37:D205–D210. doi:10.1093/nar/gkn845
Hunter S, Apweiler R, Attwood TK, Bairoch A, Bateman A et al (2009) InterPro: the integrative protein signature database. Nucleic Acids Res 37:D211–D215. doi:10.1093/nar/gkn785
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948. doi:10.1093/bioinformatics/btm404
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) Software Version 4.0. Mol Biol Evol 24:1596–1599. doi:10.1093/molbev/msm092
Bailey TL, Boden M, Buske FA, Frith M, Grant CE, Clementi L, Ren J, Li WW, Noble WS (2009) MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res 37:W202–W208. doi:10.1093/nar/gkp335
Sigrist CJ, Cerutti L, de Castro E, Langendijk-Genevaux PS, Bulliard V, Bairoch A, Hulo N (2010) PROSITE, a protein domain database for functional characterization and annotation. Nucleic Acids Res 38:D161–D166. doi:10.1093/nar/gkp885
Crooks GE, Hon G, Chandonia JM, Brenner SE (2004) WebLogo: a sequence logo generator. Genome Res 14:1188–1190. doi:10.1101/gr.849004
Kiefer F, Arnold K, Künzli M, Bordoli L, Schwede T (2009) The SWISS-MODEL repository and associated resources. Nucleic Acids Res 37:D387–D392. doi:10.1093/nar/gkn750
Lescot M, Déhais P, Thijs G, Marchal K, Moreau Y, Van de Peer Y, Rouzé P, Rombauts S (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30:325–327. doi:10.1093/nar/30.1.325
Nakano M, Nobuta K, Vemaraju K, Tej SS, Skogen JW, Meyers BC (2006) Plant MPSS databases: signature-based transcriptional resources for analyses of mRNA and small RNA. Nucleic Acids Res 34:D731–D735. doi:10.1093/nar/gkj077
Maher C, Stein L, Ware D (2006) Evolution of Arabidopsis microRNA families through duplication events. Genome Res 16:510–519. doi:10.1101/gr.4680506
Ouyang Y, Chen J, Xie W, Wang L, Zhang Q (2009) Comprehensive sequence and expression profile analysis of Hsp20 gene family in rice. Plant Mol Biol 70:341–357. doi:10.1007/s11103-009-9477-y
Blanc G, Wolfe KH (2004) Widespread paleopolyploidy in model plant species inferred from age distributions of duplicate genes. Plant Cell 16:1667–1678. doi:10.1105/tpc.021345
Gaut BS, Morton BR, McCaig BC, Clegg MT (1996) Substitution rate comparisons between grasses and palms: synonymous rate differences at the nuclear gene Adh parallel rate differences at the plastid gene rbcL. Proc Natl Acad Sci USA 93:10274–10279
Suyama M, Torrents D, Bork P (2006) PAL2NAL: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res 34:W609–W612. doi:10.1093/nar/gkl315
Comeron JM (1999) K-Estimator: calculation of the number of nucleotide substitutions per site and the confidence intervals. Bioinformatics 15:763–764. doi:10.1093/bioinformatics/15.9.763
Liu Y, Jiang HY, Chen W, Qian Y, Ma Q, Cheng B, Zhu S (2011) Genome-wide analysis of the auxin response factor (ARF) gene family in maize (Zea mays). Plant Growth Regul 63:225–234. doi:10.1007/s10725-010-9519-0
Mondragón-Palomino M, Meyers BC, Michelmore RW, Gaut BS (2002) Patterns of positive selection in the complete NBS-LRR gene family of Arabidopsis thaliana. Genome Res 12:1305–1315. doi:10.1101/gr.159402
Romanel EA, Schrago CG, Couñago RM, Russo CA, Alves-Ferreira M (2009) Evolution of the B3 DNA binding superfamily: new insights into REM family gene diversification. PLoS ONE 4:e5791. doi:10.1371/journal.pone.0005791
Guilfoyle TJ, Hagen G (2007) Auxin response factors. Curr Opin Plant Biol 10:453–460. doi:10.1016/j.pbi.2007.08.014
Demuth JP, Hahn MW (2009) The life and death of gene families. Bioessays 31:29–39. doi:10.1002/bies.080085
Colón-Carmona A, Chen DL, Yeh KC, Abel S (2000) Aux/IAA proteins are phosphorylated by phytochrome in vitro. Plant Physiol 124:1728–1738. doi:10.1104/pp.124.4.1728
Park CM (2007) Auxin homeostasis in plant stress adaptation response. Plant Signal Behav 2:306–307. doi:10.1074/jbc.M610524200
Jain M, Khurana JP (2009) Transcript profiling reveals diverse roles of auxin-responsive genes during reproductive development and abiotic stress in rice. FEBS J 276:3148–3162. doi:10.1111/j.1742-4658.2009.07033.x
Kazan K, Manners JM (2009) Linking development to defense: auxin in plant–pathogen interactions. Trends Plant Sci 14:373–382. doi:10.1016/j.tplants.2009.04.005
Vert G, Walcher CL, Chory J, Nemhauser JL (2008) Integration of auxin and brassinosteroid pathways by Auxin Response Factor 2. Proc Natl Acad Sci USA 105:9829–9834. doi:10.1073/pnas.0803996105
Song Y, You J, Xiong L (2009) Characterization of OsIAA1 gene, a member of rice Aux/IAA family involved in auxin and brassinosteroid hormone responses and plant morphogenesis. Plant Mol Biol 70:297–309. doi:10.1007/s11103-009-9474-1
Ulmasov T, Hagen G, Guilfoyle TJ (1999) Dimerization and DNA binding of auxin response factors. Plant J 19:309–319. doi:10.1046/j.1365-313X.1999.00538.x
Wenzel CL, Schuetz M, Yu Q, Mattsson J (2007) Dynamics of MONOPTEROS and PIN-FORMED1 expression during leaf vein pattern formation in Arabidopsis thaliana. Plant J 49:387–398. doi:10.1111/j.1365-313X.2006.02977.x
Tian CE, Muto H, Higuchi K, Matamura T, Tatematsu K, Koshiba T, Yamamoto KT (2004) Disruption and overexpression of auxin response factor 8 gene of Arabidopsis affect hypocotyl elongation and root growth habit, indicating its possible involvement in auxin homeostasis in light condition. Plant J 40:333–343. doi:10.1111/j.1365-313X.2004.02220.x
Ellis CM, Nagpal P, Young JC, Hagen G, Guilfoyle TJ, Reed JW (2005) AUXIN RESPONSE FACTOR1 and AUXIN RESPONSE FACTOR2 regulate senescence and floral organ abscission in Arabidopsis thaliana. Development 132:4563–4574. doi:10.1242/dev.02012
Wu MF, Tian Q, Reed JW (2006) Arabidopsis microRNA167 controls patterns of ARF6 and ARF8 expression, and regulates both female and male reproduction. Development 133:4211–4218. doi:10.1242/dev.02602
Hunter C, Willmann MR, Wu G, Yoshikawa M, de la Luz Gutierrez-Nava M, Poethig SR (2006) Trans-acting siRNA-mediated repression of ETTIN and ARF4 regulates heteroblasty in Arabidopsis. Development 133:2973–2981. doi:10.1242/dev.02491
Marin E, Jouannet V, Herz A, Lokerse AS, Weijers D, Vaucheret H, Nussaume L, Crespi MD, Maizel A (2010) miR390, Arabidopsis TAS3 tasiRNAs, and their AUXIN RESPONSE FACTOR targets define an autoregulatory network quantitatively regulating lateral root growth. Plant Cell 22:1104–1117. doi:10.1105/tpc.109.072553
Remington DL, Vision TJ, Guilfoyle TJ, Reed JW (2004) Contrasting modes of diversification in the Aux/IAA and ARF gene families. Plant Physiol 135:1738–1752. doi:10.1104/pp.104.039669
Wang Y, Deng D, Bian Y, Lv Y, Xie Q (2010) Genome-wide analysis of primary auxin-responsive Aux/IAA gene family in maize (Zea mays L). Mol Biol Rep 37:3991–4001. doi:10.1007/s11033-010-0058-6
Ameline-Torregrosa C, Wang BB, O’Bleness MS, Deshpande S, Zhu H, Roe B, Young ND, Cannon SB (2008) Identification and characterization of nucleotide-binding site-leucine-rich repeat genes in the model plant Medicago truncatula. Plant Physiol 146:5–21. doi:10.1104/pp.107.104588
Chaw SM, Chang CC, Chen HL, Li WH (2004) Dating the monocot-dicot divergence and the origin of core eudicots using whole chloroplast genomes. J Mol Evol 58:424–441. doi:10.1007/s00239-003-2564-9
Acknowledgments
We are grateful to editors and reviewers for their helpful comments. This work is supported partly by the Nature Science Foundation of Universities in Jiangsu Province (No. 09KJB180010) and the High-Level Personnel Foundation of Yangzhou University (No. nxy5286).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
11033_2011_991_MOESM6_ESM.tif
Fig. S1 Phylogeny of maize and rice ARF proteins. Ten sister pairs of maize ARF genes were emphasized in red. Scale bar 0.05 showed 0.05 amino acid substitution per site (TIFF 240 kb)
11033_2011_991_MOESM9_ESM.tif
Fig. S4 Heat map of expression patterns of ten sister pairs of maize ARF genes. SR, seedling root; SS, seedling shoot; R: root; L, leaf; T, tassel; P: pollen; E: ear 9 (TIFF 175 kb)
11033_2011_991_MOESM10_ESM.tif
Fig. S5 Phylogenesis of maize and sorghum ARF proteins. Evolutional branches of ten sister pairs of maize ARF genes except ZmARF1/ZmARF2 and their sorghum orthologs were marked in green 10 (TIFF 70 kb)
Rights and permissions
About this article
Cite this article
Wang, Y., Deng, D., Shi, Y. et al. Diversification, phylogeny and evolution of auxin response factor (ARF) family: insights gained from analyzing maize ARF genes. Mol Biol Rep 39, 2401–2415 (2012). https://doi.org/10.1007/s11033-011-0991-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-011-0991-z