Abstract
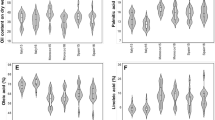
The need for new improved cultivars adapted to modern cultivation techniques and changing climatic environment requires genetic studies able to provide a better understanding of the phenotypic variance decomposition into genetic and environmental variances. In perennial crops, this decomposition requires evaluations carried out among years, trees within a genotype, samples within a tree and their interactions. The aim of this study was to perform such a detailed variance analysis of oil content components (fruit fresh weight-FFW, fruit moisture-FM, oil content on both fresh-OCFFW and dry-OCFDW basis) and fruit morphology traits (fruit maximum diameter-FMD and fruit circularity-FC) in an olive progeny issued from the cross ‘Oliviere’ × ‘Arbequina’. A total of 106 genotypes from this progeny, together with the parents cultivars, were evaluated over two consecutive years in two trees/genotype, and three samples/tree for oil content components and 30 samples/tree for fruit morphology traits. Two dataset were considered during analysis. The first one, including a limited number of genotypes without any missing data, was fully balanced and ANOVA was applied for variance components estimation from expected mean squares. The second dataset contained the full unbalanced information of the 106 genotypes and mixed linear models were built following to the same statistical model. Both analyses showed that variance among genotypes (σ 2G ) was the main contributor to total variance for FFW, OCFFW and OCFDW and variance among years (σ 2Y ) for FM. Residual error effect (σ 2ε ) was the main contributor for morphology traits, whereas variance associated with the genotype × year interaction (σ 2GY ) and tree × year interaction (σ 2TY ) represented also important variance components for most traits, and variance among trees within a genotype (σ 2T ) was negligible. Consistently, the mean broad sense heritabilities (H²) showed the lowest values for FM (0.40) and the highest values for OCFFW (0.94). An estimation of heritability and environmental variation under varying numbers of years and tree replications revealed that including more yearly replications could be more effective than adding trees replications. Finally, this study provides a global view of adequate spatial and temporal replications required to accurately develop criteria for both conventional and marker-assisted selection on fruit traits from olive crossing progenies, and more generally in fruit trees.



Similar content being viewed by others
References
Arias-Calderón R, Rouiss H, Rodríguez-Jurado D, De la Rosa R, León L (2014) Variability and heritability of fruit characters in olive progenies from open-pollination. Sci Hortic 169:94–98
Atienza SG, De la Rosa R, León L, Martín A, Belaj A (2014) Identification of QTL for agronomic traits of importance for olive breeding. Mol Breed 34:725–737
Badenes ML, Byrne DH (2011) Fruit breeding. Springer Science+Business Media, New York
Bartholome J, Salmon F, Vigneron P, Bouvet JM, Plomion C, Gion JM (2013) Plasticity of primary and secondary growth dynamics in Eucalyptus hybrids: a quantitative genetics and QTL mapping perspective. BMC Plant Biol 13:14
Belaj A, León L, Satovic Z, De la Rosa R (2011) Variability of wild olives (Olea europaea subsp europaea var. sylvestris) analyzed by agro-morphological traits and SSR markers. Sci Hortic 129:561–569
Bellini E (1992) Behaviour of some genetic characters in olive seedlings obtained by cross-breeding. Acta Hortic 317:197–208
Bellini E, Giordani E, Rosati A (2008) Genetic improvement of olive from clonal selection to cross-breeding programs. Adv Hortic Sci 22:73–86
Ben Sadok I, Celton JM, Essalouh L, Zine El Aabidine A, Garcia G, Martinez S, Grati-Kamoun N, Rebai A, Costes E, Khadari B (2013) QTL mapping of flowering and fruiting traits in olive. PLoS ONE 8(5):e62831
Collard BCY, Jahufer MZZ, Brouwer JB, Pang ECK (2005) An introduction to markers, quantitative trait loci (QTL) mapping and marker-assisted selection for crop improvement: the basic concepts. Euphytica 142:169–196
De la Rosa R, Angiolillo A, Guerrero C, Pellegrini M, Rallo L, Besnard G, Bervillé A, Martin A, Baldoni L (2003) A first linkage map of olive (Olea europaea L.) cultivars using RAPD, AFLP, RFLP and SSR markers. Theor Appl Genet 106:1273–1282
De la Rosa R, Kiran AI, Barranco D, León L (2006) Seedling vigor as a preselection criterion for short juvenile period in olive breeding. Aust J Agric Res 57:477–481
Domínguez-García MC, Belaj A, De la Rosa R, Satovic Z, Heller-Uszynska K, Kilian A, Martín A, Atienza SG (2012) Development of DArT markers in olive (Olea europaea L.) and usefulness in variability studies and genome mapping. Sci Hortic 136:50–60
Fanizza G (1982) Genetic variability and fruit carácter associations in table olives (Olea europaea). Riv Ortoflorofrutticoltura Italiana 66:115–120
Fontanazza G, Bartolozzi F, Vergari G (1998) Fs-17. Riv Frutticoltura 5:61
Hansche PE (1983) Response to selection. In: Moore JN, Janick J (eds) Methods in fruit breeding. Purdue University Press, West Lafayette, pp 154–171
Hansche PE, Beres V (1966) An analysis of environmental variability in sweet cherry. Proc Am Soc Hortic Sci 88:167–172
Iwanami H, Ishiguro M, Kotoda N, Takahashi S, Soejima J (2005) Optimal sampling strategies for evaluating fruit softening after harvest in apple breeding. Euphytica 144:169–175
Khadari B, Zine El Aabidine A, Grout C, Ben Sadok I, Doligez A, Moutier N, Santoni S, Costes E (2010) A genetic linkage map of olive based on amplified fragment length polymorphism, intersimple sequence repeat and simple sequence repeat markers. J Am Soc Hortic Sci 135:548–555
Lavee S, Haskal A, Wodner M (1986) ‘Barnea’ a new olive cultivar from first breeding generation. Olea 17:95–99
Lavee S, Avidan B (2011) Heredity diversity in populations of free-, self-, and specific cross-pollinated progenies of some olive (Olea europaea L.) cultivars. Isr J Plant Sci 59:29–37
Lavee S (2013) Evaluation of the need and present potential of olive breeding indicating the nature of the available genetic resources involved. Sci Hortic 161:333–339
León L, Rallo L, Del Río C, Martín LM (2004a) Variability and early selection on the seedling stage for agronomic traits in progenies from olive crosses. Plant Breed 123:73–78
León L, Martín LM, Rallo L (2004b) Phenotypic correlations among agronomic traits in olive progenies. J Am Soc Hortic Sci 129:271–276
Nishio S, Yamada M, Takada N, Kato H, Onoue N, Sawamura Y, Saito T (2014) Environmental variance and broad-sense heritability of nut traits in Japanese chestnut breeding. HortScience 49:696–700
R Development Core Team (2009) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. http://www.R-project.org
Rallo L, Barranco D, De la Rosa R, León L (2008) ‘Chiquitita’ olive. HortScience 43:529–531
Ruiz-Domínguez ML, Raigón MD, Prohens J (2013) Diversity for olive oil composition in a collection of varieties from the region of Valencia (Spain). Food Res Int 54:1941–1949
Sanchez G, Martinez J, Romeu J, Garcia J, Monforte AJ, Badenes ML, Granell A (2014) The peach volatilome modularity is reflected at the genetic and environmental response levels in a QTL mapping population. BMC Plant Biol 14:16
Sato A, Yamada M, Iwanami H, Hirakawa N (2000) Optimal spatial and temporal measurement repetition for reducing environmental variation of berry traits in grape breeding. Sci Hortic 85:75–83
Santos-Antunes F, León L, De la Rosa R, Alvarado J, Mohedo A, Trujillo I, Rallo L (2005) The length of the juvenile period in olive as influenced by vigor of the seedlings and the precocity of the parents. HortScience 40:1213–1215
Thaipong K, Boonprakob U (2005) Genetic and environmental variance components in guava fruit qualities. Sci Hortic 104:37–47
Troggio M, Gleave A, Salvi S, Chagne D, Cestaro A, Kumar S, Crowhurst RN, Gardiner SE (2012) Apple, from genome to breeding. Tree Genet Genomes 8:509–529
Yamada M, Yamane H, Yoshinaga K, Ukai Y (1993) Optimal spatial and temporal measurement repetition for selection in Japanese persimmon breeding. HortScience 28:838–841
Yan W, Frégeau-Reid J, Martin R, Pageau D, Mitchell-Fetch J (2015) How many test locations and replications are needed in crop variety trials for a target region? Euphytica 202:361–372
Young ND (1999) A cautiously optimistic vision for marker-assisted breeding. Mol Breed 5:505–510
Wu SB, Collins G, Sedgley M (2004) A molecular linkage map of olive (Olea europaea L.) based on RAPD, microsatellite, and SCAR markers. Genome 47:26–35
Zine El Aabidine AZ, Charafi J, Grout C, Doligez A, Santoni S, Moukhli A, Jay-Allemand C, El Modafar C, Khadari B (2010) Construction of a genetic linkage map for the olive based on AFLP and SSR markers. Crop Sci 50:2291–2302
Acknowledgments
This work was carried out in the frame of the collaboration agreement between IFAPA and INRA for cooperation on olive breeding research. This work has been partly supported by research project P11-AGR-7301 from the Andalusian Regional Government Council of Innovation and Science. L. León thanks research mobility grant PRX14/00432 of the Spanish Ministry of Education. The co-authors E. Costes and B. Khadari were supported by the Agropolis Fondation OliveMed N° 1202-066.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
León, L., Arias-Calderón, R., de la Rosa, R. et al. Optimal spatial and temporal replications for reducing environmental variation for oil content components and fruit morphology traits in olive breeding. Euphytica 207, 675–684 (2016). https://doi.org/10.1007/s10681-015-1569-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-015-1569-y