Abstract
Gynecologic (GYN) malignancies are gaining new and much-needed attention, perpetually fueling literature. Intra-/inter-tumor heterogeneity and “frightened” global distribution by race, ethnicity, and human development index, are pivotal clues to such ubiquitous interest. To advance “precision medicine” and downplay the heavy burden, data mining (DM) is timely in clinical GYN oncology. No consolidated work has been conducted to examine the depth and breadth of DM applicability as an adjunct to GYN oncology, emphasizing machine learning (ML)-based schemes. This systematic literature review (SLR) synthesizes evidence to fill knowledge gaps, flaws, and limitations. We report this SLR in compliance with Kitchenham and Charters’ guidelines. Defined research questions and PICO crafted a search string across five libraries: PubMed, IEEE Xplore, ScienceDirect, SpringerLink, and Google Scholar—over the past decade. Of the 3499 potential records, 181 primary studies were eligible for in-depth analysis. A spike (60.53%) corollary to cervical neoplasms is denoted onward 2019, predominantly featuring empirical solution proposals drawn from cohorts. Medical records led (23.77%, 53 art.). DM-ML in use is primarily built on neural networks (127 art.), appoint classification (73.19%, 172 art.) and diagnoses (42%, 111 art.), all devoted to assessment. Summarized evidence is sufficient to guide and support the clinical utility of DM schemes in GYN oncology. Gaps persist, inculpating the interoperability of single-institute scrutiny. Cross-cohort generalizability is needed to establish evidence while avoiding outcome reporting bias to locally, site-specific trained models. This SLR is exempt from ethics approval as it entails published articles.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Gynecologic (GYN) neoplasms (2022 ICD-10-CM code: C51-C58) occur in the female genital tract (FGT) and are referred to as tumor sites: vulva (C51), vagina (C52), cervix uteri (C53), corpus uteri (C54), “uterus, part unspecified” (C55), ovaries (C56), “FGT, other/unspecified” (C57), and placenta (C58) (dos Santos et al. 2023). Notwithstanding the mutual system, each depicts distinct features that, in turn, lead to long lists of differential diagnoses and prognoses (Malla et al. 2021; Hsiao et al. 2021). Pathogenesis involves a variety of intrinsic and extrinsic risk factors (RFs) above and beyond inherited faulty genes; for instance, (i) endometrial and ovarian carcinogenesis is attributed to older age (Momenimovahed et al. 2019; Zhao et al. 2021), high adult body mass index (BMI) (Cramer 2012), and/or hormone-replacement therapy exposure (Liang et al. 2022; Liu et al. 2019; ii) cervical carcinogenesis is strongly attributed to high-risk human papillomavirus (HR-HPV) infection and low socioeconomic status (Hull et al. 2020). Such RFs are inherently tied to behavioral, sociodemographic, and environmental factors, which emulates the unequal burden distribution of gynecological malignancies worldwide.
Endometrial (EC) and ovarian cancer (OC) epidemiology is nearly entwined, with a greater prevalence in high-income countries (HICs), notably in Northern and Western Europe, North America, and industrialized populations (Cramer 2012; Zhang et al. 2019; Nees et al. 2022). Under the International Agency for Research on Cancer (IARC) GLOBOCAN series, Canada and the United States of America carry the heaviest endometrial cancer burden (Brüggmann et al. 2020). By 2040, EC global incidence rates are set to leap over 50% from 2018 (Zhang et al. 2019). Ovarian cancer gauges for fewer cases (Cramer 2012), yet it is three times more lethal than breast cancer (Momenimovahed et al. 2019) as OC burden often fences in its complications, comorbidities, and treatment. The same report indicated that, by 2040, OC incidence and mortality rates are set to increase by over 36% and 50%, respectively, from 2020 (Chardin and Leary 2021; everhobbes n.d.).
Cervical cancer (CxCa) is a serious global health concern in low- and middle-income countries (LMICs), wherein 85% of cases and 87% of deaths occur (Rahman et al. 2019). Women with CxCa usually suffer from more complex and acute conditions than other severe illnesses (Krakauer et al. 2021), engendering 740 deaths per day (LaVigne et al. 2017). Again, the emerging World Health Organization (WHO) reports affirm alarming clues of an estimated 442,926 deaths by 2030 (Rahman et al. 2019), over 95% are expected in LMICs (LaVigne et al. 2017)—an upsurge despite the availability of HPV vaccination and cervical screening and management (CSM) programs (Gravitt et al. 2021). To overcome current discrepancies, WHO General Director released a global call to action for CxCa elimination as a public health issue in 2018 (Wilailak et al. 2021). In August 2020, the 73rd World Health Assembly (WHA) endorsed it (Krakauer et al. 2021).
Albeit the surfeit of GYN cancer data, the true burden remains unascertained (Medhin et al. 2020). Documenting the global burden is afflicted by the scarcity of local and national cancer registries in resource-limited settings (LaVigne et al. 2017; Varughese and Richman 2010). Commonly, underdiagnosed or underreported cases in semi-urban or rural areas are substantial impediments to proper assessment for international agency resource allocation (Varughese and Richman 2010; Fiscutean 2021). In 2021, Fiscutean (2021) report raised serious concerns regarding the gap between reality and official OC rates. Although ovarian cancer appears fairly rare in Africa, experts reckon that official records skip vast swathes of individuals and that women, no doubt, die without a diagnosis (Fiscutean 2021). Further, increasing westernization in lifestyles and behaviors is compelling emerging nations to carry the brunt of a heavier burden (Li et al. 2022). Conforming to IARC, OC alone is expected to rise over 87% in Africa by 2040—an eminent soar than anywhere else in the world (Fiscutean 2021). Whether the burden is truly proportionate to the actual overall incidence or a reporting bias, GYN oncology has evolved into a multifaceted global health challenge needing enhanced and scalable analytical approaches throughout the “precision medicine” era. Herein, data mining is introduced.
Data mining (DM), a sub-process of knowledge discovery in databases (KDD), is capturing actionable ‘hidden patterns’ from huge repositories (Idri et al. 2018a). It is perceived as a model-building process that incorporates an amalgamation of techniques and, thus, ‘robust’ models drive domain grasp and decision-making (Duque et al. 2023). Such techniques could be broadly categorized as either: (i) non-machine learning techniques, predicated on classical statistical analysis such as linear regression, principal component analysis, or discriminant analysis; and (ii) machine learning (ML) techniques, elemental of artificial intelligence (AI) in the form of, for instance, a neural network, case-based reasoning, or decision tree (Idri et al. 2018b). Data mining reckon on the aforesaid techniques, but the pursued purpose is distinctive: predictive (supervised learning, “labeled” data) or descriptive (unsupervised learning, “unlabeled” data) (Idri et al. 2018a).
Predictive analytics uses current or historical circumstances to forecast explicit values from data of interest (Razzak et al. 2019). Its implementation yields credence to tasks such as classification and regression (Idri et al. 2018a). Conversely, descriptive analytics looks for patterns and relationships in a given set of data to get a meaningful understanding (Razzak et al. 2019). Unlike predictive counterparts, descriptive models are investigative and do not attempt to predict new properties (Razzak et al. 2019). Clustering and association rules are thus designated under descriptive mining (Idri et al. 2018a).
All analytics are paramount to oncology, revolutionizing every facet of clinical practice (Luchini et al. 2021). As the health sector strives toward automation, increasingly sophisticated cancer-related knowledge could empower intelligent systems and assuredly solidify oncology and DM as viable partners in the near future. By dint of extensive heterogeneity, FGT tumors are an optimum realm for DM breakthroughs, yet little progress has been achieved. One reason for this could be unsatisfactory quality or reporting of the actual evidence, limiting its clinical applicability. The scope of implemented DM-ML in GYN oncology remains imprecise or whether advanced computational techniques enhance scientific understanding, as is the extent to which it goes beyond “numeric”.
Building on a prior mapping study (Idlahcen and Idri 2022), the present systematic review aims to assess the depth and breadth of DM literature in GYN malignancies during the past decade, with an emphasis on ML-based approaches. Matter of fact, systematic maps (SMSs) serve a key role in narrowing down a broader topic across perusals pertinent to valuable questions (Idlahcen and Idri 2022). Likewise, SMSs tend to be well-suited for outlining research gaps, depicting research trends, or bracing an exhaustive systematic literature review (SLR) (Idlahcen and Idri 2022). Notwithstanding analogous characteristics, SMSs and SLRs are distinguishable in goals and analysis schemes (Idlahcen and Idri 2022). While an SLR aims to achieve an evidence synthesis, an SMS is particularly concerned primarily with visually structuring literature into tabular and graphical summaries (Idlahcen and Idri 2022). With this in mind, the strength of the present SLR is two-fold:
-
Synthesize evidence of DM-ML literature relevant to GYN oncology by searching extensively the research published between January 1, 2011, and February 28, 2022, in five digital resources: PubMed, IEEE Xplore, ScienceDirect, Springer Link, and Google Scholar.
-
Analyze the selected articles through eight aspects:
-
1.
Publication trends, venues, and sources where the selected articles have been issued.
-
2.
Discerning the commonest tackled GYN sites-derived neoplasm with DM-ML.
-
3.
Classifying the selected articles according to contributions and empirical schemes.
-
4.
Determining the most popular data modalities and openly accessible repositories relevant to GYN oncology.
-
5.
Discerning the most encountered GYN oncology tasks by DM-ML.
-
6.
Determining the most pursued DM-ML aim in the literature.
-
7.
Highlighting the most adopted DM-ML techniques in the literature.
-
8.
Determining the most adopted performance framework to assess DM-ML models.
We anticipate the present systematic review would disclose literature gaps and ease foster the knowledge and awareness of DM-related GYN oncology practices among oncologists and data practitioners.
The remaining document is outlined as follows. Section 2 describes the research methodology, and how articles were selected and assessed. Next, the findings of the RQs are delineated in Sect. 3 and discussed in Sect. 5. Then, Sect. 6 suggests recommendations about what the audience should address based on the research findings. Section 7 sums up the present SLR.
2 Methods
A systematic literature review (SLR) is a structured, transparent, and exhaustive scientific tool to get the “lay of the land” of a topic (Greyson et al. 2019; Martinic et al. 2019). It is, therefore, a well-defined and protracted protocol that requires expertise to be conducted (van Haastrecht et al. 2021). SLR policy traces back its genesis to medicine (or evidence-based practice) (van Haastrecht et al. 2021). Since then, it has expanded across all disciplines, actively computer science subfields such as software engineering (Kitchenham et al. 2004), artificial intelligence (Sarana and Subhashini 2023), and medical (health) informatics (Hosni et al. 2019; Wadghiri et al. 2022; Melton 2017). The first Evidence-Based Software Engineering (EBSE) paradigm was proposed by Kitchenham (2004) Kitchenham (n.d)., then, through embracing (Brereton et al. 2007) strategy (2007), Kitchenham and Charters (2007) supplied clear guidelines (Keele et al. 2007) for systematic reviews’ conduct.
The following sections describe the methodology used throughout this paper.
2.1 Study design and protocol registration
The current study was undertaken as an SLR in compliance with Kitchenham and Charter’s guidelines and the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines (Page et al. 2021). The PRISMA Protocols (PRISMA-P) checklist (Moher et al. 2015). The study protocol has been registered with PROSPERO (Schiavo 2019), the international registry for systematic reviews. No ethical approval or informed consent was mandated.
2.2 Research questions
The goal of the present SLR could be explicitly translated into research questions (RQs) as follows:
-
RQ1. In which years, publication venues, and sources were the selected articles published?
-
RQ2. What is the most tackled GYN site-derived neoplasm using DM-ML?
-
RQ3. In which contribution and empirical schemes are the selected articles categorized?
-
(RQ4.1) What are the prevailing data modalities to investigate DM-ML as an adjunct to GYN oncology? (RQ4.2) Which open datasets are commonly used to investigate DM-ML as an adjunct to GYN oncology?
-
RQ5. Which medical tasks of GYN oncology practice are addressed with DM-ML?
-
RQ6. What DM-ML objectives are fulfilled for GYN oncology research?
-
RQ7. Which DM-ML techniques are commonly implemented in GYN oncology settings?
-
(RQ8.1) Which performance metrics are prevalent to evaluate DM-ML techniques as an adjunct to GYN oncology? (RQ8.2) Which validation techniques are prevalent to evaluate DM-ML techniques as an adjunct to GYN oncology? (RQ8.3) Clinicians vs. DM-ML? (RQ8.4) How perform DM-ML in GYN oncology (A case of cervix image classification for screening)?
Through answering our RQs, we seek to supply a systematic overview of the in-depth and meticulous analysis of DM-ML implications in GYN oncology.
2.3 Search strategy
Five digital libraries: (i) PubMed, (ii) IEEE Xplore, (iii) ScienceDirect, (iv) Springer Link, and (v) Google Scholar, were searched for peer-reviewed and English-language articles published from January 2011 until February 2022. We crafted the search query guided by two schemes: (i) PICO (Population, Intervention, Comparison, and Outcomes) synthesis recommended by Kitchenham and Charters and (ii) extraction of key terms/concepts from the RQs. Since no empirical assessments and tangible outcomes were conducted in the present SLR, only the initial two letters of PICO acronym and analogs were gleaned. The full scheme is supplied in Supplementary Material 1.
2.4 Study selection
An article of interest is verified as a primary study if it meets all the inclusion criteria and none of the exclusion criteria. The following inclusion (I) and exclusion (E) criteria for eligibility were used:
-
IC1. The article regards novel, or improving current, DM-ML/ GYN oncology.
-
IC2: The article scrutinizes DM-ML/ GYN oncology.
-
IC3. The article regards empirical/theoretical scrutiny of DM-ML/ GYN oncology.
-
IC4. The article pertains to [January 1st, 2011–February 28, 2022].
-
EC1. The article covers other kinds of cancer besides FGT sites.
-
EC2. The article covers polycystic ovary syndrome (PCOS) or endometriosis.
-
EC3. The article regards primarily novel, or improving current, data pre-processing approach.
-
EC4. The article regards radiomics, pathomics, or radiopathomics material.
-
EC5. The article is not presented in English.
-
EC6. The article is a duplicate.
-
EC7. The article is no longer than 6 pages (incl. references, well-marked appendices).
-
EC8. Extended abstracts, posters, or editorials.
-
EC9. The article is a non-peer-reviewed material.
-
EC10. The full-text is unavailable.
Subsequent to discarding duplicate entries through EndNote reference management software, articles were carefully screened to reject or retain the paper based on metadata, i.e. titles, abstracts, and keywords; or otherwise, the full text if metadata is unsatisfactory. The evaluation step was carried out independently by the two authors (AI and FI).
2.5 Quality assessment
Each article was retained upon four QAs: (i) QA1 assesses the clarity of the empirical results provided in the study by means of the research questions, (ii) QA2 assesses both the clarity of the empirical design and the systems set of the experiment, (iii) QA3 assess the adoption of performance measures to evaluate the study outcomes, and (iv) QA4 scoring attests the rank of the article: (i) for conferences, the ranking is based on the Computing Research and Education Association of Australasia (CORE Conference Ranking Exercise 2021), and (ii) for journals, the ranking is based on the Web of Science Journal Citation Reports (JCR 2020). We regarded article quality scoring as an exclusion criterion. Note that a similar strategy was adopted by Idri et al. (2015). (Supplementary Material 2: Tables S2A, S2B, and S2C) provide the QA criteria checklist and a full list of the selected articles and venues, including corresponding QA scores.
2.6 Data extraction
At this point, each eligible article was subjected to a data extraction framework to address the aforementioned RQs. The following are the joint fields.
-
Publication Year: calendar year.
-
Publication Venue Petersen et al. (2015): journals, conference proceedings (or book chapters).
-
Source Name: official title of the publication venue.
-
GYN neoplasm(s), in which DM-ML techniques have been performed, was/were identified for each study. The neoplasm(s) site could, for example, be ovarian, endometrial, cervical, vaginal, vulvar, etc.
-
Contribution Types (Wieringa et al. 2005): (i) Evaluation research (ER), a paper evaluating empirically a given DM-ML technique (new or existing). (ii) Solution proposal (SP), a paper proposing a DM-ML solution approach. It must therefore be novel, or markedly improve an existing approach. (iii) Validation research (VR), a paper exploring a DM-ML technique in clinical practice. (iv) review, a paper assessing the literature of DM-ML as an adjunct to GYN oncology. (v) Others, e.g. experience reports, philosophical papers, opinion papers.
-
Empirical Methods (Petersen et al. 2015): (i) Case study (CS), an empirical evaluation carried out in a real-life medical setting. (ii) Historical-based evaluation (HBE), a paper assessing the performance of a specified DM-ML technique throughout existing datasets. (iii) Survey (S), a paper based on a dataset garnered through a questionnaire.
-
Data modalities could, for example, be a whole-slide image (WSI), miRNAs-mRNAs expression profiles, GC-MS serum metabolomic fingerprint, hysteroscopy captures, mid-infrared (MIR) spectra of biofluids, cervicograms, DCE-MRI pharmacokinetic parameters, etc.
-
Publicly available dataset name.
-
Six core medical tasks (Esfandiari et al. 2014) (i) Screening (Sc), detecting illness or body dysfunction prior to as-yet-unrecognized symptoms or markers. (ii) Diagnosis (Dx), recognizing the character of a disease from its symptoms and markers. (iii) Treatment (Tx), undertaking measures to cure a patient; diagnosis outcomes serve to determine a suitable treatment. (iv) Prognosis (Px), predicting the likelihood of a disease’s course or outcome, esp. recovery, recurrence, survival, and quality of life (QoL). (v) Monitoring (M), surveilling a patient’s condition throughout time. (vi) Management (Mx), promoting health and medical services.
-
Four DM-ML objectives are considered. (i) Classification, categorizing into classes a given set of data; the output is categorical, e.g. ’case’, ’control’. (ii) Regression, predicting continuous or real values from a given set of data; the output is numerical, e.g. BMI, temperature. (iii) Clustering, disclosing inherent groupings in a given set of data, e.g. grouping patients by health perception. (iv) Association rule mining (ARM), identifying frequent patterns, co-occurrences, and correlations in a database, e.g. the relationship between LMICs and cervical cancer.
-
DM-ML techniques could, for example, be deep neural networks (dNNs), support vector machine (SVM), decision trees (DTs), k-nearest neighbors (k-NN), etc.
-
Performance metrics could, for example, be an accuracy (Acc), sensitivity (Sen), specificity (Spe), F-measure, area under a receiver operating characteristic curve (AUROC), mean absolute error (MAE), etc.
-
Validation techniques could, for example, be k-fold cross-validation (Kf-CV), leave-one-subject (patient)-out (L1SO-CV), Monte Carlo (MC-CV), Hold-out, etc.
-
The performance of the selected sample techniques.
2.7 Threats to validity
The core threats to the present SLR validity are narrated in the following two aspects.
-
1.
Study selection bias. To capture the maximum evidence and avoid any identification bias(es), we constructed a search strategy at two levels. First, gather pertinent keywords from the research questions under PICO framework. Second, exert iterative testing in which we examined the effect of alternate terms, stemming/wildcards, Boolean search conventions, and the use of assigned terms (MeSH terms) for a broader and refined search string. This resultant was performed across five digital libraries, PubMed, IEEE Xplore, ScienceDirect, Springer Link, and Google Scholar, with adjustments as necessary according to each database. Given the cross-disciplinary nature of the topic, it was prudent to utilize multifaceted libraries to access a broader range of pertinent papers, encompassing both “computer science” and “biomedicine” literature. The Google Scholar search engine functioned as a “secondary source” to ensure we rarely miss out on an article.
-
2.
Internal validity, or potential for bias(es), is a particularly important feature of systematic reviews where the scope to which the conduct achieves trustworthy results. It deals with data extraction and analysis (Idri et al. 2016). To lessen the threat of imprecise or erroneous “data extraction”, the form was completed independently by the two authors. Disagreements or doubts were settled by in-depth team discussions (AI and FI) to establish inter-rater reliability.
3 Results
In this systematic review of all DM-ML/ GYN oncology, we found 3499 candidate articles. 181 of which fulfilled eligibility criteria, were in-depth analyzed to provide valuable insights.
3.1 Study selection and characteristics: an overview
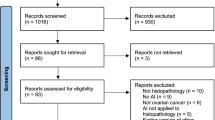
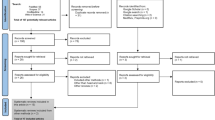
The total articles retrieved at each stage of the study selection is depicted in Fig. 1. In a prior systematic map (Idlahcen and Idri 2022), a total of 2807 primary studies published between January 2011 and June 15, 2021, were identified by searching five digital libraries as granted in Sect. 2.3. The updated search yielded 692 extra candidate articles published between June 16, 2021, and February 2022, which were appended to the earlier number, amounting to 3499 primary studies. 3142 articles were discarded following the ICs/ECs, retaining 357 articles emphasizing the application of DM-ML in GYN oncology. At last, we used the quality assessment on the 357 relevant articles. Due to low-quality scores, the process resulted in the discard of 176 articles. While 50,7% (181 art.) of the selected articles had a higher average score as per Table 1, 49.3% (176 art.) were below and thus subtracted. Ultimately, 181 articles were used.
The included articles’ data extraction is summarized in full in (Supplementary Material 3: Table S3).
3.2 RQ1. In which years, publication venues, and sources were the selected articles published?
The majority is published in journals (93.92%, 170 art.). Optimal ones (Table 2) to publish on DM-GYN oncology cope with disciplines as follows: (i) computer science -AI, (ii) obstetrics (OB)/GYN, or (iii) biomedical (clinical or health) informatics. By virtue of our QA, not CORE-ranked conference proceedings were excluded, leading to only (6.07%, 11 art.) being captured in the selection study. Accordingly, journal analysis revealed an interdisciplinary nature within the field under consideration. Figure 2 has more information on the distribution of the selected articles. We discern notable fluctuations in the yearly publication count. From 2011 to 2014, it is subjected to a gradual increase from one to ten published articles respectively, followed by a drop to eight articles each in 2015 and 2016, a progressive emerges over 2017–2018 (11–13 art.), then a sharp spike onward 2019, reaching 66.85% (121 art.) of the selected articles by 2022. While cervical neoplasm research contributed significantly to the 2019 soar by 60.53%, the other kinds remained marginally steady.
3.3 RQ2. What is the most tackled GYN site-derived neoplasm using DM-ML?
GYN tumors are a heterogeneous collection of masses arising from the female reproductive organ(s)—featuring five main types: cervical, ovarian, endometrial, vaginal, and vulvar (Lõhmussaar et al. 2020). After data extraction, we encountered several distinct types, incl. uncommon forms. To keep the report concise and easy for users to interpret, we elected to shorten GYN masses into six classes from both FIGO systems (Kehoe and Bhatla 2021) and “International Classification of Diseases (ICD)” standpoint as follows: (i) cervix uteri; (ii) ovary, Fallopian tube, and peritoneum; (iii) uterus; (iv) vagina; (v) vulva; and (vi) placenta; as shown in Table 3. Recall that a single article could examine multiple GYN neoplasms.
We witness a high focus for “cervical” neoplasms (57.6%, 118 art.), followed by “ovarian, tubal, and peritoneal” ones (24.9%, 51 art.), then “uterine” (15.1%, 31 art.) class. At last, “vaginal”, “vulvar”, and “placental” represents 1%, 0.5%, and 1%, respectively. Regarding the “ovary, Fallopian tube, or peritoneum” site, ovarian tumors garnered the most attention with 40 articles, followed by adnexal (6 art.), tubal (3 art.), and peritoneal (2 art.). While sarcoma and leiomyoma each account for five and four per uterine articles, endometrial tumors reckon 20 articles, alongside two additional art.: (i) one arising from the round ligament, and (ii) an unstated fibroid type. Such constitutes respective 64.51%, 16.13%, 12.9%, 3.23%, and 3.23% of the uterine site. Within placental tumors, throphoblastoma and choriocarcinoma are both covered in a single article.
3.4 RQ3. In which contribution and empirical schemes are the selected articles categorized?
Wieringa et al. (2005) classify research/contribution types into six broad categories: (i) evaluation research (ER), (ii) solution proposal (SP), (iii) validation research (VR), (iv) philosophical papers, (v) experience reports, and (vi) opinion papers. 84.2% (155 art.) of the selected articles were SPs either providing a new approach or improving an existing one; of which all were empirically evaluated. A further 13% (24 art.) were classified solitary as ERs assessing the performance of current or novel DM-ML. Since extensive research is labeled similarly to solution and evaluation proposals, the number of ER articles steadily exceeded that of SP papers—particularly onwards 2012. Of note, five articles (2.7%) were VR studies.
Figure 3 displays the distribution of contribution types by year. The topmost are ERs that continued to rise with the fastest expansion from 2016 to 2020 (108 art.), thus the least declines were over 2014–2016 (10 to 8 art.) and 2020–2021 (45 to 39 art.). As only articles up to February 2022 were gathered, it is unsatisfactory to induce a statement pertaining to 2022. SPs had an akin tendency, on average, up to 2017 and beyond 2018 as a little dip was noticed during 2017–2018 when compared to ERs variations. The bottommost is for VRs accounting for one article optimum, yet distribution has not altered substantially in a decade.
Three empirical types could be outlined: (i) historical-based evaluation (HBE), (ii) case study (CS), and (iii) survey. Of the empirically evaluated articles (Table 4): (i) 36.72% conducted an HBE through publicly available datasets. Such type was used in 51.77%, 47.52%, and 0.71% of ER, SP, and VR papers, respectively; (ii) 61.72% adopted a CS as it was conducted in its medical real-life context, 52.32%, 45.99%, and 1.69% amongst were respective ER, SP, and VR studies; and (iii) surveys were employed in 1.56% of the articles, of which two SPs, three ERs, and one VR.
3.5 RQ4.1. What are the prevailing data modalities to investigate ML as an adjunct to GYN oncology?
Table 5 provides insights on the investigated medical data modality(ies). Recall that medical imaging encompasses most modalities (58.3%, 130 art.). It is also worth noting that 85.08% of the selected articles regarded follow-up biopsy as ground truth (Table 6).
3.6 RQ4.2. Which open datasets are commonly used to investigate ML as an adjunct to GYN oncology?
Table 7 denotes the publicly available datasets.
3.7 RQ5. Which medical tasks of GYN oncology practice are addressed with DM-ML?
Figure 4 Illustrates the examined medical tasks in the present selection study. No “Management” medical task was undertaken by any of the selected articles. It should be noted that a single study could examine multiple medical tasks.
3.8 RQ6. What DM-ML objectives are fulfilled for GYN oncology research?
Figure 5 displays the frequency of DM-ML objectives in the present selection study. Recall that more than one objective could be examined in a single study.
3.9 RQ7. Which DM-ML techniques are commonly implemented in GYN oncology settings?
Figure 6 depicts the proportion of DM-ML techniques adopted per objective. Each technique has its strengths and limitations as summarized in Table 8. Figuring in 127 of the selected articles, neural networks (NNs) were the most prevalent DM-ML approach. Of these, 116 and 11 art. were committed to classification and regression, respectively. Solely to classification, NNs were followed by support vector machine (SVM), random forest (RF), k- nearest neighbors (k-NN), logistic regression (LR), and decision trees (DT), respective accounting for 79, 37, 37, 36, and 17 art. The rest of the classification techniques englobe LDA, naïve bayes (NB), adaboost, XGBoost (respective 14, 15, 7, and 7 art.), and others such as QDA, PLS-DA, BN, GBM, E-net, GCN, GNN, ELM to a lesser extent. NNs, comprise three types mainly: (i) standard artificial neural network (aNN), convolutional neural network (CNN), and (iii) recurrent neural network (RNN). CNNs gathered the most attention with 66 art. starting from AlexNet to Densenet-121. In regression, NNs were followed by LR, RF, SVM, DT, and XGBoost, accounting for 9, 8, 3, 2, and 2 of the selected articles, respectively. The rest of the regression techniques, such as GBM and CART, were less examined at 3% (8 art.) for each. Besides, k-means was the most prominent DM-ML clustering approach, with 12 articles implementing it. The remaining clustering techniques included: fuzzy c-means (FCM), partitioning around medoids algorithm (PAM), and hierarchical methods, e.g. hierarchical cluster analysis (HCA). Yet the ensemble algorithm for clustering cancer data (EACCD), which was designed in 2009 by D. Chen et al. to enhance the TNM staging system without altering its fundamental concepts, was observed in both Grimley et al. (2021) and Praiss et al. (2020) studies. Association-related articles employed four DM-ML techniques: fuzzy association rules mining (FARM), Apriori, and constrained-rule learning algorithm.
3.10 RQ8.1: Which performance metrics are prevalent to evaluate ML techniques as an adjunct to GYN oncology?
Figure 7 presents the used benchmark metrics to evaluate the selected articles’ models. Such are classified into (i) single scalar, e.g. accuracy, sensitivity, specificity, etc., or (ii) graphical, e.g. AUROC, AUPRC, etc. The five commonly used metrics were: (i) accuracy (Acc) (125 art., 17.7%); (i) sensitivity (Sen), recall (Re), or true-positive rate (TPR) (136 art., 19.2%); (iii) specificity (Spe) or true negative rate (TNR) (101 art., 14.3%); (iv) area under the receiver operating characteristics (AUROC) (78 art., 11%); (v) precision (Pr) or positive predictive (prognostic) value (PPV) (71 art., 10%); and (vi) F-measure (39 art., 5.5%). Nevertheless, each article tends to use a variety of performance metrics and the common combination was ‘accuracy, sensitivity, specificity, and AUROC’.
3.11 RQ8.2: Which validation techniques are prevalent to evaluate ML techniques as an adjunct to GYN oncology?
Validation schemes are used to legitimize the performance of a DM- ML model fairly and accurately. Figure 8 depicts the validation strategies reported in the present selection study, namely, (i) cross-validation (CV), (ii) hold-out or data split, and (iii) bootstrapping, accounting for 62.43% (123 art.), 28.42% (56 art.), and 2.03% (4 art.) of the selected articles, respectively. Variants such as bootstrapped Latin partition (BLP) and Kennard-Stone (K-S) algorithm were each reported in one study. In 6.1% of cases, the validation process was unstated. Recall that a single article could undergo multiple validation techniques.
As per CV, five types were used, (i) Monte Carlo (MC-CV) in one study, (ii) k-fold (k-fCV) in 93 studies, (iii) stratified k-fCV in two studies, (iv) leave-one-subject(patient)-out (L1SO-CV) in 3 studies, and (v) leave-one-out (LOO-CV) in 14 studies. The ‘k’ value was either 5 (40.7%) or 10 (49.5%) to a greater extent, instead observed were the values of 2, 3, 6, 7, 9, and 15. Of note, the CV type was unstated in 10 studies.
3.12 RQ8.3: Clinicians vs. DM-ML?
17 out of 181 articles compared the performance of the implemented DM-ML models to that of clinicians. The assessments predominantly engaged board-certified seniors with professional experience spanning from 5 to over 25 years in the domain of interest or at different professional levels (interns, in-service, and professionals). The participants, who were blinded to the pathological and clinical findings of the study, frequently conducted independent reviews of the test sets in a randomized order. The commonly used metrics were Acc, Sen, Spe, AUC, and/or interobserver agreement.
3.13 RQ8.4: How do perform DM-ML in GYN oncology (A case of cervix images classification for screening)?
Due to the selection study’s heterogeneity, we carried out an overall evaluation of the performance of DM-ML techniques used in 10 selected studies which presented 260 empirical evaluations. These selected articles are (i) ranking a quality score of 5, and (ii) relevant to cervical cancer, classification, screening, and all associated imaging, i.e. liquid-based cytology, pap smear, and cervicograms. We selected the commonly used performance metrics, i.e. accuracy, sensitivity, specificity, f-measure, and AUC, to assess the reviewed techniques as reported in Sect. 3.10. The reported values were retrieved for the selected measures to synthesize all of the evaluations of each technique. An evaluation equates to using a DM-ML technique on a dataset, which means that an article on distinct datasets provides multiple evaluations. (Supplementary Material 4: Table S4) reports the performance metrics of the 10 selected studies and Table 9 as summary. Although not representative, we observe as follows:
-
CNNs was the most assessed with 49 evaluations (9 art.).
-
The highest accuracy values were attained by aNNs, CNNs, and SVM, with respective means of 93.22%, 92.65%, and 88.73%, and low related standard deviation values.
-
Bns(C) and k-NN also achieved good accuracy values, with respective means of 83.63% and 85.23% for the same number of evaluations, 44 each. Still, high related standard deviation suggests that the values are dispersed across a greater range, raising questions about their performance.
-
aNNs reached high sensitivity and specificity values also, indicating their strength in test diagnostics.
-
CNNs may tend to falsely find a result even though they attained high specificity values due to the high associated standard deviation.
4 Discussion
4.1 DM: Changing the landscape of GYN oncology
The publications count soared markedly in 2019. The overall trend is consistent with three hypotheses: (i) deployment of digital pathology (Hanna et al. 2020), “omics” (Adamo et al. 2018), and datafication in clinical and non-clinical settings (Manteghinejad and Javanmard 2021), (ii) “precision oncology” era opened up novel pathways to urge AI-DM integration, and (iii) 2018 WHO DG global call-to-action to eliminate cervical cancer as a public health issue (Wilailak et al. 2021). Prior to 2017, no regulatory approval was issued for whole-slide scanners in primary surgical pathology, resulting in virtual microscope glass slides standing in stark contrast to digitizing radiology since the 1980s (Boyce 2017). Besides, large-scale pan-omics repositories, e.g. The Cancer Genome Atlas (TCGA), Gene Expression Omnibus (GEO), Clinical Proteomic Tumor Analysis Consortium (CPTAC), and so forth, are rapidly expanding to accommodate multi-omics with the improvement and reduced cost of mass spectrometry (MS) and high-throughput next-generation sequencing (NGS) technologies (Morganti et al. 2019; Girolamo et al. 2013). As data is a vital prerequisite, a such substantial upsurge gave impetus to the development of novel ML strategies that integrate multifaceted and pertinent datasets beyond questionnaires and classical radiology. Following 2018 WHO DG-plea, over 70 countries acted swiftly and positively toward “hidden” cervical cancer, which may partially explain the overall peak of publications in 2019—prominently on cervical cancer—as reflected in Fig. 2.
4.2 Aligning DM with GYN cancer disparities
Cervical cancer is a paradigm of global health disparity reckoning higher morbimortality rates than the combined uterine and ovarian malignancies (Medhin et al. 2020). Thus, it is apparent that “cervix uteri” was the most examined FGT site. But while the CxCa burden is readily recognized in LMICs, endometrial cancer represents a major global health disparity in developed countries (Medhin et al. 2020). Yet, “corpus uteri” endometrium was little examined. Such scrutiny entails more research as recent US evidence (Smrz et al. 2021) corroborates a perennial increase in EC incidence rates, pointedly young-onset age. Ovarian neoplasms were the second most examined, haply by cause of lethality associated with poor survival rates than those of uterine and cervical malignancies (Nie et al. 2021). Only a few investigated the less common forms of uterine and adnexal masses, but despite the lesser extent, it is still important for (i) a differential diagnosis between degenerated leiomyomas and uterine sarcomas in magnetic resonance imaging (MRI) (Suzuki et al. 2019) and (ii) case–control of adnexal masses in ultrasound images as the first-line imaging modality to satisfactorily patient triage (Holsbeke et al. 2012)—prospective research is therefore desirable. The remaining types, incl. (i) vaginal as reported by (Tian et al. 2019) and (Liu et al. 2021; ii) vulvar, round ligament, throphoblastoma, and choriocarcinoma as reported by Liu et al. (2021); (iii) tubal as reported by Liu et al. (2021), Laios et al. (2020), and Vázquez et al. (2018); and (iv) primary peritoneal as reported in Laios et al. (2020) and Vázquez et al. (2018); were incorporated alongside common types, which is insufficient to induce a statement relevant to such kinds.
We draw attention to a singularity, (i) (Kumar et al. 2014) examined the pre-/post- treatment quality of life (QoL) in CxCa patients, (ii) (Zhang and Han 2020) investigated an under-researched cancer patient population, which is pregnant or lactating women, and (iii) (Asaduzzaman et al. 2021) implicated patients suffering from ovarian or cervical cancer, with stress (Mental) disorder too. Since it is fundamental to analyze an individual as a “whole” not independently as “parts” or “organs”, it is constantly appealing to target a multimorbidity-based or a whole patient-based sample population to unravel pathogenesis insights. Besides, Kumar et al. (2014) and Asaduzzaman et al. (2021) pinpointed valuable yet neglected aspects in GYN tumors patients, i.e. mental disorders and quality of life after therapy. As it stands, extensive research is required to investigate the ability of ML-based prediction algorithms in terms of (i) capturing behavioral morbidity or subjective psychological response patterns across risk factors and (ii) forecasting the quality of life in the functional, symptom, and global health/QoL scales for finer treatment modules. Similarly, maternal tumors raise several concerns regarding the mother-fetus safety as well as the cause-specific survival of pregnant and lactating women with cancer or post-cancer, so it is of special interest to address the use of learning techniques to improve detection and care in obstetrical settings.
4.3 DM: a paucity of clinical validation
While medical data expansion paints an overall positive picture, existing DM-ML techniques still need to integrate with medical data conformations, which is congruous with the large portion of solution proposals scrutinized (84.2%). As advanced approaches are proposed to attain even better performance, expectedly, all amongst were empirically evaluated. However, we are particularly concerned about the paucity of validation research (5 art.). Since building and deploying DM models differ, it is crucial to cushion what might adversely strike the proposed model in real-world clinical settings. Such scarcity asserts that the implementation of DM techniques in GYN oncology is still limited to academic research, more research is needed to raise maturity.
It is appealing that most of the selected articles (97.28%) were empirically evaluated, with 60.03% referring to case studies, since it reflects researchers’ awareness of the value of cooperating with practitioners for successful trials and thereof generalizing across different cohorts with disparate parameters. The next 36.72% used historical-based data, accounting for 47.52% of solution proposals owing to both reliability and ease of open-access datasets use when evaluating SPs performance. Yet, the wide adoption of CS could also raise the question of the scarcity of availability and sharing of databases related to GYN Oncology. Barriers to biomedical data sharing remain major hindrances to availability, utilization, and inter-institutionalization and have been well-documented by van Panhuis et al. (2014). The resulting taxonomy concern: (i) technical barriers, including lack of data collection, no prioritization in data preservation or archiving, restrictive data formats, lack of metadata, standards, or interoperability (e.g. structure or language), along with a scarcity of analytical solutions to address such challenges although identified, hinder the potential for integrating several datasets, particularly in an international context. (ii) motivational barriers rooted in personal or institutional beliefs, including lack of incentives, opportunity costs, worry of criticisms or discredits, and disagreement on data secondary access/use. Solutions entail the establishment of trust or the formulation of transparent legal agreements. (iii) economic barriers concerning the feasibility and actual costs and damages related to data sharing and potential solutions hinge on acknowledging the value of data and implementing sustainable financing mechanisms. (iv) political barriers ingrained within health governance systems often manifest as issues related to trust, restrictive policies, and lack of guidelines. Addressing them is not a clear-cut endeavor, requiring national and global processes to foster agreement and political will. (v) legal barriers related to ownership, copyrights, and privacy protection. Resolving them relies on underlying political barriers. (vi) ethical barriers involving conflicts between moral principles and values and readily apparent in issues related to reciprocity and proportionality. Engaging in a global dialogue on the ethical principles and model legislation governing data share could resolve it. In-depth evidence is needed to broaden the understanding on sharing biomedical data. As knowledge of barriers deepens, the scope for solutions and efforts would enlarge for the benefit of global health.
Only 1.56% of the articles are based on surveys due to the strain of obtaining exhaustive and accurate information from questionnaires. Previous literature (Jo 2022) acquiesces three assumptions: (i) survey structure, (ii) narrow knowledge, and (iii) refusal to answer because of personal, sensitive, or uncomfortable concerns, notably related to GYN cancer survivors QoL.
4.4 Barriers to DM in GYN oncology
Medical imaging is a force to be reckoned with in modern medicine, it provides a DM foreground application, yet it is no wonder imaging was the most used with 58.3% of the selected articles. Yet, to join Sect. 4.3, DM-ML is still restricted. The real hurdle encountered in medical data is the inability to use straightforward approaches without any type of pre-processing to disable what might have detrimental effects on the developed model.
As for image-based data, whole-slide imaging has (i) the highest noise, (ii) staining fluctuation, (iii) artifacts, (iv) a tendency to surpass the stacking capability of GPUs, and (v) an intra- and interobserver heterogeneity, which might yield poor perception in either manual or computer-assisted assessment. Hence, implementing WSIs is a decisive endeavor requiring, at a minimum, stain normalization, artifact correction, and patch-based analysis. As hematoxylin-eosin (H &E) and immunohistochemistry staining reagents and protocols vary considerably between laboratories and often fade over time (Downing et al. 2019), it is of urgent necessity to explore prospective multi-center cohorts with varied manufacturers’ chemicals, standards, magnifications, and further modern pathology RQs. Next, digital captures from cervicography and hysteroscopic video clips come with poor quality due to blur, incomplete cervix or uterus exposure, lesions coated by vaginal discharge, heavy bleeding, and so forth (Yuan et al. 2020), resulting in a loss of clinically relevant features and a pronounced class imbalance. Digital captures entail generalization as well over multistate scenarios—notably regarding the used contrast agents in colposcopy. However, only three articles (Yuan et al. 2020; Asiedu et al. 2019; Yu et al. 2021) examined D-VIA with either D-VILI, saline, or built-in green filters. Besides, most research neglects the sequential process of colposcopy-based precancerous lesion recognition by considering only a single cervicogram, which makes it difficult to reflect the acetowhite changes in compliance with the diagnostic customs of gynecologists. Herein, Li et al. (2020) and Peng et al. (2021) provide successful cases of cervical intraepithelial neoplasia (CIN) diagnosis from time-lapsed cervicograms based on respective (i) fusion approaches at distinct post-acetic test time slots, i.e. 60 s, 90 s, 120 s, 150 s; and (ii) multimodal feature changes of registered pre- and post- acetic acid test images. Recall that most of the articles of the current selection study converted the special medical image file formats to JPEG, which is not required when loss of the finest details is forbidden. Such might alter the diagnostic accuracy in contrast to learning directly from DICOM (Urushibara et al. 2021) or Aperio SVS.
As for feature-based data, missing values are inevitable—particularly in electronic records and questionnaires (see Sect. 4.3). One straightforward approach to account for it is to remove the related instances, yielding usually to imbalanced and smaller datasets. As it stands, data must be pre-treated by either oversampling the minority class or predicting the missing values, which again is limited to biased findings. Besides, biomedical data has both multimodal and high-dimensional properties, involving common inconsistencies, redundancy, and noise in raw data. Such matter leads to the “curse of dimensionality’—for instance in transcriptome profiling, where miRNA expression microarray data contains massive genes with a little number of observations. It is therefore vital for a gene selection to minimize its size by selecting informative ones regardless of the DM techniques used.
Of note, it is worthwhile to highlight the efforts made by researchers to investigate multifaceted and cutting-edge medical data.
4.5 DM: going beyond diagnostics
“Diagnosis” underwent a higher level of scrutiny (42%), not unexpectedly with the intent of granting patients with ideal diagnostic scenarios: (i) inter- and intra-observer discrepancy-free, (ii) non- or minimally invasive, and (iii) early detected.
Visual examination of biopsied tissues is the “gold standard” for proper cancer diagnosis, yet pathological findings might differ strikingly even within a single specimen due to observer variability from human error (Genta 2014). To avert misdiagnoses, results are always subject to an expert second opinion as mandated by most pathology laboratories’ policies (Farooq et al. 2021). As the “tipping point” of digital pathology (DP), ML-based CADx(s) are able now to supply inter-rater consensus and consistent “second opinion” to clinicians. CADx(s) could even be of significant practical value in GYN oncology since FGT neoplasms exhibit salient intratumoral heterogeneity (ITH), a confounding trait driven in clinical diagnostics (Yin et al. 2019). As it stands, most studies supported the ability of EL-ML- and DL-classifiers to offer gyneco-pathologists refined interpretation for all OC, CxCa, and EC (BenTaieb et al. 2017; Sun et al. 2020; Downing et al. 2019; Yu et al. 2020a; Guo et al. 2016; Huang et al. 2020c; Li et al. 2019; Zhang et al. 2021a; Zeng et al. 2021; Cheng et al. 2020; Yang and Stamp 2021; Huang et al. 2020a; Meng et al. 2021; Xue et al. 2020a; Shin et al. 2021; Xue et al. 2019a; Zhang et al. 2021b). Counterintuitively, diagnoses are not restricted to cancer cases or healthy control, yet baseline report items wrap the histologic type, tumor grade, FIGO stage, tumor margin, lymph node status, and so forth. The foregoing exhibits subjective facets within GYN tumors, for instance, the inconsistency of differentiating endometrial adenocarcinoma from complex atypical hyperplasia within a biopsy, hysterectomy, or curettage material (Allison et al. 2008). Herein, Sun et al. (2020) proposed HIENet to classify endometrial tissue WSIs into “normal endometrium”, “polyp”, “hyperplasia”, and “adenocarcinoma”. The proposed strategy outperformed three human experts and five ML classifiers, which offers prospects for alleviating discrepancies noticed in endometrial tumor grading. Ergo, the adoption of ML techniques is projected to obviate workloads and shortages while permitting a more objective measure of clinical and surgical pathology. Subjectivity issue apart, invasiveness is debated in acquiring biopsy tissues.
Discovering non-invasive biomarkers has been a priority in cancer research for three decades, and ML-radiomics nomograms play a pivotal part in its pace (Jha et al. 2022)—H &E-based computational biomarkers are imminent and revolutionizing (Lancellotti et al. 2021). In a retrospective study of 435 women, Kawakami et al. (2019) confirmed the power of popular ML classifiers in epithelial ovarian cancer diagnosis using 32 preoperative blood biomarkers before any initial intervention. Song et al. (2018) explored 16 serum biomarker profiles in search of alternative marker pairings to lessen misdiagnosis. In another study, Aljakouch et al. (2019) revealed the potential of all Raman spectra, CARS, SHG, and TPF imaging as fast non-invasive CxCa diagnosis tools. Elias et al. (2017) suggested circulating miRNAs to establish a non-invasive OC diagnostic test, if it is validated, it could also screen women at high risk. Other research for precise diagnoses through non- or minimally invasive modalities has been undertaken, e.g. serum metabolomic fingerprints (Troisi et al. 2020), MRI (Urushibara et al. 2021; Nakagawa et al. 2019), US (Acharya et al. 2012), etc. Such synergy with DM might be a viable solution, notably, in (i) preoperative lymph node metastasis (LNM) diagnosis (Chen et al. 2021; Bedrikovetski et al. 2021; Wu et al. 2020) and (ii) worrisome-benign masses or cysts cases (Wasnik 2013). As per LNM, non-invasive and preoperative tools are compelled to discern between low-risk patients and those with substantial nodal involvement—a process generally supported by the presence of non-sentinel-lymph-node involvement in pelvic and para-aortic lymph nodes (Reijnen et al. 2020). Withal, non-invasive alternatives to biopsy referral eliminate adverse scenarios, e.g. patient discomfort, mental pressure, hemorrhage, soreness, etc (Bagaria et al. 2021). Recall that diagnoses govern therapy, postoperative prognostication, and management of cancer patients (Smeltzer et al. 2021), so properly performing ML-ML-diagnostic tools impacts the in-follow routines.
Early detection greatly affects survival rates and “avoidable deaths” (Wardle et al. 2015). It is supplied with two constituents: (i) screening and (ii) early diagnosis, or downstaging. Due to the dismal prognosis of GYN malignancies, screening (36%) is pivotal with forecast demand outweighing that of diagnosis. But while regular HPV-DNA testing, cervical cytopathology, direct visual inspection, and colposcopy, could preclude cervical cancer, screening for endometrial and ovarian cancers remains controversial. Till now no viable screening option is available for EC (Troisi et al. 2020). As per screening OC, transvaginal ultrasonography is often used in conjunction with the cancer antigen (CA)-125 biomarker (Henderson et al. 2018), yet neither has been found accurate in routine clinical use as increased preoperative serum CA-125 levels are diverse by other conditions beyond ovarian cell proliferation (Song et al. 2018). A reason behind “all except fifteen” screening-related articles aligning to cervical lesions, the remainder pertains to EC, OC, and leiomyoma. However, incorporating ML algorithms in cancer research has introduced a promising form of triage and screening, along with a unique opportunity to overcome umpteen limitations of either existing or potential screening tests. From a public health standpoint, it is imperative to leverage ML schemes for the early detection of GYN cancers. Here, Troisi et al. (2018) pilot study and related multicenter clinical validation (Troisi et al. 2020) evinced a non-invasive and low-cost endometrial cancer screening system based on metabolomics signature and ensemble classifiers. Mabwa et al. (2021) suggested the potential inclusion of dried biofluids mid-infrared spectroscopy as an EC screening tool based on ML classifiers. In another study, Barnabas et al. revealed the potential of microvesicle proteomic biomarkers in identifying all Stage-I ovarian lesions. Tsai et al. (2014) study provides the first evidence to associate ZNF509 and MTMR2 oncogenes to ovarian cancer through SVM. Elias et al. (2017) denoted that miRNAs recognized by neural networks are also found in the fallopian tube epithelium early lesions, suggesting the likelihood of pre-metastatic screening. This is significant in the sense that high-grade ovarian cancer is often detected at metastatic stages due to no early-stage-specific biomarkers and asymptomatism.
GYN cancerous cells often metastasize within the peritoneal cavity (Burg et al. 2020), with a range of events stemming from treatment regimens (Horvath et al. 2013) and disease advanced-stage. Hence, the selected articles coping with the treatment task (6.1%) frequently had connections to outcomes, complications, relapse, risk stratification, and/or decision-making of individualized treatments, noticeably: (i) predict the procedure outcome of uterine artery embolization for uterine fibroids as either symptom resolution/improvement or worsening symptoms (Luo et al. 2020), (ii) predict residual disease (R0) resection following cytoreductive surgery for advanced epithelial ovarian cancer (aEOC) (Laios et al. 2020), (iii) predict rectovaginal or vesicovaginal fistula formation when using interstitial high-dose-rate brachytherapy for locally advanced or recurrent GYN tumors unapt for intracavitary applicator treatments (Tian et al. 2019), (iv) comparing pre-treatment, post-treatment, and normal control cases by dint of OC DNA methylation data (Jiang and Liang 2020), (v) predict rectum dose-toxicity for CxCa combined external beam radiotherapy and brachytherapy (Zhen et al. 2017), (vi) predict local relapse and distant metastasis for locally advanced CxCa patients with definitive chemoradiotherapy-treated (Shen et al. 2019), (vii) predict post-treatment health related QoL in CxCa patients on the functional, symptom, and global health/QoL scales (Kumar et al. 2014), (viii) predict whether patients with locally advanced CxCa would be cured or relapsed after CRT (Torheim et al. 2014), (ix) predict CxCa recurrence by dint of DNA methylation data (Ma et al. 2021), (x) predict adverse events of radical hysterectomy in FIGO IA2–IIB CxCa patients (Kusy et al. 2013), (xi) predict platinum-free interval (PFI) of serous ovarian cancer patients by dint of histopathology and functional omics (Yu et al. 2020a), (xii) identify preoperatively EC patients at risk for LNM and poor outcome for personalized adjuvant treatment (Reijnen et al. 2020), (xiii) predict venous thromboembolism events in OC patients, e.g. venous thromboembolism (VTE) and deep vein thrombosis (DVT) (Fresard et al. 2020), and (xiv) identify high-risk of mortality OC patients by dint of genetic signatures (Hsiao et al. 2021).
Prognosis garnered 13.6% as ML-based prognostic prediction is a crucial paradigm in the realm of personalized medicine. It uncovers notably prognostic/predictive biomarkers (incl. candidate oncogenes and tumor suppressor genes), tumor recurrence, survival, and withal individualized treatment improvement (Laios et al. 2020). Monitoring received the least attention (2.3%); it requires therefore in-depth scrutiny as a vital facet of patient care entailing an innovative revolution. No management tasks-related articles were perceived.
4.6 Aligning DM analytics with GYN oncology
In theory, screening and diagnosis medical tasks are categorization matters encompassing several hurdles like FIGO staging, subtyping, TNM system, and so forth. It is therefore no wonder classification remains the subject of most research as far as screening and diagnosis are concerned in line with Sect. 3.7 data extraction. Regression (12.76%) comes second, accounting for prognosis-related use in the present selection study. Conventionally, traditional statistics analyses have been the mainstream of oncologic prognostic research, yet they are ineffective in coping with high-dimensional data and non-linear correlations (Rajula et al. 2020). Even though DM approaches are mostly based on probability and statistics, they are more resilient than parametric models allowing complex pattern identification across massive datasets with multi-input variables as accustomed to clinicians (Laios et al. 2020). Likewise, in silico ML-based regression approaches are gaining deeper insights - particularly in (i) asserting pathogenic (Cytosine-phosphate guanine) CpG sites and corresponding genes, where statistical approaches are unsuitable for DNA methylation (DNAm) data, as reported in Jiang and Liang (2020) study, and (ii) clinical outcomes such as disease-free survival (DFS), which is the utmost challenging endpoint in oncology, as reported in Wu et al. (2020) study. Clustering (11.06%) methods appear to be very useful in prognostic systems and gene expression assays. Further research of genetic clusters depending on miRNA expression patterns within training cohort samples could improve outcomes and proffer a better understanding of functional genomics (Lopez et al. 2018; Oyelade et al. 2016). Association analysis (2.98%) also permits the pinpoint of expressed methylated gene relationships in distinct samples and the shift from individual components to associative patterns amongst predisposing genes (Mallik et al. 2013; Lee et al. 2021). Hence, ARM-based algorithms distinctly improve tumor prediction and carcinogenesis, as Mallik et al. (2013) have proven for uterine leiomyoma prediction.
Most research adopted NNs, SVMs, and RFs. NNs are adequate to discern patterns, manage large or noisy data sets, and interestingly, learn in a manner reminiscent of the human brain (Yang and Wang 2020). Amongst the most implemented NN classes are CNNs. CNNs offer great prospects for addressing conventional medical CADx hurdles (Chan et al. 2020) due to their ability to (i) operate as both a feature extraction module and a classifier, (ii) to benefit from transfer learning to overcome poor generalization, overfitting, and the need for large databases supplements, more particularly for confirmed cases, and (iii) to cope with high-dimensional, heterogeneous, and complex input data as with biomedical imaging, particularly the arduous whole-slide imaging. For instance, Yang and Stamp (2021) preprocessed and patched uterine tissue biopsy WSIs from 250 potential LGESS patients for classification, in which DL outperformed classical ML models with the highest accuracy. Xue et al. (2019b) demonstrated the effectiveness of ResNet as a baseline classifier for the CIN grade classification problem using segmented and GAN-generated epithelium patches. Shin et al. (2021) a successful case of Inception V3 generalized through a style transfer technique in small image sets within the domain of ovarian digital pathology. Cheng et al. (2020) generated a structured report to support the interpretable analysis of cervical pathology images by using CNN-extracted features as input for an LSTM. Meng et al. (2021) proposed assembling a CNN-based classification, segmentation, and pseudo-labeling approach to enhance the performance on unlabeled cervical histopathology data. Zhang et al. (2021c) experiment used ResNet50 as the feature extractor coupled with several classical ML models to perform cervical tissue pathological images. Yu et al. (2020b) showcased the utility of CNNs in identifying tumor cells, and classifying both tumor grades and transcriptomic subtypes. Sun et al. (2020) developed a CAD based on a CNN and attention mechanisms to model the complicated relationships between endometrium histopathological images and related interpretations, where classical ML often fail to achieve satisfying findings. Huang et al. (2020b) investigated a novel cervical cancer classification method, analyzing the impact of fusing convolutional features from various depths and combining different classifiers on the accuracy of pathological image classification. Despite the success of DL in tackling several limitations of classical ML, most of the aforesaid raised a pressing need for causability to achieve a level of explainability and interpretability in DL pathology, aiming to develop a framework that can be seamlessly applied in clinical settings or within an inter-institutional heterogeneity as DL exhibits poor performance when used on disparate data from external sources. As performances of DL networks are proportional to the number of hidden layers—the greater the depth, the more robust a CNN Mostafa et al. (2022), going deeper would exacerbate deep neural network issues, which are akin to a ‘black box’ Mostafa et al. (2022). The inability to be interpreted yields NNs untrustworthy by the medical community. On the other hand, SVM is a potent approach used for classification and regression analyses that accommodates biological data due to their non-linear character such as genomics profiles (Huang et al. 2018; Sidey-Gibbons and Sidey-Gibbons 2019). It is recognized by the scientific community that SVM is an accurate classifier superior to neural classifiers Hirschberg et al. (2020). Yet, SVMs are greedy in terms of memory usage as well as computational time (Guo and Boukir 2015). A powerful scenario is to combine the SVM classifier with the bottleneck extracted features by a CNN as seen in Shao et al. (2020), Sun et al. (2020) studies or in the aforesaid instances. RF was used in 45 studies as it provides a robust classification with little overfitting and is simple to comprehend (Sarica et al. 2017). A further benefit of RFs is the identification of important features through the typical use of the Gini score, which allows biomarkers pinpointing as seen in Kawakami et al. (2019) study.
4.7 Assessing DM performance in GYN oncology
In literature, the preferred performance metrics are accuracy, sensitivity, specificity, and AUROC which, in turn, are in the present selection study. Whereas accuracy and precision disclose the test’s general reliability, both specificity and sensitivity highlight the likelihood of false positives (FPs) and false negatives (FNs) (Eusebi 2013). Pertaining to biomedical data, accuracy might not constitute a realistic measure since higher scores do not matter to evaluate the model as a proper medical task. Considering a simple triage of patients within imbalanced data, the model might enjoy a higher accuracy in scoring the dominant samples class, but sacrificed sensitivity in predicting the minority class (Tian et al. 2019; Wu and Zhou 2017). To get a profuse explanation, extending the evaluation by including other metrics is a must (Wu and Zhou 2017). Meanwhile, mean absolute error (MAE) and concordance index (c-index) were suitable in regression models by providing the ratio of the real and predicted value, which perfectly matches such type of task. Due to the easy but valuable handling of variations in oncological data, cross-validation was utilized largely by 62.43% of the selected articles as seen in Fig. 7. Next in line is the ’hold-out’ method, comprising 28.42% of the selected articles, as it involves the straightforward task of dividing a dataset into subsets. But while hold-out is less computationally costly, cross-validation schemes provide more meaningful insights regarding model performance on more complex and unseen data at every point, which circumvents any data bias concerns.
The ongoing shortage of human assessments underscores the inadequacy of research in DM as an adjunct to GYN oncology. More evidence, including comparative studies with clinicians, is needed, extending well beyond a mere emphasis on collaboration. This is crucial to substantiate the practical necessity in real clinical practice and to enhance DM role comprehension. For instance, Urushibara et al. (2021) demonstrated CNN potency in assisting radiologists in the interpretation of pelvic MRIs. The interobserver agreement between three radiologists (9–27 experience years) and CNN was observed to be lower than among the radiologists themselves. Such discrepancy suggests that the model might have employed a distinct perspective compared to humans, leading to differing judgments. Zhang et al. (2021d) demonstrated that the diagnostic performance of VGGNet-16 in classifying five endometrial lesions was on par with that of expert gynecologists, providing hysteroscopists with objective diagnostic evidence and potential clinical utility. Further, enhancing colposcopy precision is a crucial aspect of CIN management. Even among experienced colposcopists, the sensitivity for detecting CIN through colposcopy ranges from 81.4 to 95.7%, while specificity ranges from 34.2 to 69%. In comparison, Yuan et al. (2020) revealed that a DL diagnostic system exhibited superior performance when compared to colposcopists in the analysis of standard cervicograms for HSILs. However, its performance showed a slight decline when dealing with high-definition cervicograms. Bao et al. (2020) joined with AI-assisted cytology, achieving a level of performance in the detection of CIN2+ that was comparable to that of skilled cytologists in referring women. Integrating AI-assisted cytology into primary cytology screening, the most prevalent application in pathology, or HPV-positive triage, would improve specificity while not at the expense of sensitivity. As interobserver agreement remains a critical issue in GYN oncology, both studies have proved the potential of a stable and objective CADx system to lessen interobserver error and potentially mitigate the learning curve for less experienced gynecologists. The computerized models could also exhibit a more time-efficient performance as seen in Chen et al. (2020) study, where the DL network model, derived from MR imaging, delivered a competitive and time-efficient diagnostic performance in identifying MI depth. Exploring the DM-ML performance in therapy and its role in aiding clinical decision-making is needed. Further validation through human assessments might predict the malignancy risk of masses before surgical procedures (Gao et al. 2022) and enable better patient identification for those who can benefit the most from treatments, as seen in the promising results of Luo et al. (2020) study regarding the prediction of outcomes in uterine fibroid embolization.
Of note, it is worthwhile to highlight the efforts made by researchers to validate the implemented models.
5 Related work
In spite of the booming interest in Gynecologic AI, the topic has been marginally contemplated only in a handful of reviews. The related work primarily explores the topic of the convergence of AI and GYN oncology or categorizes it as a distinct subfield within software engineering.
Three astoundingly valuable systematic reviews scrutinized the application of AI algorithms in the context of female oncology. Such reviews encompassed varied directions, posing a challenge when attempting direct comparisons with the current review. Shrestha et al. (2022) consolidates the current state of the art in the field of gynecologic cancer imaging and provides insights for moving research and clinical integration of the analytics forward. The authors came with two pertinent challenges: generalizability and reproducibility while raising concerns about the paucity of clinicians vs. AI assessments and the relatively limited research conducted in the field of Gynecologic AI in comparison to Breast AI. Such discrepancy is particularly striking when comparing the volume of publications in the domains with the aggregated mortality rates of gynecologic cancers versus breast cancer. Still, only imaging was analyzed, which not only inhibits generalizability but pinpoints the need for further identifying other modalities. This was also apparent in Xue et al. (2022) study restricted even more to only cervical and breast cancers through DL which require further pathologies from the same anatomical site. Yet, the authors identified kindred observations in alignment with the current SLR in relation to (i) bolstering the current evidence through high-quality research such as prospective cohorts and clinical trials; (ii) generalizing prophase DL on larger and multi-institute data with a range of nationalities, ethnicities, socioeconomic status, and so forth; (iii) integrating DL into real-clinical settings throughout comprehensive healthcare systems while ensuring robustness and scientific validation for clinical and personal benefit; (iv) conducting a comparative analysis of DL against human clinicians, esp. sensitivity and specificity, as the existing study designs and reporting remain poor, which could lead to an overestimation of algorithmic performance; and (v) developing global and standardized reporting guidelines for DL in medical imaging to streamline the integration into clinical practice. Akazawa and Hashimoto (2021) revealed that a pioneer in CADx could involve utilizing Pap smears and digital colposcopy for the diagnosis of gynecologic lesions while also raising the abundance of single-institute retrospective studies concern and the high heterogeneity found. Still, the authors’ tentative assertion is based on only 71 studies within three databases and without quality appraisal for relatively poor designs and reporting, which underscores the necessity for more rigorous and comprehensive scrutiny of the available data.
Herein, it is imperative to embark on the available DM-ML quest and subsequently enhance its capabilities for heightened accuracy in potential clinical unmet ethically. Thereupon, continued reliance on systematic reviews is warranted to pinpoint research gaps, flaws, and limitations. Such extends not only to the prioritization of technology-specific reviews but also underscoring the significance of disease(site)-specific systematic reviews (Shrestha et al. 2022). While intelligent algorithmic remains in a perpetual state of evolution, our aspiration was to provide recommendations and catalyze further exploration within the topic. Such effort is intended to accelerate the adoption of the most potent technology into clinical practice at the earliest opportunity.
6 Implications for practice
Summarized evidence acknowledges the inclusion of ML modules in GYN oncology. Still, research has centered on sophisticated ML models rather than data peculiarities and unmet medical needs. An insufficient clinical trial validation could further activate fault lines, yielding attempts likely to be unsustainable systems in oncology settings. Arising from (yet not limited to) the findings of this SLR, pivotal considerations for proper DM-ML implementation in GYN oncology are:
-
1.
Generalizability (akin to external validity). The ability of an ML model to execute satisfactorily on new, independent cohorts across disparate populations, protocols, and deployment environments. When the model fails to meet this definition, it has learned biases. Lingering concerns center on too small datasets, restricted to single institutional/regional sampling, exhibiting a population non-diversity, noise/artifacts, or variables in settings. Such datasets could be socio-demographics skewed, with a process chain inadvertently encoding biases, thus not representative of the broader population. Although recommended, generalizability is not always within reach due to ethics, budgeting, and many more considerations in data sharing. Hence, hospital collaborations using model-customization techniques are needed to translate site-specific trained models to new contexts, else generalizability remains seldom practiced in literature.
-
2.
Prospective Cohorts. Data assessment of recruited subjects before outcomes, thus proving temporality, is a core factor in asserting causality. A prospective design is valued more highly in the evidence pyramid than a retrospective one (historical cohort) in which events have already occurred. It is a potent study design inferring the true benefit of ML schemes in real-world clinical settings and yielding, often, generalizable outcomes. Although encouraged, it is not always financially practical, as a prospective cohort demands eligible subjects to be followed up over a lengthy time frame. Bias or confounding ought to be prevented to the greatest extent, particularly regarding losses or individuals dropping out. Prospective cohorts remain seldom practiced in literature.
-
3.
Randomized Controlled Trials. Prospective trials in which subjects are randomly assigned to either an intervention or control group. Such are often blinded and occur after the recruitment and eligibility phasis. The act of randomization is the best clinical evidence to assess an intervention’s effects while mitigating confounding factors. While it is not practical to formally assess each model iteration through a randomized controlled trial, it would clearly quantify the clinical value of an ML system that has already reached the clinical implementation phase. Further, it provides another level of evidence to encourage ML adoption in clinical practice, as a model utility and bias can only be ascertained by deciphering the mechanistic of predictions. Due to generalizability issues, a randomized controlled trial at baseline followed by a prospective external validity is advised despite each expenditure.
-
4.
Trustworthy AI. The evolving and intricate term “trustworthy” constitutes, at its core, concerns surrounding AI-ML use, such as robustness, generalization, interpretability/explainability, fairness, transparency, accountability, responsibility, privacy, ethics, safety, reproducibility, intellectual property, so forth. Unfortunately, overly optimistic findings from intelligent systems often occur at the expense of degradation in their trustworthiness. Fostering research is required in, e.g. (i) representing global ethnic, socioeconomic, and demographic groups to mitigate equity challenges that might impact existing marginalized and underrepresented groups; (ii) reporting, data sharing, metrics, reproducibility, and traceability; and (iii) the algorithmic “black box” as a threat to core principles of procedural fairness. It is also imperative to establish international technology-specific and disease-specific guidelines for gynecologic AI.
-
5.
Human-in-the-Loop AI. While it is imperative to acknowledge the value of AI in modern clinical practice, it is equally crucial to uphold the bedrock of patient-centric tenets and clinical experience in all AI implications. A decent use involves it as a complement to physicians, not as a substitute. AI, as ’a tool, not a crutch’, should only be integrated if it satisfies the following three criteria: (i) respecting the ethical conduct and credo of clinical practice, (ii) serving solid clinical ends, and (iii) enhancing human interaction. Unfortunately, many research endeavors fall short of satisfaction, with the ’human-in-the-loop’ dimension notably absent yet quintessential component.
7 Conclusion and perspectives
The current SLR examines the relevance of ML to GYN oncology. From the 2807 potential records obtained from PubMed, IEEE Xplore, ScienceDirect, Springer Link, and Google Scholar, 181 papers published over the past decade were assessed in depth. We found that the use of ML in GYN oncology soared markedly in 2019—most notably regarding cervical neoplasms; the bulk of articles were published in journals. Most research focused on cervical cancer as it is a paradigm of global health inequity, with higher morbimortality rates than both uterine and ovarian malignancies combined. Ovarian cancer comes second by cause of lethality, then endometrial cancer since it represents a significant global health disparity in developed countries. However, few articles leveraged machine learning to tackle other types of GYN cancers entailing more research. The majority were solution proposals, all of which were empirically evaluated. However, a paucity of validation research is noticed. Most of the selected articles were empirically evaluated, with more than half referring to case studies but only a few are based on surveys. Clinical records gathered great interest, followed by Pap smears, cervicograms, liquid-based cytology, then ’biopsy, D &C, or hysterectomy’ WSI. An evolving use despite the challenges encountered in medical data. The most investigated task was the diagnosis. Minimal invasive sampling has been shown through ML promising results, however, the potential value of screening among the asymptomatic must also be evaluated. Most of the selected articles investigated the classification task. Neural networks (NNs) were prevalent and devoted to respective classification and regression. Articles employed a variety of performance metrics, with accuracy, sensitivity, specificity, and AUROC being the most popular combination. Cross-validation was used extensively in all validation schemes.
We must acknowledge the present SLR limitations. First, although we rigorously broadened the databases and search terms, it was still probable that some research was omitted. Second, as the extent is limited to English material published within the past decade, we might have missed prior research to this time (or conducted in a different language). Third, the undertaken quality assessment could have screened out valuable articles providing differing findings. Fourth, we were unable to conduct a quantitative analysis of the evidence as it fell outside this SLR scope due to the heterogeneity in our selection study. We also could not thoroughly analyze the bias risks or integrity of the reported articles due to related gaps.
Future perspectives include: (i) Signs of ovarian and endometrial cancer are hazy and commonly present in benign conditions, resulting in usually being diagnosed in late stages when survival rates are poor. Due to the rapid growth of DM-ML in cancer prediction, non-invasive and low-cost screening systems from several biomarkers sources have garnered a research interest as a promising form of triage. A critical survey of DM utility in early-stage OC/EC among the asymptomatic may provide impetus to research, providing also opportunities for drug discovery. (ii) Deep learning has undeniably exerted a substantial influence across various scientific domains in recent years. This has been evidenced in practices where approaches surpassed conventional methods, yielded commendable outcomes in previously unattainable tasks through automation, and even excelled in applications surpassing human capabilities. Now, GYN oncology is where DL progress has started to exhibit immense potential. While the promise is evident, concerns arise about the practical usability of the research output, often confined to ideal scenarios as real-life examples have recently demonstrated fallibility, raising the common question: ‘How can we place trust in neural networks to avoid errors and biases when we lack understanding of their decision-making processes?’. Most of the evidence in the current SLR concentrated on a global technicality. It would be interesting to delve in-depth into the actual evidence and what the future holds for the field of GYN deep learning. (iii) Assessing the performance of ML/DL models is imperative to evaluate the predictive ability across various populations and settings and to determine further improvements through limitations relevant to reported studies. As a future scope, we aim to systematically review, with meta-analysis, the predictive performance of the models used in this SLR study. Such will be based on the existing evidence on a particular medical task, prediction task, and GYN cancer. Based on a similar strategy adopted by Wadghiri et al. (2022), we aim to provide guidance on which performance metrics could be extracted from the selection study considering each evaluation and optimal configuration, associated statistical measures, why they are important, and how to handle circumstances when missing or poorly reported.
References
Acharya U, Mookiah M, Sree SV, Yanti R, Martis R, Saba L et al (2012) Evolutionary algorithm-based classifier parameter tuning for automatic ovarian cancer tissue characterization and classification. Ultraschall in der Medizin 35(03):237–245. https://doi.org/10.1055/s-0032-1330336
Adamo JE, Bienvenu RV, Fields FO, Ghosh S, Jones CM, Liebman M et al (2018) The integration of emerging omics approaches to advance precision medicine: how can regulatory science help? J Clin Transl Sci 2(5):295–300. https://doi.org/10.1017/cts.2018.330
Akazawa M, Hashimoto K (2021) Artificial intelligence in gynecologic cancers: current status and future challenges–a systematic review. Artif Intell Med 120:102164. https://doi.org/10.1016/j.artmed.2021.102164
Aljakouch K, Hilal Z, Daho I, Schuler M, Kraus SD, Yosef HK et al (2019) Fast and noninvasive diagnosis of cervical cancer by coherent anti-stokes Raman scattering. Anal Chem 91(21):13900–13906. https://doi.org/10.1021/acs.analchem.9b03395
Allison KH, Reed SD, Voigt LF, Jordan CD, Newton KM, Garcia RL (2008) Diagnosing endometrial hyperplasia. Am J Surg Pathol 32(5):691–698. https://doi.org/10.1097/pas.0b013e318159a2a0
Asaduzzaman S, Ahmed MR, Rehana H, Chakraborty S, Islam MS, Bhuiyan T (2021) Machine learning to reveal an astute risk predictive framework for gynecologic cancer and its impact on women psychology: Bangladeshi perspective. BMC Bioinformatics. https://doi.org/10.1186/s12859-021-04131-6
Asiedu MN, Simhal A, Chaudhary U, Mueller JL, Lam CT, Schmitt JW et al (2019) Development of algorithms for automated detection of cervical pre-cancers with a low-cost, point-of-care, pocket colposcope. IEEE Trans Biomed Eng 66(8):2306–2318. https://doi.org/10.1109/tbme.2018.2887208
Bagaria M, Wentzensen N, Clarke M, Hopkins MR, Ahlberg LJ, Guire LJM et al (2021) Quantifying procedural pain associated with office gynecologic tract sampling methods. Gynecol Oncol 162(1):128–133. https://doi.org/10.1016/j.ygyno.2021.04.033
Bao H, Bi H, Zhang X, Zhao Y, Dong Y, Luo X et al (2020) Artificial intelligence-assisted cytology for detection of cervical intraepithelial neoplasia or invasive cancer: a multicenter, clinical-based, observational study. Gynecol Oncol 159(1):171–178. https://doi.org/10.1016/j.ygyno.2020.07.099
Barnabas GD, Bahar-Shany K, Sapoznik S, Helpman L, Kadan Y, Beiner M et al (2019) Microvesicle proteomic profiling of uterine liquid biopsy for ovarian cancer early detection. Mol Cell Proteomics 18(5):865–875. https://doi.org/10.1074/mcp.ra119.001362
Bedrikovetski S, Dudi-Venkata NN, Kroon HM, Seow W, Vather R, Carneiro G et al (2021) Artificial intelligence for pre-operative lymph node staging in colorectal cancer: a systematic review and meta-analysis. BMC Cancer. https://doi.org/10.1186/s12885-021-08773-w
BenTaieb A, Li-Chang H, Huntsman D, Hamarneh G (2017) A structured latent model for ovarian carcinoma subtyping from histopathology slides. Med Image Anal 39:194–205. https://doi.org/10.1016/j.media.2017.04.008
Boyce B (2017) An update on the validation of whole slide imaging systems following FDA approval of a system for a routine pathology diagnostic service in the united states. Biotech Histochem 92(6):381–389. https://doi.org/10.1080/10520295.2017.1355476
Brereton P, Kitchenham BA, Budgen D, Turner M, Khalil M (2007) Lessons from applying the systematic literature review process within the software engineering domain. J Syst Softw 80(4):571–583. https://doi.org/10.1016/j.jss.2006.07.009
Brüggmann D, Ouassou K, Klingelhöfer D, Bohlmann MK, Jaque J, Groneberg DA (2020) Endometrial cancer: mapping the global landscape of research. J Transl Med. https://doi.org/10.1186/s12967-020-02554-y
Burg L, Timmermans M, van der Aa M, Boll D, Rovers K, de Hingh I, van Altena A (2020) Incidence and predictors of peritoneal metastases of gynecological origin: a population-based study in the Netherlands. J Gynecol Oncol. https://doi.org/10.3802/jgo.2020.31.e58
Chan H-P, Hadjiiski LM, Samala RK (2020) Computer-aided diagnosis in the era of deep learning. Med Phys. https://doi.org/10.1002/mp.13764
Chardin L, Leary A (2021) Immunotherapy in ovarian cancer: thinking beyond PD-1/PD-l1. Front Oncol. https://doi.org/10.3389/fonc.2021.795547
Chen D, Xing K, Henson D, Sheng L, Schwartz AM, Cheng X (2009) Developing prognostic systems of cancer patients by ensemble clustering. J Biomed Biotechnol. https://doi.org/10.1155/2009/632786
Chen X, Wang Y, Shen M, Yang B, Zhou Q, Yi Y et al (2020) Deep learning for the determination of myometrial invasion depth and automatic lesion identification in endometrial cancer MR imaging: a preliminary study in a single institution. Eur Radiol 30(9):4985–4994. https://doi.org/10.1007/s00330-020-06870-1
Chen C, Qin Y, Chen H, Zhu D, Gao F, Zhou X (2021) A metaanalysis of the diagnostic performance of machine learning-based MRI in the prediction of axillary lymph node metastasis in breast cancer patients. Insights into Imaging. https://doi.org/10.1186/s13244-021-01034-1
Cheng H, Wu K, Ma K, Tian J, Xu R, Gu C, Guan X (2020) Double attention for pathology image diagnosis network with visual interpretability. In: International joint conference on neural networks (IJCNN). IEEE. https://doi.org/10.1109/ijcnn48605.2020.9206603
Cramer DW (2012) The epidemiology of endometrial and ovarian cancer. Hematol Oncol Clin N Am 26(1):1–12. https://doi.org/10.1016/j.hoc.2011.10.009
dos Santos FLC, Wojciechowska U, Michalek IM, Didkowska J (2023) Survival of patients with cancers of the female genital organs in Poland, 2000–2019. Sci Rep. https://doi.org/10.1038/s41598-023-35749-6
Downing MJ, Papke DJ, Tyekucheva S, Mutter GL (2019) A new classification of benign, premalignant, and malignant endometrial tissues using machine learning applied to 1413 candidate variables. Int J Gynecol Pathol 39(4):333–343. https://doi.org/10.1097/pgp.0000000000000615
Duque J, Moreira JJ, Costa J (2023) Data mining to support decision-making-a research approach. Intelligent sustainable systems. Springer, Singapore, pp 553–563
Elias KM, Fendler W, Stawiski K, Fiascone SJ, Vitonis AF, Berkowitz RS, et al (2017) Diagnostic potential for a serum miRNA neural network for detection of ovarian cancer. eLife. https://doi.org/10.7554/elife.28932
Esfandiari N, Babavalian MR, Moghadam A-ME, Tabar VK (2014) Knowledge discovery in medicine: current issue and future trend. Expert Syst Appl 41(9):4434–4463. https://doi.org/10.1016/j.eswa.2014.01.011
Eusebi P (2013) Diagnostic accuracy measures. Cerebrovasc Dis 36(4):267–272. https://doi.org/10.1159/000353863
everhobbes (n.d.) Ovarian Cancer Key Stats* - worldovariancancercoalition. org. https://worldovariancancercoalition.org/about-ovarian-cancer/key-stats/. Accessed 22 May 2022
Farooq A, Abdelkader A, Javakhishivili N, Moreno GA, Kuderer P, Polley M et al (2021) Assessing the value of second opinion pathology review. Int J Qual Health Care. https://doi.org/10.1093/intqhc/mzab032
Fiscutean A (2021) Clarifying the burden of ovarian cancer. Nature 600(7889):S48–S49. https://doi.org/10.1038/d41586-021-03719-5
Fresard ME, Erices R, Bravo ML, Cuello M, Owen GI, Ibanez C, Rodriguez- Fernandez M (2020) Multi-objective optimization for personalized prediction of venous thromboembolism in ovarian cancer patients. IEEE J Biomed Health Inform 24(5):1500–1508. https://doi.org/10.1109/jbhi.2019.2943499
Gao Y, Zeng S, Xu X, Li H, Yao S, Song K et al (2022) Deep learning-enabled pelvic ultrasound images for accurate diagnosis of ovarian cancer in china: a retrospective, multicentre, diagnostic study. Lancet Digital Health 4(3):e179–e187. https://doi.org/10.1016/s2589-7500(21)00278-8
Genta RM (2014) Same specimen, different diagnoses. Adv Anat Pathol 21(3):188–190. https://doi.org/10.1097/pap.0000000000000023
Girolamo F, Lante I, Muraca M, Putignani L (2013) The role of mass spectrometry in the “omics" era. Curr Org Chem 17(23):2891–2905. https://doi.org/10.2174/1385272817888131118162725
Gravitt PE, Silver MI, Hussey HM, Arrossi S, Huchko M, Jeronimo J et al (2021) Achieving equity in cervical cancer screening in low- and middle-income countries (LMICs): strengthening health systems using a systems thinking approach. Prev Med 144:106322. https://doi.org/10.1016/j.ypmed.2020.106322
Greyson D, Rafferty E, Slater L, MacDonald N, Bettinger JA, Dubé È, MacDonald SE et al (2019) Systematic review searches must be systematic, comprehensive, and transparent: a critique of perman. BMC Public Health. https://doi.org/10.1186/s12889-018-6275-y
Grimley PM, Liu Z, Darcy KM, Hueman MT, Wang H, Sheng L et al (2021) A prognostic system for epithelial ovarian carcinomas using machine learning. Acta Obstet Gynecol Scand 100(8):1511–1519. https://doi.org/10.1111/aogs.14137
Guo L, Boukir S (2015) Fast data selection for SVM training using ensemble margin. Pattern Recogn Lett 51:112–119. https://doi.org/10.1016/j.patrec.2014.08.003
Guo P, Banerjee K, Stanley RJ, Long R, Antani S, Thoma G et al (2016) Nuclei-based features for uterine cervical cancer histology image analysis with fusion-based classification. IEEE J Biomed Health Inform 20(6):1595–1607. https://doi.org/10.1109/jbhi.2015.2483318
Hanna MG, Reuter VE, Ardon O, Kim D, Sirintrapun SJ, Schüffler PJ et al (2020) Validation of a digital pathology system including remote review during the COVID-19 pandemic. Mod Pathol 33(11):2115–2127. https://doi.org/10.1038/s41379-020-0601-5
Henderson JT, Webber EM, Sawaya GF (2018) Screening for ovarian cancer. JAMA 319(6):595. https://doi.org/10.1001/jama.2017.21421
Hirschberg C, Edinger M, Holmfred E, Rantanen J, Boetker J (2020) Image-based artificial intelligence methods for product control of tablet coating quality. Pharmaceutics 12(9):877. https://doi.org/10.3390/pharmaceutics12090877
Holsbeke CV, Calster BV, Bourne T, Ajossa S, Testa AC, Guerriero S et al (2012) External validation of diagnostic models to estimate the risk of malignancy in adnexal masses. Clin Cancer Res 18(3):815–825. https://doi.org/10.1158/1078-0432.ccr-11-0879
Horvath S, George E, Herzog TJ (2013) Unintended consequences: surgical complications in gynecologic cancer. Womens Health 9(6):595–604. https://doi.org/10.2217/whe.13.60
Hosni M, Abnane I, Idri A, de Gea JMC, Alemán JLF (2019) Reviewing ensemble classification methods in breast cancer. Comput Methods Programs Biomed 177:89–112. https://doi.org/10.1016/j.cmpb.2019.05.019
Hsiao Y-W, Tao C-L, Chuang EY, Lu T-P (2021) A risk prediction model of gene signatures in ovarian cancer through bagging of GA-XGBoost models. J Adv Res 30:113–122. https://doi.org/10.1016/j.jare.2020.11.006
Huang S et al (2018) Applications of support vector machine (SVM) learning in cancer genomics. Cancer Genomics Proteomics. https://doi.org/10.21873/cgp.20063
Huang P, Tan X, Chen C, Lv X, Li Y (2020) AF-SENet: classification of cancer in cervical tissue pathological images based on fusing deep convolution features. Sensors 21(1):122. https://doi.org/10.3390/s21010122
Huang P, Zhang S, Li M, Wang J, Ma C, Wang B, Lv X (2020) Classification of cervical biopsy images based on LASSO and EL-SVM. IEEE Access 8:24219–24228. https://doi.org/10.1109/access.2020.2970121
Hull R, Mbele M, Makhafola T, Hicks C, Wang S, Reis R et al (2020) Cervical cancer in low and middle-income countries (review). Oncol Lett 20(3):2058–2074. https://doi.org/10.3892/ol.2020.11754
Idlahcen F, Idri A (2022) Systematic map of data mining for gynecologic oncology. Information systems and technologies. Springer, pp 466-475
Idri A, Amazal F, Abran A (2015) Analogy-based software development effort estimation: a systematic mapping and review. Inf Softw Technol 58:206–230. https://doi.org/10.1016/j.infsof.2014.07.013
Idri A, Hosni M, Abran A (2016) Systematic literature review of ensemble effort estimation. J Syst Softw 118:151–175. https://doi.org/10.1016/j.jss.2016.05.016
Idri A, Benhar H, Fernández-Alemán J, Kadi I (2018) A systematic map of medical data preprocessing in knowledge discovery. Comput Methods Programs Biomed 162:69–85. https://doi.org/10.1016/j.cmpb.2018.05.007
Idri A, Chlioui I, Ouassif BE (2018) A systematic map of data analytics in breast cancer. In: Proceedings of the Australasian computer science week multiconference. ACM. https://doi.org/10.1145/3167918.3167930
Jha AK, Mithun S, Purandare NC, Kumar R, Rangarajan V, Wee L, Dekker A (2022) Radiomics: a quantitative imaging biomarker in precision oncology. Nucl Med Commun 43(5):483–493. https://doi.org/10.1097/mnm.0000000000001543
Jiang H-K, Liang Y (2020) Penalized logistic regression based on l1/2 penalty for high-dimensional DNA methylation data. Technol Health Care 28:161–171. https://doi.org/10.3233/thc-209016
Jo S (2022) The use of multiple imputation to handle missing data in secondary datasets: suggested approaches when missing data results from the survey structure. INQUIRY. https://doi.org/10.1177/00469580221088627
Kanavati F, Hirose N, Ishii T, Fukuda A, Ichihara S, Tsuneki M (2022) A deep learning model for cervical cancer screening on liquid-based cytology specimens in whole slide images. Cancers 14(5):1159. https://doi.org/10.3390/cancers14051159
Kawakami E, Tabata J, Yanaihara N, Ishikawa T, Koseki K, Iida Y et al (2019) Application of artificial intelligence for preoperative diagnostic and prognostic prediction in epithelial ovarian cancer based on blood biomarkers. Clin Cancer Res 25(10):3006–3015. https://doi.org/10.1158/1078-0432.ccr-18-3378
Keele S, et al (2007) Guidelines for performing systematic literature reviews in software engineering. Technical report, ver. 2.3 EBSE technical report. ebse
Kehoe S, Bhatla N (2021) FIGO cancer report 2021. Int J Gynecol Obstet 155(S1):5–6. https://doi.org/10.1002/ijgo.13882
Kitchenham B (n.d.) Evidence-based Software Engineering – keele.ac.uk. https://www.keele.ac.uk/research/ourresearch/computerscienceandmathematics/evidence-basedsoftwareengineering/#!. Accessed 22 May 2022
Kitchenham B, Dyba T, Jorgensen M (2004) Evidence-based software engineering. In: Proceedings 26th international conference on software engineering. IEEE Comput. Soc., pp 273–281. https://doi.org/10.1109/icse.2004.1317449
Krakauer EL, Kwete X, Kane K, Afshan G, Bazzett-Matabele L, Bien-Aimé DDR et al (2021) Cervical cancer-associated suffering: Estimating the palliative care needs of a highly vulnerable population. JCO Glob Oncol 7:862–872. https://doi.org/10.1200/go.21.00025
Kumar S, Rana ML, Verma K, Singh N, Sharma AK, Maria AK et al (2014) PrediQt-cx: Post treatment health related quality of life prediction model for cervical cancer patients. PLoS ONE 9(2):e89851. https://doi.org/10.1371/journal.pone.0089851
Kusy M, Obrzut B, Kluska J (2013) Application of gene expression programming and neural networks to predict adverse events of radical hysterectomy in cervical cancer patients. Med Biol Eng Comput 51(12):1357–1365. https://doi.org/10.1007/s11517-013-1108-8
Laios A, Gryparis A, DeJong D, Hutson R, Theophilou G, Leach C (2020) Predicting complete cytoreduction for advanced ovarian cancer patients using nearest-neighbor models. J Ovarian Res. https://doi.org/10.1186/s13048-020-00700-0
Lancellotti C, Cancian P, Savevski V, Kotha SRR, Fraggetta F, Graziano P, Tommaso LD (2021) Artificial intelligence & tissue biomarkers: advantages, risks and perspectives for pathology. Cells 10(4):787. https://doi.org/10.3390/cells10040787
LaVigne AW, Triedman SA, Randall TC, Trimble EL, Viswanathan AN (2017) Cervical cancer in low and middle income countries: addressing barriers to radiotherapy delivery. Gynecol Oncol Rep 22:16–20. https://doi.org/10.1016/j.gore.2017.08.004
Lee CKH, Tse YK, Ho G, Chung S (2021) Uncovering insights from healthcare archives to improve operations: an association analysis for cervical cancer screening. Technol Forecast Soc Chang 162:120375. https://doi.org/10.1016/j.techfore.2020.120375
Li C, Chen H, Zhang L, Xu N, Xue D, Hu Z et al (2019) Cervical histopathology image classification using multilayer hidden conditional random fields and weakly supervised learning. IEEE Access 7:90378–90397. https://doi.org/10.1109/access.2019.2924467
Li Y, Chen J, Xue P, Tang C, Chang J, Chu C et al (2020) Computer-aided cervical cancer diagnosis using timelapsed colposcopic images. IEEE Trans Med Imaging 39(11):3403–3415. https://doi.org/10.1109/tmi.2020.2994778
Li S, Chen H, Zhang T, Li R, Yin X, Man J et al (2022) Spatiotemporal trends in burden of uterine cancer and its attribution to body mass index in 204 countries and territories from 1990 to 2019. Cancer Med 11(12):2467–2481. https://doi.org/10.1002/cam4.4608
Liang LA, Einzmann T, Franzen A, Schwarzer K, Schauberger G, Schriefer D et al (2021) Cervical cancer screening: comparison of conventional pap smear test, liquid-based cytology, and human papillomavirus testing as stand-alone or cotesting strategies. Cancer Epidemiol Biomarkers Prev 30(3):474–484. https://doi.org/10.1158/1055-9965.epi-20-1003
Liang Y, Jiao H, Qu L, Liu H (2022) Association between hormone replacement therapy and development of endometrial cancer: results from a prospective US cohort study. Front Med. https://doi.org/10.3389/fmed.2021.802959
Liu Y, Ma L, Yang X, Bie J, Li D, Sun C et al (2019) Menopausal hormone replacement therapy and the risk of ovarian cancer: a meta-analysis. Front Endocrinol. https://doi.org/10.3389/fendo.2019.00801
Liu X, Xiao Z, Song Y, Zhang R, Li X, Du Z (2021) A machine learning-aided framework to predict outcomes of anti-PD-1 therapy for patients with gynecological cancer on incomplete post-marketing surveillance dataset. IEEE Access 9:120464–120480. https://doi.org/10.1109/access.2021.3107498
Lõhmussaar K, Boretto M, Clevers H (2020) Human-derived model systems in gynecological cancer research. Trends Cancer 6(12):1031–1043. https://doi.org/10.1016/j.trecan.2020.07.007
Lopez C, Tucker S, Salameh T, Tucker C (2018) An unsupervised machine learning method for discovering patient clusters based on genetic signatures. J Biomed Inform 85:30–39. https://doi.org/10.1016/j.jbi.2018.07.004
Luchini C, Pea A, Scarpa A (2021) Artificial intelligence in oncology: current applications and future perspectives. Br J Cancer 126(1):4–9. https://doi.org/10.1038/s41416-021-01633-1
Luo Y-H, Xi IL, Wang R, Abdallah HO, Wu J, Vance AZ et al (2020) Deep learning based on MR imaging for predicting outcome of uterine fibroid embolization. J Vasc Interv Radiol 31(6):1010-1017.e3. https://doi.org/10.1016/j.jvir.2019.11.032
Ma J-H, Huang Y, Liu L-Y, Feng Z (2021) An 8-gene DNA methylation signature predicts the recurrence risk of cervical cancer. J Int Med Res 49(5):030006052110184. https://doi.org/10.1177/03000605211018443
Mabwa D, Gajjar K, Furniss D, Schiemer R, Crane R, Fallaize C et al (2021) Mid-infrared spectral classification of endometrial cancer compared to benign controls in serum or plasma samples. Analyst 146(18):5631–5642. https://doi.org/10.1039/d1an00833a
Malla RR, Patnala K, Kumar DKG, Marni R (2021) Drug resistance in gynecologic cancers: emphasis on noncoding RNAs and drug efflux mechanisms. Overcoming drug resistance in gynecologic cancers. Elsevier, pp 155–168. https://doi.org/10.1016/b978-0-12-824299-5.00018-6
Mallik S, Mukhopadhyay A, Maulik U, Bandyopadhyay S (2013) Integrated analysis of gene expression and genome-wide DNA methylation for tumor prediction: an association rule mining-based approach. In: IEEE symposium on computational intelligence in bioinformatics and computational biology (CIBCB). IEEE. https://doi.org/10.1109/cibcb.2013.6595397
Manteghinejad A, Javanmard SH (2021) Challenges and opportunities of digital health in a post-covid19 world. J Res Med Sci 26
Martinic MK, Pieper D, Glatt A, Puljak L (2019) Definition of a systematic review used in overviews of systematic reviews, metaepidemiological studies and textbooks. BMC Med Res Methodol. https://doi.org/10.1186/s12874-019-0855-0
Medhin LB, Tekle LA, Achila OO, Said S (2020) Incidence of cervical, ovarian and uterine cancer in eritrea: data from the national health laboratory, 2011–2017. Sci Rep. https://doi.org/10.1038/s41598-020-66096-5
Melton BL (2017) Systematic review of medical informatics-supported medication decision making. Biomed Inform Insights. https://doi.org/10.1177/1178222617697975
Meng Z, Zhao Z, Li B, Su F, Guo L (2021) A cervical histopathology dataset for computer aided diagnosis of precancerous lesions. IEEE Trans Med Imaging 40(6):1531–1541. https://doi.org/10.1109/tmi.2021.3059699
Moher D, Shamseer L, Clarke M, Ghersi D, Liberati A, Petticrew M et al (2015) Preferred reporting items for systematic review and meta-analysis protocols (PRISMAp) 2015 statement. Syst Rev. https://doi.org/10.1186/2046-4053-4-1
Momenimovahed Z, Tiznobaik A, Taheri S, Salehiniya H (2019) Ovarian cancer in the world: epidemiology and risk factors. Int J Women’s Health 11:287–299. https://doi.org/10.2147/ijwh.s197604
Morganti S, Tarantino P, Ferraro E, D’Amico P, Viale G, Trapani D, et al (2019) Role of next-generation sequencing technologies in personalized medicine. P5 eHealth: an agenda for the health technologies of the future. Springer, pp 125–154
Mostafa S, Mondal D, Beck MA, Bidinosti CP, Henry CJ, Stavness I (2022) Leveraging guided backpropagation to select convolutional neural networks for plant classification. Front Artif Intell. https://doi.org/10.3389/frai.2022.871162
Nakagawa M, Nakaura T, Namimoto T, Iyama Y, Kidoh M, Hirata K et al (2019) A multiparametric MRI based machine learning to distinguish between uterine sarcoma and benign leiomyoma: comparison with 18f-FDG PET/CT. Clin Radiol 74(2):167.e1-167.e7. https://doi.org/10.1016/j.crad.2018.10.010
Nees LK, Heublein S, Steinmacher S, Juhasz-Böss I, Brucker S, Tempfer CB, Wallwiener M (2022) Endometrial hyperplasia as a risk factor of endometrial cancer. Arch Gynecol Obstet 306(2):407–421. https://doi.org/10.1007/s00404-021-06380-5
Nie X, Song L, Li X, Wang Y, Qu B (2021) Prognostic signature of ovarian cancer based on 14 tumor microenvironment-related genes. Medicine 100(28):e26574. https://doi.org/10.1097/md.0000000000026574
Oyelade J, Isewon I, Oladipupo F, Aromolaran O, Uwoghiren E, Ameh F et al (2016) Clustering algorithms: their application to gene expression data. Bioinformatics Biol Insights. https://doi.org/10.4137/bbi.s38316
Page MJ, McKenzie JE, Bossuyt PM, Boutron I, Hoffmann TC, Mulrow CD et al (2021) The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ. https://doi.org/10.1136/bmj.n71
Peng G, Dong H, Liang T, Li L, Liu J (2021) Diagnosis of cervical precancerous lesions based on multimodal feature changes. Comput Biol Med 130:104209. https://doi.org/10.1016/j.compbiomed.2021.104209
Petersen K, Vakkalanka S, Kuzniarz L (2015) Guidelines for conducting systematic mapping studies in software engineering: an update. Inf Softw Technol 64:1–18. https://doi.org/10.1016/j.infsof.2015.03.007
Praiss AM, Huang Y, Clair CMS, Tergas AI, Melamed A, Khoury-Collado F et al (2020) Using machine learning to create prognostic systems for endometrial cancer. Gynecol Oncol 159(3):744–750. https://doi.org/10.1016/j.ygyno.2020.09.047
Rahman R, Clark MD, Collins Z, Traore F, Dioukhane EM, Thiam H et al (2019) Cervical cancer screening decentralized policy adaptation: an African rural-context-specific systematic literature review. Glob Health Action 12(1):1587894. https://doi.org/10.1080/16549716.2019.1587894
Rajula HSR, Verlato G, Manchia M, Antonucci N, Fanos V (2020) Comparison of conventional statistical methods with machine learning in medicine: diagnosis, drug development, and treatment. Medicina 56(9):455. https://doi.org/10.3390/medicina56090455
Razzak MI, Imran M, Xu G (2019) Big data analytics for preventive medicine. Neural Comput Appl 32(9):4417–4451. https://doi.org/10.1007/s00521-019-04095-y
Reijnen C, Gogou E, Visser NCM, Engerud H, Ramjith J, van der Putten LJM et al (2020) Preoperative risk stratification in endometrial cancer (ENDORISK) by a Bayesian network model: a development and validation study. PLoS Med 17(5):e1003111. https://doi.org/10.1371/journal.pmed.1003111
Rodriguez JPM, Rodriguez R, Silva VWK, Kitamura FC, Corradi GCA, de Marchi ACB, Rieder R (2022) Artificial intelligence as a tool for diagnosis in digital pathology whole slide images: a systematic review. J Pathol Inform 13:100138. https://doi.org/10.1016/j.jpi.2022.100138
Sarana A, Subhashini R (2023) A systematic review of explainable artificial intelligence models and applications: recent developments and future trends. Decis Analyt J 7:100230. https://doi.org/10.1016/j.dajour.2023.100230
Sarica A, Cerasa A, Quattrone A (2017) Random forest algorithm for the classification of neuroimaging data in Alzheimer’s disease: a systematic review. Front Aging Neurosci. https://doi.org/10.3389/fnagi.2017.00329
Schiavo JH (2019) PROSPERO: an international register of systematic review protocols. Med Ref Serv Q 38(2):171–180. https://doi.org/10.1080/02763869.2019.1588072
Shao J, Zhang Z, Liu H, Song Y, Yan Z, Wang X, Hou Z (2020) DCE-MRI pharmacokinetic parameter maps for cervical carcinoma prediction. Comput Biol Med 118:103634. https://doi.org/10.1016/j.compbiomed.2020.103634
Shen W-C, Chen S-W, Wu K-C, Hsieh T-C, Liang J-A, Hung Y-C et al (2019) Prediction of local relapse and distant metastasis in patients with definitive chemoradiotherapy-treated cervical cancer by deep learning from [18f]-fluorodeoxyglucose positron emission tomography/computed tomography. Eur Radiol 29(12):6741–6749. https://doi.org/10.1007/s00330-019-06265-x
Shin SJ, You SC, Jeon H, Jung JW, An MH, Park RW, Roh J (2021) Style transfer strategy for developing a generalizable deep learning application in digital pathology. Comput Methods Programs Biomed 198:105815. https://doi.org/10.1016/j.cmpb.2020.105815
Shrestha P, Poudyal B, Yadollahi S, Wright DE, Gregory AV, Warner JD et al (2022) A systematic review on the use of artificial intelligence in gynecologic imaging–background, state of the art, and future directions. Gynecol Oncol 166(3):596–605. https://doi.org/10.1016/j.ygyno.2022.07.024
Sidey-Gibbons JAM, Sidey-Gibbons CJ (2019) Machine learning in medicine: a practical introduction. BMC Med Res Methodol. https://doi.org/10.1186/s12874-019-0681-4
Smeltzer MP, Lee Y-S, Faris Nicholas RM, Fehnel C, Akinbobola O, Meadows-Taylor M et al (2021) Trends in accuracy and comprehensiveness of pathology reports for resected NSCLC in a high mortality area of the united states. J Thorac Oncol 16(10):1663–1671. https://doi.org/10.1016/j.jtho.2021.06.027
Smrz SA, Calo C, Fisher JL, Salani R (2021) An ecological evaluation of the increasing incidence of endometrial cancer and the obesity epidemic. Am J Obstet Gynecol 224(5):506.e1-506.e8. https://doi.org/10.1016/j.ajog.2020.10.042
Song H-J, Yang E-S, Kim J-D, Park C-Y, Kyung M-S, Kim Y-S (2018) Best serum biomarker combination for ovarian cancer classification. BioMed Eng. https://doi.org/10.1186/s12938-018-0581-6
Stoler MH (2001) Interobserver reproducibility of cervical cytologic and histologic interpretations. JAMA 285(11):1500. https://doi.org/10.1001/jama.285.11.1500
Sun H, Zeng X, Xu T, Peng G, Ma Y (2020) Computer-aided diagnosis in histopathological images of the endometrium using a convolutional neural network and attention mechanisms. IEEE J Biomed Health Inform 24(6):1664–1676. https://doi.org/10.1109/jbhi.2019.2944977
Suzuki A, Aoki M, Miyagawa C, Murakami K, Takaya H, Kotani Y et al (2019) Differential diagnosis of uterine leiomyoma and uterine sarcoma using magnetic resonance images: a literature review. Healthcare 7(4):158. https://doi.org/10.3390/healthcare7040158
Tian Z, Yen A, Zhou Z, Shen C, Albuquerque K, Hrycushko B (2019) A machine-learning-based prediction model of fistula formation after interstitial brachytherapy for locally advanced gynecological malignancies. Brachytherapy 18(4):530–538. https://doi.org/10.1016/j.brachy.2019.04.004
Torheim T, Malinen E, Kvaal K, Lyng H, Indahl UG, Andersen EKF, Futsaether CM (2014) Classification of dynamic contrast enhanced MR images of cervical cancers using texture analysis and support vector machines. IEEE Trans Med Imaging 33(8):1648–1656. https://doi.org/10.1109/tmi.2014.2321024
Troisi J, Sarno L, Landolfi A, Scala G, Martinelli P, Venturella R et al (2018) Metabolomic signature of endometrial cancer. J Proteome Res 17(2):804–812. https://doi.org/10.1021/acs.jproteome.7b00503
Troisi J, Raffone A, Travaglino A, Belli G, Belli C, Anand S et al (2020) Development and validation of a serum metabolomic signature for endometrial cancer screening in postmenopausal women. JAMA Netw Open 3(9):e2018327. https://doi.org/10.1001/jamanetworkopen.2020.18327
Tsai M-H, Chen M-Y, Huang SG, Hung Y-C, Wang H-C (2014) A bio-inspired computing model for ovarian carcinoma classification and oncogene detection. Bioinformatics 31(7):1102–1110. https://doi.org/10.1093/bioinformatics/btu782
Urushibara A, Saida T, Mori K, Ishiguro T, Sakai M, Masuoka S et al (2021) Diagnosing uterine cervical cancer on a single t2-weighted image: comparison between deep learning versus radiologists. Eur J Radiol 135:109471. https://doi.org/10.1016/j.ejrad.2020.109471
van Haastrecht M, Sarhan I, Ozkan BY, Brinkhuis M, Spruit M (2021) SYMBALS: a systematic review methodology blending active learning and snowballing. Front Res Metr Anal. https://doi.org/10.3389/frma.2021.685591
van Panhuis WG, Paul P, Emerson C, Grefenstette J, Wilder R, Herbst AJ et al (2014) A systematic review of barriers to data sharing in public health. BMC Public Health. https://doi.org/10.1186/1471-2458-14-1144
Varughese J, Richman S (2010) Cancer care inequity for women in resource-poor countries. Rev Obstet Gynecol 3(3):122–132
Vázquez MA, Mariño IP, Blyuss O, Ryan A, Gentry-Maharaj A, Kalsi J et al (2018) A quantitative performance study of two automatic methods for the diagnosis of ovarian cancer. Biomed Signal Process Control 46:86–93. https://doi.org/10.1016/j.bspc.2018.07.001
Wadghiri M, Idri A, Idrissi TE, Hakkoum H (2022) Ensemble blood glucose prediction in diabetes mellitus: a review. Comput Biol Med 147:105674. https://doi.org/10.1016/j.compbiomed.2022.105674
Wardle J, Robb K, Vernon S, Waller J (2015) Screening for prevention and early diagnosis of cancer. Am Psychol 70(2):119–133. https://doi.org/10.1037/a0037357
Wasnik AP (2013) Multimodality imaging of ovarian cystic lesions: review with an imaging based algorithmic approach. World J Radiol 5(3):113. https://doi.org/10.4329/wjr.v5.i3.113
Wieringa R, Maiden N, Mead N, Rolland C (2005) Requirements engineering paper classification and evaluation criteria: a proposal and a discussion. Requirements Eng 11(1):102–107. https://doi.org/10.1007/s00766-005-0021-6
Wilailak S, Kengsakul M, Kehoe S (2021) Worldwide initiatives to eliminate cervical cancer. Int J Gynecol Obstet 155(S1):102–106. https://doi.org/10.1002/ijgo.13879
Wu W, Zhou H (2017) Data-driven diagnosis of cervical cancer with support vector machine-based approaches. IEEE Access 5:25189–25195. https://doi.org/10.1109/access.2017.2763984
Wu Q, Wang S, Zhang S, Wang M, Ding Y, Fang J et al (2020) Development of a deep learning model to identify lymph node metastasis on magnetic resonance imaging in patients with cervical cancer. JAMA Netw Open 3(7):e2011625. https://doi.org/10.1001/jamanetworkopen.2020.11625
Xue Y, Zhou Q, Ye J, Long LR, Antani S, Cornwell C, et al (2019a) Synthetic augmentation and feature-based filtering for improved cervical histopathology image classification. arXiv:1907.10655
Xue Y, Zhou Q, Ye J, Long LR, Antani S, Cornwell C, et al (2019b) Synthetic augmentation and feature-based filtering for improved cervical histopathology image classification. arXiv:1907.10655
Xue D, Zhou X, Li C, Yao Y, Rahaman MM, Zhang J et al (2020) An application of transfer learning and ensemble learning techniques for cervical histopathology image classification. IEEE Access 8:104603–104618. https://doi.org/10.1109/access.2020.2999816
Xue P, Ng MTA, Qiao Y (2020) The challenges of colposcopy for cervical cancer screening in LMICs and solutions by artificial intelligence. BMC Med. https://doi.org/10.1186/s12916-020-01613-x
Xue P, Tang C, Li Q, Li Y, Shen Y, Zhao Y et al (2020) Development and validation of an artificial intelligence system for grading colposcopic impressions and guiding biopsies. BMC Med. https://doi.org/10.1186/s12916-020-01860-y
Xue P, Wang J, Qin D, Yan H, Qu Y, Seery S, et al (2022) Deep learning in image-based breast and cervical cancer detection: a systematic review and meta-analysis. npj Digit Med. https://doi.org/10.1038/s41746-022-00559-z
Xue P, Xu H-M, Tang H-P, Wu W-Q, Seery S, Han X et al (2023) Assessing artificial intelligence enabled liquid-based cytology for triaging HPV positive women a population based crosssectional study. Acta Obstet Gynecol Scand 102(8):1026–1033. https://doi.org/10.1111/aogs.14611
Yang X, Stamp M (2021) Computer-aided diagnosis of low grade endometrial stromal sarcoma (LGESS). Comput Biol Med 138:104874. https://doi.org/10.1016/j.compbiomed.2021.104874
Yang GR, Wang X-J (2020) Artificial neural networks for neuroscientists: a primer. Neuron 107(6):1048–1070. https://doi.org/10.1016/j.neuron.2020.09.005
Yin F-F, Zhao L-J, Ji X-Y, Duan N, Wang Y-K, Zhou J-Y et al (2019) Intra-tumor heterogeneity for endometrial cancer and its clinical significance. Chin Med J 132(13):1550–1562. https://doi.org/10.1097/cm9.0000000000000286
Yu K-H, Hu V, Wang F, Matulonis UA, Mutter GL, Golden JA, Kohane IS (2020) Deciphering serous ovarian carcinoma histopathology and platinum response by convolutional neural networks. BMC Med. https://doi.org/10.1186/s12916-020-01684-w
Yu Y, Ma J, Zhao W, Li Z, Ding S (2021) MSCI: a multistate dataset for colposcopy image classification of cervical cancer screening. Int J Med Informatics 146:104352. https://doi.org/10.1016/j.ijmedinf.2020.104352
Yuan C, Yao Y, Cheng B, Cheng Y, Li Y, Li Y et al (2020) The application of deep learning based diagnostic system to cervical squamous intraepithelial lesions recognition in colposcopy images. Sci Rep. https://doi.org/10.1038/s41598-020-68252-3
Zeng H, Chen L, Zhang M, Luo Y, Ma X (2021) Integration of histopathological images and multi-dimensional omics analyses predicts molecular features and prognosis in high-grade serous ovarian cancer. Gynecol Oncol 163(1):171–180. https://doi.org/10.1016/j.ygyno.2021.07.015
Zhang Z, Han Y (2020) Detection of ovarian tumors in obstetric ultrasound imaging using logistic regression classifier with an advanced machine learning approach. IEEE Access 8:44999–45008. https://doi.org/10.1109/access.2020.2977962
Zhang S, Gong T-T, Liu F-H, Jiang Y-T, Sun H, Ma X-X et al (2019) Global, regional, and national burden of endometrial cancer, 1990–2017: results from the global burden of disease study, 2017. Front Oncol. https://doi.org/10.3389/fonc.2019.01440
Zhang H, Chen C, Gao R, Yan Z, Zhu Z, Yang B et al (2021) Rapid identification of cervical adenocarcinoma and cervical squamous cell carcinoma tissue based on Raman spectroscopy combined with multiple machine learning algorithms. Photodiagn Photodyn Ther 33:102104. https://doi.org/10.1016/j.pdpdt.2020.102104
Zhang S, Chen C, Chen C, Chen F, Li M, Yang B et al (2021) Research on application of classification model based on stack generalization in staging of cervical tissue pathological images. IEEE Access 9:48980–48991. https://doi.org/10.1109/access.2021.3064040
Zhang Y, Wang Z, Zhang J, Wang C, Wang Y, Chen H et al (2021) Deep learning model for classifying endometrial lesions. J Transl Med. https://doi.org/10.1186/s12967-020-02660-x
Zhao J, Hu Y, Zhao Y, Chen D, Fang T, Ding M (2021) Risk factors of endometrial cancer in patients with endometrial hyperplasia: implication for clinical treatments. BMC Women’s Health. https://doi.org/10.1186/s12905-021-01452-9
Zhen X, Chen J, Zhong Z, Hrycushko B, Zhou L, Jiang S et al (2017) Deep convolutional neural network with transfer learning for rectum toxicity prediction in cervical cancer radiotherapy: a feasibility study. Phys Med Biol 62(21):8246–8263. https://doi.org/10.1088/1361-6560/aa8d09
Author information
Authors and Affiliations
Contributions
Both authors contributed to the creation of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors report no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Idlahcen, F., Idri, A. & Goceri, E. Exploring data mining and machine learning in gynecologic oncology. Artif Intell Rev 57, 20 (2024). https://doi.org/10.1007/s10462-023-10666-2
Published:
DOI: https://doi.org/10.1007/s10462-023-10666-2