Abstract
In this work, we study and analyze the aggregate death counts of COVID-19 reported by the United States Centers for Disease Control and Prevention (CDC) for the fifty states in the United States. To do this, we derive a stochastic model describing the cumulative number of deaths reported daily by CDC from the first time Covid-19 death is recorded to June 20, 2021 in the United States, and provide a forecast for the death cases. The stochastic model derived in this work performs better than existing deterministic logistic models because it is able to capture irregularities in the sample path of the aggregate death counts. The probability distribution of the aggregate death counts is derived, analyzed, and used to estimate the count’s per capita initial growth rate, carrying capacity, and the expected value for each given day as at the time this research is conducted. Using this distribution, we estimate the expected first passage time when the aggregate death count is slowing down. Our result shows that the expected aggregate death count is slowing down in all states as at the time this analysis is conducted (June 2021). A formula for predicting the end of Covid-19 deaths is derived. The daily expected death count for each states is plotted as a function of time. The probability density function for the current day, together with the forecast and its confidence interval for the next four days, and the root mean square error for our simulation results are estimated.
Similar content being viewed by others
1 Introduction
Several mathematical models (Wu et al. 2020; Stutt et al. 2020; Linka et al. 2020; Okuonghae and Omame 2020; Ndairou et al. 2020; Ladde et al. 2020; Otunuga 2020; Mummert and Otunuga 2019; Otunuga 2018; Santosh 2020) have been developed to study the transmission of the COVID-19 virus caused by the virus species ”severe acute respiratory syndrome-related corona virus”, named SARS-CoV-2. The airborne transmission occurs by inhaling droplets loaded with SARS-CoV-2 particles that are expelled by infectious people. According to Wu et al. (2020), the ”severe acute respiratory syndrome coronavirus” (SARS-CoV) and the ”Middle East respiratory syndrome coronavirus” (MERS-CoV) are two other novel coronaviruses that emerged as major global health threats since 2002. Several public health interventions have been put in place to eradicate or reduce the spread of the disease. According to CDC, the first U.S. laboratory-confirmed caseFootnote 1 of COVID-19 in the U.S. was recorded on January 20, 2020 from the samples taken 2 days earlier in Washington state. The first COVID-19 death in the United States was first reported in the same state by CDC on February 29, 2020. As of June 24, 2021, the total number of Covid-19 cases in the United States was reported by CDC to be 33, 437, 643, resulting in about 601, 221 deathsFootnote 2. On December 11 and December 18, 2020, the United States Food and Drug Administration (FDA)Footnote 3 issued an Emergency Use Authorization (EUA) for the Pfizer-BioNTech and the Moderna COVID-19 vaccine, respectively, in the United States. An EUA for the third vaccine, the Johnson and Johnson (J &J) vaccine, was first issued in the United States on February 27, 2021Footnote 4. A pause on the usage of the J &J vaccine was recommended by CDC and the FDA on April 13, 2021 due to the serious blood clots (a condition called thrombosis) in six women between the ages of 18 and 49 years with thrombocytopenia syndrome (TTS)Footnote 5 following the usage of the vaccine. As of June 7, 2021, about \(51.5\%\) of the total population of the United States have received at least one dose of the vaccination, with \(41.9\%\) fully vaccinated. The study done in this work is to check the effects of these interventions. That is, with these vaccines, we check if the number of deaths resulting from the Covid-19 disease slows down as time proceeds and as the number of those who are vaccinated increases.
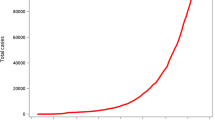
The trajectory of the aggregate death counts in most states in the United States follows the same dynamics. At first, it follows a somewhat exponential trajectory, with its growth slowing down at some point and speeding up at other points. With public health interventions like vaccination mitigating the growth of the virus, this pattern is expected to continue until a certain steady-state is reached. This dynamic follows roughly the well known Verhulst logistic equation and its generalization (Beddington and May 1977; Li and Wang 2010; Prajneshu 1980; Pelinovsky et al. 2020; Pella and Tomlinson 1969; Wang et al. 2020). The model was first derived by Verhulst (1838) to study population growth. Some other methods (Baud et al. 2020; Bhapkar et al. 2020; Satpathy et al. 2021; Kaciroti et al. 2021) have been developed to estimate mortality following the Covid-19 infection. In this work, we consider the logistic model
where \(t_{0}\ge 0\), \(N_{0}>0\), N(t) denotes the total number of Covid-19 death counts at time t, \(\mathcal {K}\) is the maximum number of Covid-19 deaths, and \(\bar{\varvec{\mu }}\) is the per capita initial growth rate, to interpret the aggregate number of COVID-19 death trajectories in the United States. We see from (1.1) that \(\frac{dN}{dt}>0\) on \(\left( 0,\mathcal {K}\right)\) and \(\frac{d^{2}N}{dt^{2}}=\frac{\bar{\varvec{\mu }}^{2}}{\mathcal {K}^{2}}\left( \mathcal {K}-N\right) \left( \mathcal {K}-2N\right) N\). From this, we have \(\frac{d^{2}N}{dt^{2}}>0\) on the interval \((0,\mathcal {K}/2)\), and \(\frac{d^{2}N}{dt^{2}}<0\) on \((\mathcal {K}/2,\mathcal {K})\). That is, the trajectory of the aggregate number speeds up in the interval \((0,\mathcal {K}/2)\) and slows down in the interval \((\mathcal {K}/2,\mathcal {K})\). Also, since N(t) represents the aggregate number of deaths at a given time t, it follows that the daily number of deaths is at the maximum at the time when \(N(t)=\mathcal {K}/2\).
From this analysis, if the current day aggregate death counts in time series is more than \(\mathcal {K}/2\), then we know that its growth is slowing down and the virus spread has been controlled. Otherwise, the virus is spreading, with speeding growth. The problem with using this model to analyze the aggregate number of cases is that it fails to account for the fluctuations or perturbations in the data resulting from fluctuations in the rates of infection or death. These noises/fluctuations can be caused by many factors like the rates at which Covid-19 testing is done, vaccination rates, mask use per capital, social behavior, public health intervention (Linka et al. 2020), and so on. In addition, CDCFootnote 6 reported on their websites that counting exact aggregate confirmed and probable COVID-19 cases and deaths is not possible due to delays in reporting from different voluntary jurisdictions. The number of death cases reported on CDC’s website might not be complete because it takes several weeks for death records to be processed, coded, submitted, and tabulated on the National Center for Health Statistics (NCHS). As a result, these cause the counts to fluctuate substantially, with a possibility of a negative number of probable cases reported on a given day if more probable cases were disproven than were initially reported on that day.
Several authors (Lv et al. 2019; Li and Wang 2010; Lungu and Øksendal 1997; Yang et al. 2019; Gardiner 1985) have worked on model (1.1) and its extension to a stochastic model. In this work, we derive a stochastic model governing the aggregate number of Covid-19 death counts in the United States by extending model (1.1) to a stochastic case. The proposed model is better in the sense that it captures fluctuations in the aggregate counts better than the widely used logistic model (1.1) with lesser root mean square error. Our aim in this study is to determine whether the virus infection and death counts resulting from the infection are still growing sharply or slowing down. Upon analyzing the COVID-19 data collected from CDC, we see that the path of the aggregate number of Covid-19 death counts over time follows a logistic model with irregular trajectories. We assume this fluctuation is caused by many factors such as listed above, causing the per capita growth rate to fluctuate over time. We account for the fluctuations in this rate by extending the deterministic logistic model (1.1) to a stochastic model. This is done by assuming the parameter \(\bar{\varvec{\mu }}\) is not constant over time, but fluctuates about a particular mean value. In order to estimate the epidemiological parameters \(\mathcal {K}\) (the carrying capacity of Covid-19 death counts), \(\bar{\varvec{\mu }}\) (the per capita initial growth rate), and the death rate noise intensity, we derive the transition probability density function for the aggregate number of death counts and apply a Maximum Likelihood Estimate (MLE) scheme. The distribution is also used to calculate the expected aggregate count at a particular period in time. Using the parameter estimates for the fitted data, we forecast the cumulative number of Covid-19 deaths and provide a \(95\%\) confidence interval for the forecast.
The organization of the work done is as follows: In Sect. 2, we derive a stochastic model describing the cumulative number of deaths by assuming \(\bar{\varvec{\mu }}\) is not constant, but changes with time and fluctuates around a mean value. We show that the model is well defined, with a unique closed-form solution. In Sect. 3, the transition probability density function for the aggregate number of death counts is derived. Using the MLE scheme, we estimate the epidemiological parameters in the model. Since the aggregate count N(t) is random, we calculate the expected number of Covid-19 aggregate death counts for each states in the United States. We show that with probability one the aggregate count remains in the interval \((0,\mathcal {K})\) if it starts from there. In Sect. 4, the expected first hitting time when the aggregate count N(t) reaches \(\mathcal {K}-\epsilon\) is calculated for some small positive constant \(\epsilon >0\). We also estimate the expected first passage time when the aggregate death counts started slowing down. Numerical simulations, forecast, and analysis of the aggregate death counts for the fifty states in the United States are carried out in Sect. 5. The summary of the work done is given in Sect. 6.
2 Methodology
2.1 Data Sources
The Covid-19 aggregate death counts in the United States is collected from the United States Centers for Disease Control and Prevention (CDC) website and provided by the CDC Case Task ForceFootnote 7. The data was collected for the period ranging from January 22, 2020 to June 24, 2021, and it includes the date of counts, state/jurisdiction, total/aggregate number of death cases (including total confirmed and probable deaths), number of new and new probable deaths with the date and time the records were created. The definition of each of these can be found on the CDC’s website\(^7\).
2.2 Modeling the Covid-19 Aggregate Death Counts
In this section, we describe the dynamics of the aggregate number of deaths in the United States by extending the well-known deterministic logistic model (1.1) to a stochastic differential equation. Analysis of the data (see Figs. 1, 2) shows that its growth rate fluctuates with time. Some works (Lagarto and Braumann 2014; Mazzuco et al. 2018; Zocchetti and Consonni 1994) have been done in analyzing the distribution of mortality rate. The dynamics of male and female crude death rates of the Portuguese population over the period 1940–2009 were modeled in the work of Lagarto and Braumann (2014) using a bi-dimensional stochastic Gompertz model with correlated Wiener processes. Zocchetti and Consonni (1994) showed in their work that when the number of deaths is sufficiently elevated, the Gauss distribution (also referred to as the normal distribution) can be used as a good approximation distribution for the variability in the mortality rate. In their work, Mazzuco et al. (2018) derived a new model for mortality rate based on the mixture of a half-normal distribution with a generalization of the skew-normal distribution. The Wiener process (Khasminskii 2012; Kloeden and Platen 1995; Mao 2007; Øksendal 2003), often called Brownian motion, is a real-valued continuous-time stochastic process \(\{W(t): t\ge 0\}\) defined on a probability space \((\Omega , \mathcal {F},\mathbb {P})\) with stationary independent Gaussian increments such that \(W(0)=0\) with probability one, \(W(t+\Delta t)-W(t)\) is normally distributed with mean 0 and variance \(\Delta t\), and \(W(t)-W(s)\) is independent of the past random variable W(u), \(0\le u\le s\). In addition, the random variables \(\{W(t_{j})-W(s_{j}),\ j=1,2,\cdots ,n\}\) are jointly independent, for \(0\le s_{1}<t_{1}\le s_{2}<t_{2}\cdots \le s_{n}<t_{n}<\infty\). The independence and stationarity of the increment, together with the continuous (almost everywhere) sample path of the process W(t) lead to its great tractability, making it one of the most important stochastic process in continuous time used in biological systems to model perturbed epidemiological parameters. Following a similar assumption made in the work of Prajneshu (1980), Yang et al. (2019), and Otunuga (2021b), where a logistic stochastic population model with the population’s transition probability density function were derived to study the distribution of a population subjected to a continuous spectrum of disturbances, with fluctuations in the intrinsic growth rate, we assume the dynamics of \(\bar{\varvec{\mu }}\) shouldn’t be constant over time, but instead be driven by fluctuations that can be modeled to follow a process of the form
where W(t) is a standard Wiener process, \(\mu\) is the average death counts per capita initial growth rate, \(\sigma\) is the noise intensity, and \(\circ\) is the Stratonovich integral symbol (Arnold 1974). We use the Stratonovich calculus instead of the Itô calculus to describe this dynamic simply because it obeys the traditional rule of chain rule and allows white noise to be treated as a regular derivative of a Brownian or Wiener process (West et al. 1979; Wong and Zakai 1965). For more reading on the Itô and Stratonovich calculus, we direct readers to the work of Kloeden and Platen (1995) and Øksendal (2003). Substituting into (1.1), the proposed stochastic differential equation (SDE) for the aggregate number of deaths is given by
where \(N_{0}>0\), \(\mathcal {K}\) is the carrying death capacity, \(\mu\) and \(\sigma\) are as described in (2.1). We note here that the stochasticity added in model (2.2) leads to a non-monotonic cumulative death sample path, a property that makes it able to capture irregularities in the sample path better than the deterministic counterpart, with a smaller root mean square error. The interpretation of the stochastic differential equation as a Stratonovich differential equation follows the work of Otunuga (2019, 2021a). We shall later compare model (2.2) with its deterministic equivalent (1.1) and show that model (2.2) performs better in capturing the trajectory of (and the noise in) the aggregate death counts. We convert (2.2) into a Itô stochastic differential equation as
In order to show that model (2.3) is biologically feasible, we show in Theorem 2 (using Corollary 3.1 of Khasminskii (2012)) that the solution N(t) of (2.3) exists, and it remains in \((0,\mathcal {K})\) with probability one whenever it starts from there. A statement of the corollary, together with definition of some terminologies in the corollary are given below.
Definition 1
L-operator.
Given a one-dimensional stochastic differential equation
we define the L-operator (Mao 2007) associated with (2.4) as
If L acts on a nonnegative function V(t, x) which is continuously differentiable with respect to t and twice continuously differentiable with respect to x, then
The usefulness of the expression for LV(t, x) is seen in the Itô Lemma (a Lemma which gives the formula for the stochastic analogue of the chain rule or change of variable rule in calculus), which simply states that the stochastic differential of V(t, x) is given by
We see here that LV(t, x) is the drift part of the differential dV(t, x).
Theorem 1
(From Corollary 3.1 of Khasminskii (2012))
Let \(D_{n}\) be an increasing sequence of open sets whose closure are contained in an open set D such that \(\bigcup D_{n} =D\). Suppose that the drift and diffusion coefficients f(t, x) and g(t, x), respectively, in (2.4) satisfy the Lipschitz and linear growth conditions in \((0,\infty )\times D_{n}\) and there exists a function V(t, x), twice continuously differentiable in x and continuously differentiable in t in the domain \((0,\infty )\times D\), which satisfies
for some positive constant c. Then for every random variable \(x(t_{0})\) independent of \(W(t)-W(t_{0})\), there exists a solution x(t) of (2.4) which is an almost surely continuous stochastic process and is unique up to equivalenceFootnote 8 provided that \(\mathbb {P}(x(t_{0})\in D)=1\). Moreover the solution satisfies the relation
Since the drift and diffusion coefficients of (2.3) are non-linear, the classical existence and uniqueness theorem of SDE (Kloeden and Platen 1995; Khasminskii 2012; Mao 2007; Øksendal 2003) does not apply. We use Theorem 1 to prove the existence and uniqueness of the solution of (2.3) in the interval \((0,\mathcal {K})\) in Theorem 2.
Theorem 2
Let the stochastic differential equation (2.3) be given for any \(t_{0}\ge 0\) and initial value \(N_{0}\in (0,\mathcal {K})\) independent of \(W(t)-W(t_{0})\). Then there exists a unique global positive solution \(N:[t_{0},\infty )\rightarrow \Re ^{+}\) such that with probability one \(N(t)\in (0,\mathcal {K})\). That is,
Proof
Define the sequence \(\{D_{r}\}\) by
Clearly, \(\{D_{r}\}\) is an increasing sequence of open sets whose closures are contained in \((0,\mathcal {K})\). The drift and diffusion coefficients \(b(N)=\frac{\mu }{\mathcal {K}}\left( \mathcal {K}-N\right) N+\frac{\sigma ^{2}}{2\mathcal {K}^{2}} \left( \mathcal {K}-N\right) \left( \mathcal {K}-2N\right) N\) and \(g(N)=\frac{\sigma }{\mathcal {K}} \left( \mathcal {K}-N\right) N\), respectively, of (2.3) satisfy the Lipschitz and linear growth conditions locally in \(D_{r}\). Define a function \(V: \left( 0,\mathcal {K}\right) \rightarrow \Re ^{+}\) by
Applying the L-operator ((2.5)) associated with (2.3) on V, we have
where \(\mathcal {C}=\mu +3\sigma ^{2}/2\). For any \(x\in D\backslash D_{r}=\left( 0,\frac{1}{r}\right] \bigcup \left[ \mathcal {K}-\frac{1}{r},\mathcal {K}\right)\), we have
The result follows from Theorem 1. \(\square\)
We give the exact solution of (2.3) in the following theorem. This solution will be used in the simulation process to plot the sample path of the death counts.
Theorem 3
For any given initial value \(N_{0}\in (0,\mathcal {K})\), the exact solution of the SDE (2.3) is obtained as
where
Proof
As shown in Theorem (2), there is a unique global positive solution \(N(t)\in (0,\mathcal {K})\) for all \(t\ge t_{0}\) with probability one. Define
It follows from (2.3) and (2.6) that
with solution
The result follows by substituting (2.10) into (2.8) and solving for N(t). \(\square\)
Remark 1
Theorem 2 shows that the aggregate number of deaths cannot grow past a particular value, \(\mathcal {K}\), with probability one if its starting point is in \((0,\mathcal {K})\). If \(0< N_{0} <\mathcal {K}\), then it follows from Theorems 2 and 3 that the feasible epidemiological region of interest for the solution N(t) is the set
The following theorem shows that if \(\mu >\sigma ^{2}/2\) and the process N(t) starts from \((0,\mathcal {K})\), then it converges almost surely to a random variable \(N_{\infty }\) with finite expectation.
Theorem 4
For any given initial condition \(N_{0}\in \left( 0,\mathcal {K}\right)\), the process N(t) is a submartingale if \(\mu >\sigma ^{2}/2\). That is,
Furthermore, there exists a random variable \(N_{\infty }\) such that \(\mathbb {E}\left[ \left( N_{\infty }\right) \right] <\infty\) and
Proof
It follows from (2.2) that for \(s\le t\), we have
If \(\mu >\sigma ^{2}/2\), we obtain
Clearly, we see from (2.7) that \(0<N(t)<\mathcal {K}\) for all \(t\ge 0\). Hence, \(\sup \limits _{t\ge 0}\mathbb {E}\left[ \left( N(t)^{+}\right) \right] <\infty\). By the Martingale Convergence Theorem, we have \(\mathbb {E}\left( N_{\infty }\right) <\infty\) and equation (2.12) is satisfied. \(\square\)
Remark 2
The Martingale Convergence Theorem is a stochastic analogue of the Monotone Convergence Theorem. Here, we also see that a submartingale property is a stochastic analogue of a non-decreasing sequence. Since aggregate death count is expected to be increasing with time and the process N(t) is stochastic, we need a condition that will guarantee a stochastic analogue of an increasing function. Theorem 4 gives such condition. It shows that the expected aggregate death counts is an increasing function provided \(N_{0}\in (0,\mathcal {K})\) and \(\mu >\sigma ^{2}/2\). It has been shown in several works (Mendez et al. 2012; Otunuga 2018, 2020) that the presence of environmental perturbations can affect the dynamic nature of biological systems if the noise intensity grows beyond a certain value. The latter condition shows that the noise intensity, \(\sigma\), of the environmental perturbation must not be allowed to grow beyond a certain function of the average death count’s growth rate \(\mu\) if the submartingale property is to be maintained. Following Theorem 4, we assume for the rest of this work that the noise intensity \(\sigma <\sqrt{2\mu }\).
We discuss and analyze the distribution of the random aggregate number of death counts process N(t) by first deriving its transition probability density function. The distribution will be used to estimate the epidemiological parameters \(\mathcal {K}\), \(\mu\), and \(\sigma\), and to calculate the expected aggregate number of death counts at a particular time t in the United States. These estimates will later be used in simulating and forecasting the total death counts.
3 Probability Distribution of the Aggregate Number of Deaths Following (2.3)
Let \(p_{N}(n|t,N_{0})\) represents the transition probability density function (PDF) for the aggregate death counts N(t) given t and \(N_{0}\). Following the results in (2.8) and (2.10), the transition probability density function \(p_{N}(n|t,N_{0})\) is obtained as
The purpose of deriving this PDF is to be able to estimate the epidemiological parameters in model (2.3) using the MLE scheme.
3.1 Parameter Estimates
Let T be a number corresponding to the current date the Covid-19 aggregate data is collected, and \(t_{0}<t_{1}<\cdots <t_{m}=T\) be a partition P of the interval \([t_{0},T]\). Denote \(N(t_{j})\) by \(N_{j}\) and let \(N_{0}, N_{1}, N_{2},\cdots ,N_{m}\) be samples satisfying (2.3) at a given time. Let \(\Delta t_{j}=t_{j}-t_{j-1}\), \(j=1,2,\cdots ,m\). The likelihood and log-likelihood functions \(L\left( \Theta |N\right)\) and \(\mathcal {L}\left( \Theta |N\right)\), respectively, of the samples are obtained using (3.1), the transformation (2.8), and the distribution of u in (2.10), as
and
where \(\Theta =\{\mathcal {K}, \mu , \sigma \}\) represents the parameter set to be estimated. The maximum likelihood estimates \(\hat{\mathcal {K}}\), \(\hat{\mu }\), \(\hat{\sigma }^{2}\) of \({\mathcal {K}}\), \({\mu }\), \({\sigma ^{2}}\) are estimated from (3.2) as
where \(\hat{\mathcal {K}}\) satisfies
Remark 3
The initial point \(N_{0}\) can also be estimated for better simulation result. In this case, the estimates \(\hat{\mathcal {K}}\), \(\hat{\mu }\), \(\hat{\sigma }^{2}\), \(\hat{N}_{0}\) of \({\mathcal {K}}\), \({\mu }\), \({\sigma ^{2}}\), and \(N_{0}\) reduce to
where \(\hat{\mathcal {K}}\) satisfies
3.2 Expected and Simulated Number of Deaths
Since the aggregate death counts N(t) is a random process, it is important to calculate the expected number of total deaths at each given time. Given the initial value \(N_{0}\), the expected aggregate number of deaths, denoted \(\mathbb {E}\left[ N(t)|N_{0}\right]\), at time t is calculated from (3.1) as
We show in the next theorem that for each time t, the expected death counts falls in \((0,\mathcal {K})\) if the death counts starts there.
Theorem 5
If \(N_{0}\in (0,\mathcal {K})\), then \(0<\mathbb {E}[N(t)|N_{0}]< \mathcal {K}\).
Proof
If \(N_{0}\in (0,\mathcal {K})\), then it follows from (3.7) that
\(\square\)
Theorem 5 shows that the expected aggregate number of deaths will always be in the feasible region \(\mathcal {T}\) if the initial point \(N_{0}\) starts from there. The following theorem gives the total deaths expected on the long run and shows condition under which this number converges to the point \(N=\mathcal {K}\) using Theorem 5.3 of Khasminskii (2012).
Theorem 6
If \(\mu >\sigma ^{2}/2\) and \(N_{0}\in (0,\mathcal {K})\), then the total deaths expected on the long run is \(\mathcal {K}\).
Proof
Consider the random process \(z(t)=\mathcal {K}-N(t)\). It follows from (3.1) that the probability density function \(p_{z}(z|t,z_{0})\) of the random variable z given the initial point \(z_{0}\) is obtained as
For any \(\delta >0\),
and
We deduce from the Squeeze Theorem that
Hence, the result follows from Khasminskii (2012). \(\square\)
4 Predicting the End of Covid-19 Death
Since N(t) denotes the aggregate death counts at a particular time t, it follows that dN/dt will describe the daily death counts. Following Theorems 2 and 6, we know that the daily death counts converges to zero as N(t) converges asymptotically to \(\mathcal {K}\). So, in order to calculate an approximate time that people will stop dying of Covid-19, we need to calculate, in an \(\epsilon >0\) neighborhood of \(\mathcal {K}\), a time when the aggregate number \(\mathcal {K}-\epsilon\) is reached. For some positive small constant \(\epsilon\), define the open interval
Following Theorem 6, we plan to calculate the first-hitting-time \(\tau _{\epsilon }\) until the process N(t) enters \(\mathcal {A}_{\epsilon }\).
Definition 2
We define the first passage time \(\tau _{\epsilon }\) as
Let \(g(t)=\mathbb {P}\left( \tau _{\epsilon }\le t\right)\) and \(f_{\tau _{\epsilon }}(t)=dg(t)/dt\) be the First Passage Time Density (FPTD), the probability that the aggregate number of death counts N(t) has first reached a point \(\mathcal {K}{-\epsilon }\) at exactly time t. We derive \(f_{\tau _{\epsilon }}(t)\) and the expected first hitting time in the theorem below for small \(\epsilon\).
Theorem 7
If \(N_{0}\in \left( 0, \mathcal {K}\right)\), then the probability \(f_{\tau _{\epsilon }}(t)\) that the aggregate number of death counts N(t) has first reached a point \(\mathcal {K}-\epsilon\) at exactly time t is obtained as
Furthermore, for t0=0, the expected first hitting time \(\mathbb {E}\left( \tau _{\epsilon }\right)\) is obtained as
Proof
From this, and the Fundamental Theorem of Calculus, we have
and for t0=0,
\(\square\)
Equation (4.3) and Theorem 7 can be used to calculate an approximate expected time when people in the United States will stop dying of the Covid-19 by making \(\epsilon\) to be as small as possible.
Remark 4
For the solution \(N_{\sigma =0}(t)=\frac{\mathcal {K}}{1+\frac{\mathcal {K}-N_{0}}{N_{0}}e^{-\mu t}}\) of (1.1), the time \(T_{\epsilon }=\frac{1}{\mu }\ln \left( \frac{\left( \mathcal {K}-\epsilon \right) \left( \mathcal {K}-N_{0}\right) }{\epsilon N_{0} }\right)\) at which \(N_{\sigma =0}(t)=\mathcal {K}-\epsilon\) satisfies \(T_{\epsilon }=\mathbb {E}\left( \tau _{\epsilon }\right)\).
As shown in Sect. 1 with respect to model (1.1), the aggregate death count’s growth speeds up in the interval \((0,\mathcal {K})\) and slows down in the interval \((\mathcal {K}/2,\mathcal {K})\). This shows that the maximum number of daily deaths can be calculated as \(\max \left( \frac{dN}{dt}\right) =\frac{\varvec{\mu }}{4}\mathcal {K}\), occurring at time \(T_{\mathcal {K}/2}=\frac{1}{\varvec{\mu }}\ln \left( \frac{\mathcal {K}-N_{0}}{N_{0}}\right) {=\mathbb {E}\left( \tau _{\mathcal {K}/2}\right) }\). In the next theorem, we calculate the expected first time when the aggregate death counts reaches the size \(\mathcal {K}/2\) for the stochastic case.
Corollary 8
If \(N_{0}\in \left( 0,\mathcal {K}\right)\), we have \(\lim \limits _{\epsilon \rightarrow 0^{+}}\mathbb {E}\left( \tau _{\epsilon }\right) =+\infty\). That is, the process N(t) never reaches the point \(\mathcal {K}\). Also, \(\mathbb {E}\left( \tau _{\mathcal {K}/2}\right) =\frac{1}{\mu }\ln \left( \frac{\left( \mathcal {K}-N_{0}\right) }{ N_{0} }\right) .\)
Proof
The proof follows from (4.3) and Theorem 7. \(\square\)
The expected first time passage \(\mathbb {E}\left( \tau _{\mathcal {K}/2}\right)\) is analogous to the deterministic time \(T_{\mathcal {K}/2}\) when the process N(t) starts slowing down. That is, the time where the maximum number of daily deaths occurs.
As discussed in Sect. 2, we showed, with respect to the deterministic model (1.1), the current day’s aggregate death count’s growth is slowing down starting from the moment when the current day’s aggregate count is more than \(\mathcal {K}/2\). Otherwise, the virus is spreading, with speeding growth. Numerical results in Figs. 7 and 8 also show, for the stochastic case, that the expected aggregate count slows down starting from the time \(\mathbb {E}\left( \tau _{\mathcal {K}/2}\right)\).
5 Numerical Simulation and Forecast for the Aggregate Death Counts in the United States
As discussed earlier, W(t) is a Wiener process that depends continuously on \(t\in [0,T]\) with independent increment property such that \(W(t_{0}=0)=0\), \(W(t)-W(s)\sim \sqrt{t-s}\ N(0,1)\) for \(0\le s<t\le T\), where N(0, 1) is the standard normal distribution. Let \(\Delta W_{j}=W_{j}-W_{j-1}\), \(j=1,2,\cdots ,m\), where \(W_{j+1}=W(t_{j+1})\), \(t_{j}=t_{0}+(j-1)\Delta t_{j}\). We discretize the Wiener processes \(\Delta W_{j}\) with time step \(\Delta t_{j}\), and \(W(t_{j})\) as \(\Delta W_{j}\sim \sqrt{\Delta t_{j}}\ N(0,1)\) and \(W(t_{j})=W(t_{j-1})+\Delta W_{j}\), \(j=2,3,\cdots ,m\) with \(W(t_{1})=\Delta W_{1}\).
Let \(\hat{N}_{j}\) be the discretized aggregate death counts satisfying the solution (2.7) at time \(t_{j}\), \(j=1,2,\cdots ,m\). We estimate \(\hat{N}_{j}\) using (2.7) as
where \(\widehat{\mathcal {K}}, \hat{\mu }, \hat{\sigma }\) are calculated in Sect. 3. In order to generate pseudo samples for each point \(\hat{N}_{j}\), we define \(\Delta W_{j}^{l}=W_{j}^{l}-W_{j-1}^{l}\), \(j=1,2,\cdots ,m\), \(l=1,2,\cdots ,L\), for sample size m and L number of simulations. Using Milstein scheme (Gaines and Lyons 1994), the l-th discretized solution \(N_{j}^{l}\equiv N(t_{j})^{l}\) of (2.3) satisfies
for \(j=1,2,\cdots ,m\), \(l=1,2,\cdots ,L\). We use the estimate (5.1), together with the estimated parameters \(\hat{\mathcal {K}}\), \(\hat{\mu }\) and \(\hat{\sigma }\) in (3.5)–(3.6) to fit the aggregate COVID-19 death counts in the fifty states in the United States from the period when the first death count is recorded to June 20, 2021, and also to forecast from June 21, 2021 to June 24, 2021. Model (5.2) and (3.1) are used to generate the probability density function for the aggregate death counts for each time \(t_{j}\). Let \(\hat{N}_{j,\sigma =0}\) denote the deterministic equivalent of (5.1), which is the discretization of solution of (1.1) with \(\sigma =0\). The parameters in \(\hat{N}_{j,\sigma =0}\) are also estimated using the Non-Linear Least Square estimate scheme (Coleman and Li 1996; May 1963) for model comparison purposes. Denote the root mean square error for the deterministic and stochastic discretization scheme by \(\text{ RMSE }\) and \(\text{ RAMSE }\), respectively. We define
where \(\left\{ N(t_{j})\right\} _{j=1}^{m}\) is the real aggregate death counts data. In order to show the superiority of model (2.3) over (1.1), we compare the root mean square errors \(\text{ RMSE }\) and \(\text{ RAMSE }\) and show that model (2.3) has a smaller root mean square error.
Table 1 shows the parameter estimates for the stochastic model (2.3) together with the root mean square errors \(\text{ RMSE }\) and \(\text{ RAMSE }\) in (5.3) for the deterministic and stochastic cases, respectively, using the Covid-19 aggregate death counts in the United States for the period when the first death case is reported to June 20, 2021. Here, \(N_{0}\) denotes the estimate of the starting value when the first death case is reported. The expected first time the aggregate death counts is more than half its carrying capacity is calculated. The root mean square error \(\text{ RMSE }\) was calculated by first estimating the parameters in the deterministic model (1.1) using the Non-Linear Least Square estimate scheme (May 1963; Coleman and Li 1996). The estimated parameters for the deterministic model are not reported in this work. Within the analysis period, we see that the expected aggregate death counts started slowing down around mid December when the first vaccine was administered for most states. A quick comparison of \(\text{ RMSE }\) and \(\text{ RAMSE }\) in Table 1 shows that the stochastic model (2.3) performs better than the deterministic model (1.1) in describing the trajectory of the aggregate count of Covid-19 in the United States. In order to minimize space, we only show the real and simulated death counts for 24 out of 50 states in the United States in Figs. 1 and 2.
Figures 1 and 2 show the real and simulated aggregate death counts for some of the fifty states in the United States. In order to forecast the aggregate death counts from June 21, 2021 to June 24, 2021, we analyze the data starting from June 4, 2021 to June 20, 2021. The parameter estimates are shown in Table 2.
Table 2 contains parameter estimates derived using data set from June 4, 2021 to June 20, 2021. Here, \(N_{0}\) denotes the estimate of the starting value for June 4, 2021. These estimates are used in the forecast for the aggregate death counts from June 21, 2021 to June 24, 2021. In Table 2, \(N_{06/21/2021}\) denotes the forecast aggregate death counts estimate for June 21, 2021. The \(95\%\) confidence interval for the forecast estimate is also calculated and presented in Figs. 3 and 4.
Figures 3 and 4 show simulated and forecast estimate for the aggregate death counts of Covid-19 in the United States. The parameter estimates used for the simulation are given in Table 2 using the data set for June 4, 2021 to June 20, 2021. These parameters are used to forecast the aggregate death counts for June 21, 2021 to June 24, 2021.
In order to verify the validity of the obtained probability density function (3.1), we show in the following graphs the comparison of the probability density function for the random variable \(N_{T}^{l}\) given in (5.2) for \(l=10,000\) simulations with the probability density function in (3.1) by setting \(t=t_{m}=T\). The time \(t=T\) corresponds to the day: June 24, 2021.
By generating the histogram of \(\{N_{T}^{l}\}_{l=1}^{10,000}\) in (5.2), we show the comparison of the graphs of the probability density function for the random variable \(N_{T}^{l}\) and \(p_{N}(N|T,N_{0})\) in Figs. 5 and 6. The graphs show that the probability density function concentrates on a particular value. To know what this value is, we calculate the expected value of the aggregate death counts obtained in (3.7) for each of the fifty states in the United States and noticed the probability density function concentrates on the expected aggregate count \(\mathbb {E}\left( N(T)\right)\) on June 24, 2021, with \(t=t_{m}=T\) denoting June 24, 2021. We also noticed that this value is close to the equilibrium point \(\mathcal {K}\), which is the maximum aggregate death counts as at the time this research is conducted. Define
The comparison of the value \(N_{T,\max }\) where the probability density function \(p_{N}(N|T,N_{0})\) concentrates on, with the expected aggregate count on June 24, 2021 is shown in Table 3. The plot of the expected aggregate death counts is plotted in Figs. 7 and 8 as a function of time.
Using the result obtained in (3.7), we plot the graph of the expected value of the aggregate death counts \(\mathbb {E}\left( N(t)|N_{0}\right)\) with time for each states in the United States in Figs. 7 and 8. The graph shows that the expected value of the aggregate death counts is increasing for \(t\ge 0\). We also see from the graphs that the expected aggregate death counts started slowing down sometimes around the month of December 2020 for most states. This number is still slowing down as at the time this analysis is carried out (June 2021).
6 Summary and Discussion
In this work, we study and analyze the aggregate death counts N(t) resulting from the Covid-19 virus infection in the United States. Recent studies use the deterministic logistic model to analyze this counts by assuming the death count’s per capita growth rate of the Covid-19 virus is constant over time. Our studies show that this is not the case. We assume, based on some analysis, that this growth rate can be affected by external perturbations causing fluctuations that can be modeled as a white noise described using a Wiener process. This assumption is used to modify and extend the existing logistic model to a stochastic differential equation. We analyze this model by first showing that it has a unique solution and its solution is bounded, with probability one. Analogue to an increasing function, we show that the process N(t) is a submartingale that converges almost surely to a random variable on the long run if \(\mu\) is greater than \(\sigma ^{2}/2\). By calculating the first hitting time when the aggregate death counts reach \(\mathcal {K}-\epsilon\) for small \(\epsilon >0\), we calculate an approximate expected time when the death counts slow down for each state in the United States. To do this, we first calculate the probability of the first passage time \(\tau _{\epsilon }\) described in (4.2), and later calculate the expected first passage time. Our result shows that the aggregate death counts is now slowing down as at June 2021 when this analysis is conducted. By comparing the estimate \(\mathcal {K}/2\) with the estimate N(T) and \(\mathbb {E}\left[ N({T})|N_{0}\right]\) for the current day, June 24, 2021, our analysis shows that the Covid-19 death crisis slows down in the month of June in most states in the United States.
By deriving the transition probability density function for the process N(t), we show that the expected aggregate death counts is bounded, and approaches \(\mathcal {K}\) asymptotically. We also studied the distribution of the counts for the time T when this analysis is conducted and our result shows that the distribution concentrates on the expected aggregate count for that day (June 24, 2021).
Using the Maximum Likelihood Estimate scheme, we estimate the epidemiological parameters \(\mathcal {K}\), \(\mu\), and \(\sigma\) using (3.3)–(3.4). These are used together with the Milstein scheme (5.2) to simulate and forecast the aggregate death counts for each states in the United States. A \(95\%\) confidence interval is provided for the forecast result. To show that our model (2.3) performs better than existing model (1.1), we compare the root mean square errors \(\text{ RAMSE }\) and \(\text{ RMSE }\) for the stochastic and deterministic models, respectively, and show that \(\text{ RAMSE }<\text{ RMSE }\). This research is still ongoing and an update will be provided when available.
Notes
https://www.cdc.gov/museum/timeline/covid19.html, accessed 06.04.2022
https://covid.cdc.gov/covid-data-tracker, accessed 06.24.2021 at 9:09PM.
https://www.healthline.com/health/vaccinations/johnson-and-johnson-vaccine, accessed 06.07.2021.
Two solutions \(x_{1}:[t_{0},T]\rightarrow \Re ^{+}\) and \(x_{2}:[t_{0},T]\rightarrow \Re ^{+}\) are said to be equivalent if \(\mathbb {P}\left( x_{1}(t)=x_{2}(t)\ \text{ for } \text{ all } t\in [t_{0},T]\right) =1\).
References
Arnold L (1974) Stochastic differential equations: theory and applications. Wiley, New York
Beddington JR, May RM (1977) Harvesting natural populations in a randomly fluctuating environment. Science 197:463–465
Baud D, Qi X, Nielsen-Saines K, Musso D, Pomar Léo, Favre Guillaume (2020) Real estimates of mortality following COVID-19 infection. Lancet Infect Dis 20(7):773. https://doi.org/10.1016/S1473-3099(20)30195-X
Bhapkar HR, Mahalle PN, Dey N, Santosh KC (2020) Revisited COVID-19 mortality and recovery rates: are we missing recovery time period? J Med Syst 44(12):202
Coleman TF, Li Y (1996) An interior, trust region approach for nonlinear minimization subject to bounds. SIAM J Optim 6:418–45
Gaines JG, Lyons TJ (1994) Random generation of stochastic area integrals. SIAM J Appl Math 54(4):1132–1146
Gardiner CW (1985) Handbook of stochastic methods for physics, chemistry and the natural sciences. Springer-Verlag, New York
Lv J, Liu H, Zou X (2019) Stationary distribution and persistence of a stochastic predator-prey model with a functional response. J Appl Anal Comput 9(1):1–11
Kaciroti NA, Lumeng C, Parekh V, Boulton ML (2021) A bayesian mixture model for predicting the COVID-19 related mortality in the United States. Am J Trop Med Hyg 104(4):1484–1492
Khasminskii R (2012) Stochastic stability of differential equations, 2nd edn. Springer-Verlag, Berlin Heidelberg, p 66
Kloeden PE, Platen E (1995) Numerical solution of stochastic differential equations. Springer-Verlag, New York
Lagarto S, Braumann CA (2014) Modeling human population death rates: ABi-dimensional stochastic Gompertz model with correlated wiener processes. In: Pacheco A, Santos R, Oliveira M, Paulino C (eds) New advances in statistical modeling and applications. Studies in theoretical and applied statistics. Springer, Cham
Li W, Wang K (2010) Optimal harvesting policy for general stochastic logistic population model. J Math Anal Appl 368:420–428
Linka K, Peirlinck M, Kuhl E (2020) The reproduction number of COVID-19 and its correlation with public health interventions. Comput Mech 66(4):1035–1050. https://doi.org/10.1007/s00466-020-01880-8
Lungu EM, Øksendal B (1997) Optimal harvesting from a population model in a Stochastic Crowded Environment. Math Biosci 145:47–75
May RM (1963) An algorithm for least-squares estimation of nonlinear parameters. SIAM J Appl Math 11(2):431–41
Mao X (2007) Stochastic differential equations and applications, 2nd edn. Horwood, Chichester
Mazzuco S, Scarpa B, Zanotto L (2018) A mortality model based on a mixture distribution function. Popul Stud 72(3):1–10
Mendez V, Campos D, Horsthemke W (2012) Stochastic fluctuations of the transmission rate in the susceptible-infected-susceptible epidemic model. Phys Rev E 86:011919
Ndairou F, Area I, Nieto JJ, Torres DFM (2020) Mathematical modeling of COVID-19 transmission dynamics with a case study of Wuhan. Chaos Solitons Fractals 135:109846
Okuonghae D, Omame A (2020) Analysis of a mathematical model for COVID-19 population dynamics in Lagos, Nigeria. Chaos Solitons Fractals 139:110032
Øksendal B (2003) Stochastic differential equations, An introduction with applications. Springer-Verlag, Berlin Heidelberg, New York
Ladde GS, Otunuga OM, Ladde NS (2020) Local lagged adapted generalized method of moments Dynamic Process. U.S. Patent Number: 10719578
Otunuga OM (2021) Time-dependent probability distribution for the number of infection in a stochastic SIS model: case study COVID-19. Chaos Solitons Fractals 147:110983
Otunuga OM (2021) Time-dependent probability density function for general stochastic logistic population model with harvesting effort. Phys A 573:1–33
Otunuga OM (2020) Qualitative analysis of a stochastic SEITR epidemic model with multiple stages of infection and treatment. Infect Dis Modell 5:61–90
Otunuga OM (2019) Closed-form probability distribution of number of infections at a given time in a stochastic SIS epidemic model. Heliyon 5:1–12
Mummert A, Otunuga OM (2019) Parameter identification for a stochastic SEIRS epidemic model: case study influenza. J Math Biol 79(2):705–729. https://doi.org/10.1007/s00285-019-01374-z
Otunuga OM (2018) Global stability for a 2n + 1 dimensional HIV/AIDS epidemic model with treatments. Math Biosci 5:138–52
Pelinovsky E, Kurkin A, Kurkina O, Kokoulina M, Epifanova A (2020) Logistic equation and COVID-19. Chaos Solitons Fractals 140:110241
Pella JS, Tomlinson PK (1969) A generalised stock-production model. Bull Int Am Trop Tuna Commun 13:421–496
Prajneshu (1980) Time dependent solution of the logistic model for population growth in random environment. J Appl Prob 17:1083–1086
Santosh KC (2020) COVID-19 prediction models and unexploited data. J Med Syst 44:170
Satpathy S, Mangla M, Sharma N, Deshmukh H, Mohanty S (2021) Predicting mortality rate and associated risks in COVID-19 patients. Spat Inf Res 29(4):455–464
Stutt Rojh, Retkute R, Bradley M, Gilligan CA, Colvin J (2020) A modelling framework to assess the likely effectiveness of facemasks in combination with ‘lock-down’ in managing the COVID-19 pandemic. Proc R Soc A 476:20200376
Wang P, Zheng X, Li J, Zhu B (2020) Prediction of epidemic trends in COVID-19 with logistic model and machine learning technics. Chaos Solitons Fractals 139:110058
Verhulst Pierre-Francois (1838) Notice sur la loi que la population poursuit dans son accroissement. Correspondance Mathématique et Physique. 10:113-121. Retrieved 3 Dec 2014
West BJ, Bulsara AR, Lindenberg K, Seshadri V, Shuler KE (1979) Stochastic processes with non-additive fluctuations: I. Itô and Stratonovich calculus and the effects of correlations. Phys A 97(2):211–233
Wong E, Zakai M (1965) On the convergence of ordinary integrals to stochastic integrals. Ann Math Stat 36(5):1560–1564
Wu JT, Leung K, Leung GM (2020) ’ ’Nowcasting and forecasting the potential domestic and international spread of the 2019-nCoV outbreak originating in Wuhan, China: a modelling study. Lancet 35(10225):689–697
Yang B, Cai Y, Wang K, Wang W (2019) Optimal harvesting policy of logistic population model in a randomly fluctuating environment. Phys A 526:120817
Zocchetti C, Consonni D (1994) Mortality rate and its statistical properties. Med Lav 85(4):327–43
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Otunuga, O.M., Otunuga, O. Stochastic Modeling and Forecasting of Covid-19 Deaths: Analysis for the Fifty States in the United States. Acta Biotheor 70, 25 (2022). https://doi.org/10.1007/s10441-022-09449-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10441-022-09449-z