Abstract
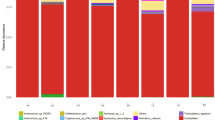
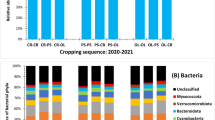
Biocontrol strategies using organic substrates such as wood fibers and biocontrol agents such as Trichoderma are currently developed to control soil pathogens such as Fusarium oxysporum. Nonetheless, such biocontrol methods give discording results, notably because microbial communities of organic substrates actually are not taken into account. Therefore, there is a lack of information concerning the variability of microbial composition related to the organic substrate type. Here we studied peat, wood and coir fibers, that are substrates known for their different biocontrol efficiency against Fusarium wilt of cucumber. We analyzed in microcosms the microbial composition of wood fibers, coir fibers and peat, incubated up to 60 days, by using an amplicon-sequencing approach based on 16S rRNA gene for bacteria and the internal transcribed spacer (ITS) for fungi. Diversity was assessed by sequencing the 16S rRNA for bacteria and ITS2 region for fungi. Results showed that bacterial richness was threefold higher for coir fiber and peat than for wood fiber. Fungal richness was three times higher for wood and coir fibers compared to peat. Bacterial and fungal patterns showed a dominance of α- and γ- Proteobacteria and Sordariomycetes for coir fiber; β- and γ-Proteobacteria and Eurotiomycetes for wood fibers; Flavobacteria, Leotiomycetes and Sordariomycetes for peat. In conclusion, results show that substrates have different microbial composition. Finally, for a proper use of a biocontrol strategy is important to take into account the type of substrate.


Similar content being viewed by others
References
Bardin SD, Huang HC, Moyer JR (2004) Control of Pythium damping-off of sugar beet by seed treatment with crop straw powders and a biocontrol agent. Biol Control 29:453–460. doi:10.1016/j.biocontrol.2003.09.001
Barret M, Briand M, Bonneau S, Préveaux A, Valière S, Bouchez O, Hunault G, Simoneau P, Jacques M-A (2015) Emergence Shapes the Structure of the Seed Microbiota. Appl Environ Microbiol 81:1257–1266. doi:10.1128/AEM.03722-14
Bernard L, Mougel C, Maron P-A, Nowak V, Lévêque J, Henault C, Haichar FEZ, Berge O, Marol C, Balesdent J, Gibiat F, Lemanceau P, Ranjard L (2007) Dynamics and identification of soil microbial populations actively assimilating carbon from 13C-labelled wheat residue as estimated by DNA- and RNA-SIP techniques. Environ Microbiol 9:752–764. doi:10.1111/j.1462-2920.2006.01197.x
Bonanomi G, Antignani V, Capodilupo M (2010) Identifying the characteristics of organic soil amendments that suppress soilborne plant diseases. Soil Biol Biochem 42:136–144. doi:10.1016/j.soilbio.2009.10.012
Chaparro JM, Sheflin AM, Manter DK, Vivanco JM (2012) Manipulating the soil microbiome to increase soil health and plant fertility. Biol Fertil Soils 48:489–499. doi:10.1007/s00374-012-0691-4
de Boer W, Folman LB, Summerbell RC, Boddy L (2005) Living in a fungal world: impact of fungi on soil bacterial niche development. FEMS Microbiol Rev 29:795–811. doi:10.1016/j.femsre.2004.11.005
Domeño I, Irigoyen I, Muro J (2011) Comparison of traditional and improved methods for estimating the stability of organic growing media. Sci Hortic 130:335–340. doi:10.1016/j.scienta.2011.07.012
Faessel L, Nassr N, Lebeau T, Walter B (2008) Effects of the Plant Defence Inducer, Acibenzolar-S-Methyl, on Hypocotyl Rot of Soybean Caused by Rhizoctonia solani AG-4. J Phytopathol 156:236–242. doi:10.1111/j.1439-0434.2007.01367.x
Fierer N, Bradford MA, Jackson RB (2007) Toward an ecological classification of soil bacteria. Ecology 88:1354–1364. doi:10.1890/05-1839
Gamalero E, Cesaro P, Cicatelli A, Todeschini V, Musso C, Castiglione S, Fabiani A, Lingua G (2012) Poplar clones of different sizes, grown on a heavy metal polluted site, are associated with microbial populations of varying composition. Sci Total Environ 425:262–270. doi:10.1016/j.scitotenv.2012.03.012
Koohakan P, Ikeda H, Jeanaksorn T, Tojo M, Kusakari S-I, Okada K, Sato S (2004) Evaluation of the indigenous microorganisms in soilless culture: occurrence and quantitative characteristics in the different growing systems. Sci Hortic 101:179–188. doi:10.1016/j.scienta.2003.09.012
Kozich JJ, Westcott SL, Baxter NT, Highlander SK, Schloss PD (2013) Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq illumina sequencing platform. Appl Environ Microbiol 79:5112–5120. doi:10.1128/AEM.01043-13
Kuramae EE, Yergeau E, Wong LC, Pijl AS, van Veen JA, Kowalchuk GA (2012) Soil characteristics more strongly influence soil bacterial communities than land-use type. FEMS Microbiol Ecol 79:12–24. doi:10.1111/j.1574-6941.2011.01192.x
Laslo É, György É, Mara G, Tamás É, Ábrahám B, Lányi S (2012) Screening of plant growth promoting rhizobacteria as potential microbial inoculants. Crop Prot 40:43–48. doi:10.1016/j.cropro.2012.05.002
Lebeau T (2011) Bioaugmentation for in situ soil remediation: how to ensure the success of such a process. In: Singh A, Parmar N, Kuhad RC (eds) Bioaugmentation, biostimulation and biocontrol, soil biology. Springer, Berlin, pp 129–186. doi:10.1007/978-3-642-19769-7_7
Lecomte C, Alabouvette C, Edel-Hermann V, Robert F, Steinberg C (2016) Biological control of ornamental plant diseases caused by Fusarium oxysporum: a review. Biol Control 101:17–30. doi:10.1016/j.biocontrol.2016.06.004
Lluch J, Servant F, Païssé S, Valle C, Valière S, Kuchly C, Vilchez G, Donnadieu C, Courtney M, Burcelin R, Amar J, Bouchez O, Lelouvier B (2015) The characterization of novel tissue microbiota using an optimized 16S metagenomic sequencing pipeline. PLoS ONE 10:1–22. doi:10.1371/journal.pone.0142334
Mercier J, Manker DC (2005) Biocontrol of soil-borne diseases and plant growth enhancement in greenhouse soilless mix by the volatile-producing fungus Muscodor albus. Crop Prot 24:355–362. doi:10.1016/j.cropro.2004.09.004
Montagne V, Charpentier S, Cannavo P, Capiaux H, Grosbellet C, Lebeau T (2015) Structure and activity of spontaneous fungal communities in organic substrates used for soilless crops. Sci Hortic 192:148–157. doi:10.1016/j.scienta.2015.06.011
Montagne V, Capiaux H, Cannavo P, Charpentier S, Renaud S, Liatard E, Grosbellet C, Lebeau T (2016) Protective effect of organic substrates against soil-borne pathogens in soilless cucumber crops. Sci Hortic 206:62–70. doi:10.1016/j.scienta.2016.04.035
Muyzer G, de Waal EC, Uitterlinden AG (1993) Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol 59:695–700
Russell JB, Rychlik JL (2001) Factors that alter rumen microbial ecology. Science 292:1119–1122. doi:10.1126/science.1058830
Sabuquillo P, De Cal A, Melgarejo P (2010) Development of a dried Penicillium oxalicum conidial formulation for use as a biological agent against Fusarium wilt of tomato: selection of optimal additives and storage conditions for maintaining conidial viability. Biol Control 54:221–229. doi:10.1016/j.biocontrol.2010.05.010
Schellenberger S, Kolb S, Drake HL (2010) Metabolic responses of novel cellulolytic and saccharolytic agricultural soil Bacteria to oxygen. Environ Microbiol 12:845–861. doi:10.1111/j.1462-2920.2009.02128.x
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Horn DJV, Weber CF (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541. doi:10.1128/AEM.01541-09
Turnbaugh PJ, Ridaura VK, Faith JJ, Rey FE, Knight R, Gordon JI (2009) The effect of diet on the human gut microbiome: a metagenomic analysis in humanized gnotobiotic mice. Sci Transl Med 1:6ra14-6ra14. doi:10.1126/scitranslmed.3000322
Vivant A-L, Garmyn D, Maron P-A, Nowak V, Piveteau P (2013) Microbial diversity and structure are drivers of the biological barrier effect against Listeria monocytogenes in soil. PLoS ONE 8:1–11. doi:10.1371/journal.pone.0076991
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protocols 38:315–322
Acknowledgements
We wish to thank the Florentaise firm, the ANRT (Association Nationale de la Recherche et de la Technologie—CIFRE convention No. 2012/1062) for funding this work. We are particularly grateful to the NED team (UMR1289 TANDEM) and the GeT-PLaGE plateform in Toulouse using Illumina Miseq technology (Génopole Toulouse-Midi-pyrénées, INRA Auzeville, 24 Chemin de Borde Rouge-Auzeville, CS52627, 31326 Castanet-Tolosan, France).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
No conflict of interest declared.
Rights and permissions
About this article
Cite this article
Montagne, V., Capiaux, H., Barret, M. et al. Bacterial and fungal communities vary with the type of organic substrate: implications for biocontrol of soilless crops. Environ Chem Lett 15, 537–545 (2017). https://doi.org/10.1007/s10311-017-0628-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10311-017-0628-0