Abstract
Objectives
Rheumatoid arthritis (RA) is a chronic systemic autoimmune disorder characterized by progressive synovial inflammation and joint destruction, with a largely unknown etiology. Studies have suggested that autophagy and its expression may be involved in the pathogenesis of RA; however, autophagy-related genes in RA are still largely unidentified. Therefore, in this study, we aimed to identify and validate autophagy-related genes in RA.
Methods
We identified differentially expressed autophagy-related genes between patients with RA and healthy individuals using gene expression profiles in the GSE55235 dataset and R software. Subsequently, correlation analysis, protein–protein interaction, gene ontology enrichment, and Kyoto Encyclopedia of Genes and Genomes pathway enrichment analyses were carried out using these differentially expressed autophagy-related genes. Finally, our results were validated by examining the expression of differentially expressed autophagy-related hub genes in clinical samples using qRT-PCR.
Results
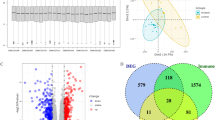
We identified 52 potential autophagy-related genes in RA based on bioinformatic analyses. Ten hub genes, CASP8, CTSB, TNFSF10, FADD, BAX, MYC, FOS, CDKN1A, GABARAPL1, and BNIP3, were validated to be differentially expressed and may serve as valuable prognostic markers and new potential therapeutic targets for RA via the regulation of autophagy.
Conclusions
Our results may help improve the understanding of RA pathogenesis. Autophagy-related genes in RA could be valuable biomarkers for diagnosis and prognosis and they might be exploited clinically as therapeutic targets in the future.
Key Points • CASP8, CTSB, TNFSF10, FADD, BAX, MYC, FOS, CDKN1A, GABARAPL1, and BNIP3 may be autophagy-related hub genes correlated with the pathogenesis of RA. |






Similar content being viewed by others
References
Wei K, Nguyen HN, Brenner MB (2021) Fibroblast pathology in inflammatory diseases. J Clin Invest 131(20). https://doi.org/10.1172/JCI149538
Zhao J, Jiang P, Guo S et al (2021) Apoptosis, autophagy, NETosis, necroptosis, and pyroptosis mediated programmed cell death as targets for innovative therapy in rheumatoid arthritis. Front Immunol 12:809806. https://doi.org/10.3389/fimmu.2021.809806
Jin M, Zhang Y (2020) Autophagy and autoimmune diseases. Adv Exp Med Biol 1207:405–408. https://doi.org/10.1007/978-981-15-4272-5_28
Martinez J, Cunha LD, Park S et al (2016) Noncanonical autophagy inhibits the autoinflammatory, lupus-like response to dying cells. Nature 533:115–119. https://doi.org/10.1038/nature17950
Dumit VI, Kuttner V, Kappler J et al (2014) Altered MCM protein levels and autophagic flux in aged and systemic sclerosis dermal fibroblasts. J Invest Dermatol 134:2321–2330. https://doi.org/10.1038/jid.2014.69
Frech T, De Domenico I, Murtaugh MA et al (2014) Autophagy is a key feature in the pathogenesis of systemic sclerosis. Rheumatol Int 34:435–439. https://doi.org/10.1007/s00296-013-2827-8
Zhang Y, Vasheghani F, Li YH et al (2015) Cartilage-specific deletion of mTOR upregulates autophagy and protects mice from osteoarthritis. Ann Rheum Dis 74:1432–1440. https://doi.org/10.1136/annrheumdis-2013-204599
Karami J, Masoumi M, Khorramdelazad H et al (2020) Role of autophagy in the pathogenesis of rheumatoid arthritis: latest evidence and therapeutic approaches. Life Sci 254:117734. https://doi.org/10.1016/j.lfs.2020.117734
Woetzel D, Huber R, Kupfer P et al (2014) Identification of rheumatoid arthritis and osteoarthritis patients by transcriptome-based rule set generation. Arthritis Res Ther 16:R84. https://doi.org/10.1186/ar4526
Kasai M, Tanida I, Ueno T et al (2009) Autophagic compartments gain access to the MHC class II compartments in thymic epithelium. J Immunol 183:7278–7285. https://doi.org/10.4049/jimmunol.0804087
Ciccacci C, Perricone C, Alessandri C et al (2018) Evaluation of ATG5 polymorphisms in Italian patients with systemic lupus erythematosus: contribution to disease susceptibility and clinical phenotypes. Lupus 27:1464–1469. https://doi.org/10.1177/0961203318776108
International Consortium for Systemic Lupus Erythematosus G, Harley JB, Alarcon-Riquelme ME et al (2008) Genome-wide association scan in women with systemic lupus erythematosus identifies susceptibility variants in ITGAM, PXK, KIAA1542 and other loci. Nat Genet 40:204–210. https://doi.org/10.1038/ng.81
Kamel AM, Badary MS, Mohamed WA et al (2020) Evaluation of autophagy-related genes in Egyptian systemic lupus erythematosus patients. Int J Rheum Dis 23:1226–1232. https://doi.org/10.1111/1756-185X.13910
Mahil SK, Twelves S, Farkas K et al (2016) AP1S3 Mutations cause skin autoinflammation by disrupting keratinocyte autophagy and up-regulating IL-36 production. J Invest Dermatol 136:2251–2259. https://doi.org/10.1016/j.jid.2016.06.618
Igci M, Baysan M, Yigiter R et al (2016) Gene expression profiles of autophagy-related genes in multiple sclerosis. Gene 588:38–46. https://doi.org/10.1016/j.gene.2016.04.042
Sorice M, Iannuccelli C, Manganelli V et al (2016) Autophagy generates citrullinated peptides in human synoviocytes: a possible trigger for anti-citrullinated peptide antibodies. Rheumatology (Oxford) 55:1374–1385. https://doi.org/10.1093/rheumatology/kew178
Manganelli V, Recalchi S, Capozzi A et al (2018) Autophagy induces protein carbamylation in fibroblast-like synoviocytes from patients with rheumatoid arthritis. Rheumatology (Oxford) 57:2032–2041. https://doi.org/10.1093/rheumatology/key174
An Q, Yan W, Zhao Y et al (2018) Enhanced neutrophil autophagy and increased concentrations of IL-6, IL-8, IL-10 and MCP-1 in rheumatoid arthritis. Int Immunopharmacol 65:119–128. https://doi.org/10.1016/j.intimp.2018.09.011
van Loosdregt J, Rossetti M, Spreafico R et al (2016) Increased autophagy in CD4(+) T cells of rheumatoid arthritis patients results in T-cell hyperactivation and apoptosis resistance. Eur J Immunol 46:2862–2870. https://doi.org/10.1002/eji.201646375
Kato M, Ospelt C, Gay RE et al (2014) Dual role of autophagy in stress-induced cell death in rheumatoid arthritis synovial fibroblasts. Arthritis Rheumatol 66:40–48. https://doi.org/10.1002/art.38190
Huang RZ, Zheng J, Liu FL et al (2021) A novel autophagy-related marker for improved differential diagnosis of rheumatoid arthritis and osteoarthritis. Front Genet 12:743560. https://doi.org/10.3389/fgene.2021.743560
Kumar P, Yao LJ, Saidin S et al (2018) Molecular mechanisms of autophagic memory in pathogenic T cells in human arthritis. J Autoimmun 94:90–98. https://doi.org/10.1016/j.jaut.2018.07.014
Xu C, Vitone GJ, Inoue K et al (2019) Identification of a novel role for Foxo3 Isoform2 in osteoclastic inhibition. J Immunol 203:2141–2149. https://doi.org/10.4049/jimmunol.1900707
Dai Y, Ding J, Yin W et al (2018) Increased autophagy enhances the resistance to tumor necrosis factor-alpha treatment in rheumatoid arthritis human fibroblast-like synovial cell. Biomed Res Int 2018:4941027. https://doi.org/10.1155/2018/4941027
Zhu L, Wang H, Wu Y et al (2017) The autophagy level is increased in the synovial tissues of patients with active rheumatoid arthritis and is correlated with disease severity. Mediators Inflamm 2017:7623145. https://doi.org/10.1155/2017/7623145
Okada Y, Wu D, Trynka G et al (2014) Genetics of rheumatoid arthritis contributes to biology and drug discovery. Nature 506:376–381. https://doi.org/10.1038/nature12873
Wang H, Wang Z, Wang L et al (2020) IL-6 promotes collagen-induced arthritis by activating the NLRP3 inflammasome through the cathepsin B/S100A9-mediated pathway. Int Immunopharmacol 88:106985. https://doi.org/10.1016/j.intimp.2020.106985
Singh AK, Haque M, Madarampalli B et al (2021) Ets-2 propagates IL-6 trans-signaling mediated osteoclast-like changes in human rheumatoid arthritis synovial fibroblast. Front Immunol 12:746503. https://doi.org/10.3389/fimmu.2021.746503
Yang X, Wang J, Liu C et al (2005) Cleavage of p53-vimentin complex enhances tumor necrosis factor-related apoptosis-inducing ligand-mediated apoptosis of rheumatoid arthritis synovial fibroblasts. Am J Pathol 167:705–719. https://doi.org/10.1016/S0002-9440(10)62045-7
Mouasni S, Gonzalez V, Schmitt A et al (2019) The classical NLRP3 inflammasome controls FADD unconventional secretion through microvesicle shedding. Cell Death Dis 10:190. https://doi.org/10.1038/s41419-019-1412-9
Misra S, Bagchi A, Sarkar A et al (2021) Methotrexate and theaflavin-3, 3’-digallate synergistically restore the balance between apoptosis and autophagy in synovial fibroblast of RA: an ex vivo approach with cultured human RA FLS. Inflammopharmacology 29:1427–1442. https://doi.org/10.1007/s10787-021-00857-0
Lee YZ, Guo HC, Zhao GH et al (2020) Tylophorine-based compounds are therapeutic in rheumatoid arthritis by targeting the caprin-1 ribonucleoprotein complex and inhibiting expression of associated c-Myc and HIF-1alpha. Pharmacol Res 152:104581. https://doi.org/10.1016/j.phrs.2019.104581
Yamashita T, Yao Z, Li F et al (2007) NF-kappaB p50 and p52 regulate receptor activator of NF-kappaB ligand (RANKL) and tumor necrosis factor-induced osteoclast precursor differentiation by activating c-Fos and NFATc1. J Biol Chem 282:18245–18253. https://doi.org/10.1074/jbc.M610701200
Gang X, Xu H, Si L et al (2018) Treatment effect of CDKN1A on rheumatoid arthritis by mediating proliferation and invasion of fibroblast-like synoviocytes cells. Clin Exp Immunol 194:220–230. https://doi.org/10.1111/cei.13161
Kim JK, Kim YS, Lee HM et al (2018) GABAergic signaling linked to autophagy enhances host protection against intracellular bacterial infections. Nat Commun 9:4184. https://doi.org/10.1038/s41467-018-06487-5
Kammouni W, Wong K, Ma G et al (2007) Regulation of apoptosis in fibroblast-like synoviocytes by the hypoxia-induced Bcl-2 family member Bcl-2/adenovirus E1B 19-kd protein-interacting protein 3. Arthritis Rheum 56:2854–2863. https://doi.org/10.1002/art.22853
Funding
This study was supported by the National Nature Science Foundation of China (grant number 82071752).
Author information
Authors and Affiliations
Contributions
Q-h Y and D-d F designed the experiments; D-d F and P-y T collected and analyzed the data; L J and P-y T collected the clinical samples and performed the molecular experiments; D-d F wrote the manuscript; Q-h Yu and Y Q revised this paper.
Corresponding author
Ethics declarations
Disclosures
None.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fan, Dd., Tan, Py., Jin, L. et al. Bioinformatic identification and validation of autophagy-related genes in rheumatoid arthritis. Clin Rheumatol 42, 741–750 (2023). https://doi.org/10.1007/s10067-022-06399-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10067-022-06399-2