Abstract
Soil functioning is closely linked to the interactions between biological communities with the physical environment. Yet, the impact of plant community attributes on metabolic processes promoting soil nutrient cycling remains largely unknown. We hypothesized that the plant community acts as a regulating agent of nutrient mobilization in soils according to the phylogenetic and morpho-functional traits of plant species of which it is composed. Rhizosphere soils were collected in autumn and spring under 32 tree and shrub species in two Mediterranean mixed forests (four plots in each) located in southern Spain, and nine soil enzymatic activities related to C, N and P mobilization were assessed. Phylogeny and morpho-functional traits of plant species were recorded and their imprint in soil enzymatic activities across forests was determined. The results showed a plant phylogenetic signal for N mobilization in both forests, while it varied across forests for non-labile C and P mobilization. The plant phylogenetic signals were primarily driven by lineages that diversified through the Miocene, about 25 Myr ago. In addition, leaf traits and plant’s mycorrhizal type explained soil enzymatic activities independently from phylogeny. C and P mobilization increased under ectomycorrhizal plants, whilst enhanced N mobilization did occur under arbuscular mycorrhizal ones. The plant community composition led to a different carbon and nutrient mobilization degree, which in turn was mediated by distinct microbial communities mirroring differentiated resource-acquisition strategies of plants. Our results highlight the role of plant traits and mycorrhizal interactions in modulating carbon and nutrient cycling in Mediterranean mixed forest soils.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Highlights
-
Plant community structure determines soil carbon and nutrient cycling in Mediterranean forests.
-
A plant community phylogenetic signal is revealed on soil enzymatic activity at a regional scale.
-
Leaf traits and mycorrhizal association type predict soil enzymatic activity.
Introduction
Carbon and nutrient cycling are essential processes for forest ecosystem functioning (Baldrian 2017). The plant species pool and their trait assortment are critical for nutrient turnover regulation (De Groote and others 2018). In fact, above- and below-ground traits may provide important clues about biogeochemical footprints of species within a given habitat, and particularly those associated with leaves and roots, whose composition and phenology can trigger shifts in soil metabolism (Cheeke and others 2017; Urbina and others 2017). Studying the effect of individual plant species and their traits on nutrient cycling and ecosystem dynamics is crucial for understanding ecosystem response to perturbations, notably under the current global change scenario (Chen and others 2017; Wang and others 2021).
Beyond their potential to determine nutrient turnover, traits can reveal ecological differences between species, and likely phylogenetic signals accounting for additional information on the structure of plant communities (de Bello and others 2017), as closely relatives would tend to have similar life-history traits, metabolic rates, phenology or responses to disturbances (Chen and others 2017; López-Angulo and others 2018). As a result, the contribution of plant communities to ecological processes in forests may come determined from their traits (for example, based on species size, physiology, leaf and root habits) (Liese and others 2017) and their phylogenetic composition, that is, position respect to the rest of species across the community phylogeny (López-Angulo and others 2018). However, both the potential weight of plant traits and phylogenetic composition in predicting processes related to carbon and nutrient cycling in forests, and whether these cycling processes can report a phylogenetic signal from plant communities, are still poorly understood (de Bello and others 2017).
In terrestrial ecosystems, plant communities can also shape the structure and functionality of associated microbial communities (Pérez-Izquierdo and others 2019; Bahram and others 2020). Soil microorganisms are key drivers of nutrient mobilization through targeting plant organic matter inputs (that is, litter and root exudates) (Baldrian 2017; Midgley and Sims 2020). By releasing exo-enzymes capable to convert complex organic compounds into simpler and more soluble molecules (Allison and Vitousek 2005), soil microorganisms are directly involved in carbon (C), phosphorus (P) and nitrogen (N) cycling in soils. For example, saprotrophic and mycorrhizal fungi substantially contribute to C cycling in forest soils, as decomposers, or by directly obtaining C from plants (Allison and Vitousek 2005; Tedersoo and others 2020). Mycorrhizal fungi are likewise well-known to facilitate soil P and N mobilization (Tedersoo and others 2020; Lebreton and others 2021), although rates can considerably differ among mycorrhizal types (Averill and others 2019). In fact, mycorrhizal types may greatly vary in their effect on main ecosystem processes, as leading to higher soil C to N ratios in ectomycorrhizal-dominated forests than in arbuscular mycorrhizal-dominated ones (Brzostek and others 2015; Sulman and others 2017). These cues have even been observed when phylogenetic information is incorporated into plant-soil feedback models (Senior and others 2018; Segnitz and others 2020), which reinforces the idea that plant community phylogenetic structure might be key to understand other essential ecological processes such as C and nutrient cycling in soils (Keller and Phillips 2019). This phylogenetic approach remains particularly scarce in Mediterranean forests (Pérez-Izquierdo and others 2019), despite these ecosystems are characterized by highly diverse plant communities with complex phylogenetic and ecological relationships (Alcántara and others 2019; Perea and others 2021).
In this study, we focused on Mediterranean mixed forests to investigate potential links of plant phylogeny and morpho-functional traits with metabolic processes affecting carbon and nutrient cycling in the soil. Given the important role of plants and spatial–temporal factors on soil functioning (Purahong and others 2016; Pérez-Izquierdo and others 2018), we expected (i) to observe a main role of plant species identity on soil enzymatic activities related to carbon and nutrient cycling, together with that of local conditions and seasonality. Whether this was the case, we expected (ii) to detect a plant phylogenetic signal (that is, closely related plant species would tend to display similar carbon and nutrient cycling rates) (Cavender-Bares and others 2009; López-Angulo and others 2018), and (iii) to find phylogenetic independent trait-soil function relationships indicative of the plant community economics spectrum (Phillips and others 2013) (for example, between the labile carbon and nutrient enzymatic activity with fast-growing related traits).
To elucidate these questions, we evaluated different above- and below-ground traits (that is, plant size, leaf structure and habit, and mycorrhizal type) of representative woody and semi-woody plant species of Mediterranean mixed forests, and we determined a range of enzymatic activities involved in C, P and N cycling in soils beneath these plants.
Material and Methods
Study Area and Sampling Design
The study was conducted in secondary mixed forests of Sierras de Cazorla, Segura y Las Villas (Segura, hereafter) and Monte La Sierra (Jaén, hereafter) natural parks located in the province of Jaén, Southern Spain. Both natural parks (hereinafter referred to as ‘forests’) are approximately 100 km distant and have a continental Mediterranean-type climate, at which drought, rainfall seasonal regimes and fire recurrence have largely conditioned the structure and dynamics of biological communities (Rundel and others 2016; Pérez-Valera and others 2018). In the study area, annual mean rainfall ranges between 800–1000 mm and mean annual temperature between 10–12°C. Plant communities of these forests have been exhaustively characterized in previous studies (Alcántara and others 2018, 2019; Perea and others 2021), with pine and oak species dominating the uppermost canopy layer (Pinus nigra subsp. salzmanii J. F. Arnold, Quercus ilex L., Quercus faginea Lam., and Quercus pyrenaica Willd., in Segura and Pinus halepensis Miller, Q. ilex, and Q. faginea in Jaén), and high diversity of understory tree and tall shrub (for example, species of Crataegus, Juniperus, Sorbus, Prunus, Phillyrea, and Pistacia) and short shrub species (for example, genera Thymus, Cistus, Phlomis, Genista and Rosmarinus).
The sampling design was spatially-nested and consisted of eight 50 × 50 m sites distributed across the two natural parks (4 sites in Jaén, 4 sites in Segura) (following the experimental design detailed in Pulgar and others 2017; Alcántara and others 2018). To capture part of the temporal variation, the sampling was carried out in autumn 2016 and spring 2017 (Table S1). The survey included a total of 32 species: 21 in Jaén and 16 in Segura, 5 of which were found in both forests (Table S1). Adults of representative plant species were randomly selected for soil sampling, with a minimum distance of 10 m between individuals. We removed the litter layer and dug 10 × 10 × 20 cm holes in N, SE and SW orientations, 5–50 cm far from the stem of plants. After tracing the main roots of the target plant individual, we collected soil surrounding secondary and fine roots. Soil subsamples were kept at 4°C and transferred to the laboratory. A total of 339 soil samples were collected: 151 in spring and 188 in autumn, 179 in Jaén and 160 in Segura, and ranging between 30 and 59 per site and between 4 and 32 per plant species according to the number of individuals found of each species (Table S1).
Measurement of Extracellular Enzymatic Activities Involved in soil Carbon and Nutrient Cycling
The three soil subsamples per each plant individual were pooled into a single composite sample, roots were removed, and the remaining soil was homogenized and 2 mm sieved. A fresh soil portion was used to determine gravimetric soil moisture by lost weight of samples oven-dried at 105°C for 48 h. A second soil portion was preserved at −20°C for enzymatic analyses, while the rest of soil was air-dried to determine pH (1:5, w:v in H2O) and soil organic matter (SOM) content by loss on ignition at 400°C for 4 h.
Nine potential enzymatic activities related to C, N and P cycling in forest soils (Pierre-Emmanuel and others 2016; Pérez-Izquierdo and others 2017) were determined in the soil samples, under optimal controlled conditions: β-glucosidase (EC 3.2.1.3) and cellobiohydrolase (EC 3.2.1.91) that are glucoside- and cellulose-hydrolyzing enzymes, respectively, and considered as proxies of labile C cycling; β-xylosidase (EC 3.2.1.37) and β-glucuronidase (EC 3.2.1.31) that are hemicellulose-hydrolyzing enzymes and related to recalcitrant C cycling; acid and alkaline phosphatase (EC 3.1.3.2) activities that hydrolyze P-containing organic compounds promoting P cycling; and chitinase (EC 3.2.1.14) and leucine-aminopeptidase (EC 3.4.11.1) activities as proxies of N cycling, respectively cleaving chitin and polypeptides, and hence mobilizing N-enriched residues. In addition, laccase activity (EC 1.10.3.2), a lignin-oxidizing enzyme involved in recalcitrant C cycling was also measured. We followed the experimental procedure described in Pérez-Izquierdo and others (2017) to measure these enzymes. Briefly, 1 g of rhizosphere soil was incubated (100 rpm, at 25°C overnight) in the corresponding buffer (Tris-maleate 40 mM, pH 8 for leucine aminopeptidase; Tris–acetate 10 mM, pH 11 for alkaline phosphatase; and Tris–acetate 10 mM, pH 4.5 for the rest of enzymes). After soil incubation, all enzymes except laccase were determined by fluorogenic assays that were performed by using the substrates methylcoumarin (AMC) for leucine aminopeptidase and methylumbelliferone (MU) for the rest of enzymes. For laccase, a photometric assay was carried out by using 2,2-Azino-bis 3-ethylbenzothiazoline-6-sulfonic acid (ABTS). Enzymatic activities were measured with a Victor microplate reader (Perkin-Elmer Life Sciences, Massachusetts, USA), at wavelengths of excitation/emission of 355/460 nm for fluorogenic assays and 415 nm for laccase activity. Results were expressed at pmol mg soil−1 h−1 SOM−1. Soil enzymatic activities were grouped by carbon or nutrient cycle (β-glucosidase + cellobiohydrolase as proxy for labile C cycling; β-glucuronidase + β-xylosidase + laccase for non-labile C cycling; acid + alkaline phosphatase for P cycling; and chitinase + leucine-aminopeptidase for N cycling), and also by enzyme ratios (C:N, C:P and N:P enzymatic activities). These groupings are well-established biological indicators of the microbial nutrient demand and linked to carbon and nutrient cycling in soils (Sinsabaugh and others 2008, 2009; Waring and others 2014).
Plant Community Morpho-Functional and Phylogenetic Traits
The morpho-functional traits of plants of our study were obtained from the trait database previously built by Perea and others (2021) in the same mixed forests. The dataset is based on measured traits of randomly-selected adult individuals across the same sites, including height (m), basal area, and equivalent basal diameter (EBD, that is, calculated as a mean when individuals had more than one stem, m2), ratio height to EBD, canopy area (m2) and volume (m3), branches density, specific leaf area (SLA), leaf morphology (for example, acicular, lobed, lanceolate, palmate), leaf water content, leaf habit (perennial vs deciduous), leaf size (length, area, fresh weight, dry matter), leaf chemical composition (C, N, H content, C to N ratio). Additionally, we classified plant species by their mycorrhizal type, that is, ECM and AM, according to literature (Bueno and others 2017; Brundrett and Tedersoo 2020). At each forest, the dataset accounted a total of 20 morpho-functional traits for each plant species.
The phylogenetic information of the plant communities studied was extracted from Alcántara and others (2018). We subset the 32 plant species of our study and used the respective phylogenetic distance-based matrix of each forest (built with 21 plant species in Jaén and 16 in Segura) for calculations. Phylogenetic relatedness is considered as a good proxy of functional similarity because traits are usually conserved across lineages (Webb and others 2002; Cavender-Bares and others 2009). To use the phylogeny as function predictor, phylogenetic distances of plant communities of each forest were decomposed via a Principal Coordinate Analysis (PCoA) (de Bello and others 2017) using the pcoa function with the lingoes correction (ape R package). The resulting eigenvectors (20 for Jaén and 15 for Segura), which gathered the weight and placement of taxa across the phylogeny (de Bello and others 2017; LeRoy and others 2020), were used as phylogenetic traits.
Data Analysis
Prior to analyses, data of enzymatic activities were log-transformed for normalization and to avoid biases in ordination approaches (Pierre-Emmanuel and others 2016). All analyses conducted in this study were carried out with functions and packages of the R free software v.4.1.1. (R Core Team 2021).
To determine the effects of the plant species on carbon and nutrient cycling in soils (hypothesis 1), we firstly fitted Linear Mixed-Effect Models (lme4 R package) setting up forest and season as fixed factors, and plant species and site nested in forest as random factors. The dependent variables were soil properties (pH, SOM, soil moisture), and carbon or nutrient cycles and enzyme ratios (C:N, C:P and N:P enzymatic activities). In all models, p-values were corrected by the Bonferroni method. To show the found trends, the relative soil enzymatic activity (%) for each C, N, P cycle and enzyme ratios were calculated as the respective cycle enzymes for a given factor (forest, plant species and season) divided by the total enzymatic activity within that given factor.
To further detect if the plant species printed a phylogenetic signal on soil carbon and nutrient cycling (hypothesis 2) (that is, the tendency of related plant species to resemble each other more than species drawn at random from the same phylogenetic tree), the corresponding plant phylogenetic tree and nutrient cycling matrix of each forest were first merged with the phylo4d function in phylosignal R package (Keck and others 2016). The phylogenetic signal was separately tested per forest by carbon and nutrient cycle (labile C, non-labile C, N, P) and enzyme ratios (C:N, C:P, N:P), by using the phyloSignal function with the Blomberg’s K method (Münkemüller and others 2012; Keck and others 2016), which tests the null hypothesis of absence of phylogenetic signal over a given variable. The rejection of the null hypothesis allowed to confirm that closely related plant species had more similar carbon and nutrient cycling rates in their soil-associated environment (López-Angulo and others 2018). Furthermore, since the phylogenetic signal might be directed by specific clades (that is, local indicators of phylogenetic association – LIPA), we calculated the Local Moran’s Index (LMI), which may detect autocorrelation hotspots across a given plant phylogeny (Keck and others 2016). LMI can show significant positive values (that is, the trait is more similar among closely relatives than expected by chance) or negative values (that is, the trait is more divergent among distantly relatives than expected by chance). The lipaMoran function was used to determine LMI and autocorrelation values were plotted per plant species at each forest with the dotplot.phylo4d function in the phylosignal R package.
To evaluate whether plant phylogenetic traits (based on PCoA eigenvectors) and morpho-functional traits affected soil carbon and nutrient cycling, we performed Redundancy Analyses, which allowed to select the trait predictors and phylogenetic compositional variables that best explained variations in soil enzymatic activities while discarding redundant ones. The redundancy analyses were carried out on the enzymatic activity matrix and using both PCoA eigenvectors and morpho-functional trait matrices per forest (Jaén and Segura) with the rda function in vegan R package. Before conducting these analyses, covarying plant community trait predictors were filtered with the ordistep function and the variance inflation factor (VIF) analysis by using the vif.cca function in vegan. Values in VIF > 5 indicated the predictors that were highly correlated with others and had to be dropped from the constraint ordination. After filtering un-correlated and significant plant community predictors, we carried out a variance partitioning per forest to measure the relative contribution of plant phylogeny and traits in explaining soil enzymatic activities from spatial factors (that is, sites within each forest) and soil properties (pH, organic matter –SOM- and soil moisture). Variance partitioning was analysed by using the varpart R function. Un-correlated and significant plant community predictors were further projected within the redundancy analysis plots per forest.
Results
Drivers of Enzymatic Activities Involved in Soil Carbon and Nutrient Cycling
Linear mixed-effect models revealed that forest type was a major factor explaining the variations in carbon and nutrient cycling and enzyme ratios in soils (Table S2). In general, a significantly higher soil enzymatic activity was found in Segura than in Jaén, with the exception of enzymes related to P cycling that did not vary across forests (Figure 1a). The season had a slighter effect than forest type, with a higher N:P enzyme ratio in spring than in autumn (Figure 1b). The models also revealed that plant species were a significant driver of overall C, N and P cycling and enzyme ratios.
Relative soil enzymatic activity (%) per nutrient cycle and cycling ratio depending on a forest (Jaén, Segura) and b season (Autumn, Spring). Relative soil enzymatic activity was calculated as the respective cycle enzymes for a given factor (forest and season) divided by the sum of values of enzymatic activities within that factor. Enzyme values per cycle corresponded to: Labile C=β-glucosidase + cellobiohydrolase; Non-labile C=β-xylosidase + β-glucuronidase + laccase; N = chitinase + leucine-aminopeptidase; P = acid + alkaline phosphatases. Significance is noted next to the bar of each variable, in each graphic, p-value: ***p < 0. 001, **p < 0 .01, *p < 0.05, .p < 0.10, ns = not significant, corrected by the Bonferroni method.
Regarding soil properties (Table S2), pH did markedly depend on the forest but also on the plant species, whereas variations in SOM content were mainly determined by spatial–temporal factors. Both pH and SOM were significantly higher in Jaén (pH 7.59 ± 0.03 and SOM 18.2 ± 1.1%) than Segura (pH 6.20 ± 0.06 and SOM 8.5 ± 0.3%). Soil pH was slightly lower in spring (6.86 ± 0.07) than in autumn (7.01 ± 0.07), whereas the opposite was observed for SOM (14.9 ± 1.1% in spring and 12.6 ± 0.8% in autumn).
When the linkages between enzymes and C soil cycling (that is, hydrolytic or oxidative C enzymes) were explored, a significant positive relationship between N-related and C-hydrolytic enzymes was found (F1335 = 74.57, p < 0.001), while no relation with C-oxidative enzyme was detected (F1335 = 1.09, p = 0.30) (Figure S1). However, significant effects of plant species were further revealed in both C-related enzyme groups (F31307 = 2.30, p < 0.001 for C-hydrolytic enzymes; F31307 = 2.78, p < 0.001 for C-oxidative enzyme).
Phylogenetic Signal of Plant Community on Soil Carbon and Nutrient Cycling
A phylogenetic signal (that is, the tendency of phylogenetically related species to resemble each other) on soil enzymatic activities was found (Table 1; Figure 2). Plant species had a phylogenetic imprinting effect on non-labile C cycling in Jaén, on soil N cycling at both forests (marginal in Jaén), and on P cycling in Segura. A significant plant species phylogenetic signal was also revealed on the C:N enzyme ratio in Jaén, and on the N:P enzyme ratio in Jaén (marginal) and Segura.
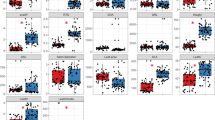
Relative enzymatic activity (%) related to carbon (C), nitrogen (N) and phosphorous (P) cycles of soils collected under the different plant species distributed across the phylogenetic tree (see complete species names in Table S1). Relative soil enzymatic activity was calculated as the respective cycle enzymes for a given plant species divided by the sum of values of that cycle for all plant species. Cycle enzyme values corresponded to: Labile C = β-glucosidase + cellobiohydrolase; Non-labile C = β-xylosidase + β-glucuronidase + laccase; N = chitinase + leucine-aminopeptidase; P = acid + alkaline phosphatases. Further data standardization per plant species within cycles was done relative to the sum of all relative enzyme values of all plant species, to make bars comparable in the graphic. Legend shows the colors associated with each carbon or nutrient cycle. The bar below the phylogenetic tree indicates the adjusted 50 million year-scale.
In Jaén, the Local Moran’s Index revealed hotspots of positive autocorrelation (that is, phylogenetic conservatism) of Pistacia spp. and Lavandula latifolia on soil labile C cycling (Figure S2a), as well as of Quercus spp. and other Lamiales (Rosmarinus officinalis and Thymus mastichina) on N cycling. A similar positive autocorrelation trend was observed in Quercus spp. on non-labile C cycling, C:N and C:P enzyme ratio. In Segura (Figure S2b), significant positive autocorrelations were only detected for Cistus spp. on C and P cycling as well as on C:N enzyme ratio. Negative autocorrelations (that is, phylogenetic divergence) were revealed for the legume shrub Cytisus scoparius, T. mastichina and species belonging to Rosales (Prunus spinosa and Rosa sp.) on N cycling and C:N enzyme ratio, and for and Berberis hispanica also on C:N enzyme ratio.
Plant Trait Predictors of Carbon and Nutrient Cycling in Soils
Both morpho-functional traits and phylogeny (based on PCoA axes) of plant species significantly predicted the enzymatic activity of soils (F9169 = 5.35, p = 0.001, R2adj = 0.16 in Jaén; F10144 = 6.37, p = 0.001, R2adj = 0.29 in Segura) (Table S3). Concerning the phylogenetic composition of plant communities, one PCoA axis per study site was a non-redundant predictor of a total of 20–15 PCoA axes generated at Jaén and Segura, respectively (Figure 4; Table S3). From the total subset of 20 plant morpho-functional traits considered, only a few were non-redundant, that is, leaf habit and mycorrhizal type in Jaén and leaf C, leaf length and mycorrhizal type in Segura.
Variance partitioning (Figure 3) revealed that un-correlated plant community traits were main predictors of soil enzymatic activity variation in both forests (0.15–0.27 of explained variance in Jaén and Segura, respectively), followed by soil properties (0.10 in Jaén and 10.5 in Segura). PCoAs (that is, phylogeny) had a lower contribution in soil enzymatic activity variance (0.04 in Jaén and 0.01 in Segura), and mainly when interacted with plant traits in Jaén and soil properties in Segura. Spatial factors (that is, sites within each forest) did not contribute to explain any variance of soil enzymatic activity (Figure 3).
Amount of enzymatic activity variance explained by the plant community phylogeny, morpho-functional traits, sites and soil properties in the two forest studied, Jaén and Segura. Variance partitioning was carried out with non-redundant plant phylogeny variables (PCoA axes) and traits, the study sites (four within each forest) and soil properties (pH, organic matter –SOM- and gravimetric soil moisture), and plotted in a Venn diagram per forest that shows the relative contribution of predictors in explaining enzymatic activity variations (up to 1). Negative values are not plotted within the diagrams. Residuals were 0.75 and 0.59 in Jaén and Segura, respectively.
In Jaén, the redundancy analysis showed that N-related activities negatively co-varied with PCoA 9 (Figure 4a). This PCoA axis mainly represented phylogenetic distances between C. albidus and Pistacia spp. and, at a lower extent, with species belonging to Rosaceae, Rhamnus lycioides and Ruscus aculeatus (Figure S3a). C and P-related activities did positively co-vary with PCoA 9. Concerning non-redundant morpho-functional traits, non-labile C and P cycling positively co-varied with ECM type whilst N cycling did it with AM type, and labile C did it with perennial plants although at a lower extent than the rest of observed co-variations at this forest (Figure 4a). In Segura (Figuer 4b), N activity positively co-varied with AM type, whilst non-labile C and P cycling positively did it with the plant’s leaf length, leaf C content and with mycorrhizal type (ECM). PCoA 15 showed a low representation within the redundancy analysis, and mainly represented phylogenetic distances among Cistus species (Figure S3b). In plant communities of both forests, plant’s mycorrhizal type was the major trait explaining carbon and nutrient cycling in soils (Figure 4; Table S3).
Plant community phylogenetic and morpho-functional traits related to carbon and nutrient cycling in soils analysed by Redundancy Analysis. Only non-redundant predictors were presented: a in Jaén, one phylogenetic predictor (out of a total of n = 20 PCoA axes) and the morpho-functional traits leaf habit (perennial or deciduous) and mycorrhizal type (arbuscular mycorrhizal –AM– or ectomycorrhizal –ECM–); and b in Segura, four phylogenetic predictors (out of a total of n = 15 PCoA axes), and the morpho-functional traits leaf length (cm), leaf C content and mycorrhizal type (AM, ECM). In the graphic of each forest, soil enzymatic activities related to labile and non-labile C, N and P cycling (green vectors), plant phylogenetic predictors (blue vectors) and morpho-functional traits (orange vectors) are projected, and the direction and length of the vector indicate the effect strength of each variable. See Figure S3 to relate scores of each PCoA axis to plant species.
Discussion
Under the assumption that plant species control the composition of their associated soil microbiota and this, in turn, is key in explaining soil functions (Aponte and others 2013; Pérez-Izquierdo and others 2019), we aimed to closely examine the effect of plant species and their traits in relation to carbon and nutrient cycling in soil. Our results have revealed that plant species act as regulating agents of soil carbon and nutrient cycling by imprinting a phylogenetic signal on soil enzymatic activity. Accordingly, soil carbon and nutrient cycling can be explained by local plant community phylogeny and morpho-functional traits. As expected, all these results are strongly modulated by the local habitat conditions, mainly linked to soil pH, organic matter and moisture. Overall, local plant community phylogenetic traits, together with plant species leaf traits (habit, length and C content) and mycorrhizal type (AM or ECM), are major predictors of soil enzymatic activities in the Mediterranean mixed forests studied.
Drivers of Carbon and Nutrient Cycling
The forest type together with the plant species had a clear effect on soil enzymatic activities in the mixed forest studied, while the season had only a slight effect. Local-context effects have been widely demonstrated at different forest ecosystems (Tedersoo and others 2016), including the Mediterranean basin (Pérez-Izquierdo and others 2017, 2019). Soil properties such as pH and SOM content did notably vary among forests, both being greater in Jaén than in Segura. These soil parameters are recognized effectors of soil carbon and nutrient cycling by conditioning an acidic/alkaline environment and the quantity/quality of substrates, both essential factors to promote extracellular enzymatic activities (Martina and Baldrian 2011; Stock and others 2019). Compared with Jaén, the activities of enzymes related to carbon and nitrogen mobilization were higher in Segura where, in addition, a lower content of SOM was observed. This could indicate an influence of the organic matter properties such as recalcitrance (Theuerl and Buscot 2010; Weintraub and others 2013), that limited resource availability for microorganisms in Segura soils. Independently of the forest, the plant species identity was an important driver of soil carbon and nutrient cycling in these Mediterranean mixed forests, as observed in other forest systems (Aponte and others 2013; Pérez-Izquierdo and others 2017; Haghverdi and Kooch 2019). As Chen and Sinsabaugh (2021) showed, some linkages between N-enzymes and soil C cycling were observed, particularly with hydrolytic C-enzymes, being indicative of soil microbial functionality shifts on the enzymatic activities. Nevertheless, our results revealed a significant effect of the plant species on both hydrolytic and oxidative C-related enzymes, reinforcing that plant community attributes were major driver of soil enzymatic activities.
Plant Community Phylogenetic Signal on Carbon and Nutrient Cycling in Soil
A phylogenetic signal of plant species on soil carbon and nutrient cycling was revealed in the studied mixed forests that is, closely related plant species tended to show similar carbon and nutrient cycling and enzyme ratios. Indeed, our results evidenced that soil enzymatic activity was constrained by the phylogenetic structure of local plant communities. These imprinting on soil nutrient cycling could be explained by the different phylogenetic relationships of the plant species pool and/or by the total number of plant species present in each forest (Münkemüller and others 2012; Davies and others 2013). In particular, phylogenetic conservatism was observed in Quercus spp., Pistacia spp., and species of Lamiales in Jaén and similarly in Cistus spp., C. scoparius and Rosa sp. in Segura. It is noteworthy that these plant taxa diversified in recent times, that is, since the Miocene (~ 25 M.y.). This period is known by suffering numerous climate changes that could have influenced the diversification of several plant lineages in the Mediterranean basin, leading to phylogenetically conserved ecological niches and traits (Grivet and others 2013; Benítez-Benítez and others 2018; Alonso and others 2019). In Segura, phylogenetic divergence was also observed in some clades with respect to the N cycling and C:N enzyme ratio (B. hispanica, C. scoparius, P. spinosa, Rosa sp., T. mastichina). These divergence patterns showed in the outermost species within plant lineages would also support the strong phylogenetic signal detected in the most recent taxa, as pointed out by Alonso and others (2019) on phylogenetically-conserved molecular mechanisms (cytosine methylation). As enzyme ratios are used as proxies of microbial nutrient demand and acquisition strategies (Sinsabaugh and others 2008), the phylogenetic signal detected on ratios of enzyme activities would also indicate a plant phylogenetic conservatism of soil microbial functional groups, particularly in the most recent plant taxa. The differences across forests might be also suggesting an interaction with the local context in order to elucidate these signals. The evolution and co-evolution of plants and soil microbes with resource-use traits underlie when decomposition and production are N-limited (Wooliver and others 2019; Chen and Sinsabaugh 2021). Further investigations involving the role of plant phylogeny and evolutionary history on determining the soil microbiome and its functions would be needed in our studied forests.
Influence of Phylogenetic and Morpho-Functional Traits on Carbon and Nutrient Cycling
Phylogenetic and morpho-functional traits of plant communities did significantly relate to soil carbon and nutrient cycling in the mixed forests studied. The direction of the phylogenetic traits (PCoA axes in redundancy analyses) evidenced a clear influence of the local plant phylogenetic association on soil carbon and nutrient cycling, highlighting the role of the plant community phylogenetic structure on the biogeochemical niche configuration (de Bello and others 2017; Urbina and others 2017). Habit, length and C content of leaves and mycorrhizal type were among the main traits related to soil carbon and nutrient cycling, pointing to distinct organic matter quality and microbial community structure associated with the plant communities of Jaén and Segura. The mycorrhizal type of plants was particularly determinant on the soil enzymatic activities, pointing to the mediating role of ectomycorrhizal (recalcitrant carbon and phosphorous) and arbuscular mycorrhizal (nitrogen) fungi in soil nutrient mobilization. This agrees with the results of Brzostek and others (2015), that showed the magnitude of root-induced changes in decomposition and carbon and nutrient cycling in a temperate forest was partly explained by the mycorrhizal association of the dominant trees.
A predominant influence of leaf traits of local plant species on soil enzymatic activity was also revealed, in particular of leaf habit (perennial vs. deciduous) in Jaén and leaf length and C content in Segura. As happening with belowground inputs, leaves are main contributors of organic matter to soils, and associated traits are well-established predictors of soil enzymatic activity, especially those related to C and N cycling (Poorter and Bongers 2006; Courty and others 2007; Domínguez and others 2012). The positive co-variations among leaf traits, mainly in Segura, with C cycling suggests differences in SOM recalcitrance of the plant species involved (for example, Pinus spp., Cistus spp.) and the mediation of ectomycorrhizal fungi in the decomposition process (as determined by Smith and Wan 2019; Meeds and others 2021). In both forests, N cycling was consistently related to arbuscular mycorrhizal type. Increasing evidence is mounting about the important role of AM symbiosis in N mobilization (Hodge and Fitter 2010; Bukovská and others 2018). Since AM fungal genomes do not contain powerful exo-enzymatic systems capable of mobilizing organic compounds (Tisserant and others 2012; Morin and others 2019), the contrasted carbon and nutrient cycling promotion of ECM versus AM plants that we observed could be reflecting that other microorganisms associated with AM fungi are facilitating this process (Hodge and Fitter 2010; Fernandez and Kennedy 2016; Bukovská and others 2018; Lebreton and others 2021).
Our findings might elucidate underlying mechanisms operating on soil carbon and nutrient cycling, such as the rhizosphere priming effects (Kuzyakov 2010), as we observed that enzymatic activities related to carbon mobilization is highly dependent on organic matter quantity and quality. Decomposition in a high recalcitrant SOM context (that is, ECM plants in our study sites) was preferentially directed to C mobilization. This could be likely performed by dominant ECM fungal communities with great N use efficiency, hence increasing the C:N enzyme ratio, as described in Carteron and others (2021). By contrast, the soil activity under low recalcitrant SOM (that is, AM plants) could allow other decomposer microbial communities the access to SOM sources that promoted N-cycling activities, hence diminishing C:N activity ratio, as reviewed in Fernandez and others (2022). However, the scenario is probably more complex and other important mechanisms could be also operating for example, AM fungal regulation of decomposition rates by favouring soil aggregate formation (Tian and others 2016; Frey 2019), or competition/facilitation between saprotrophic and ECM/AM fungal communities (Fernandez and Kennedy 2016).
Conclusions
Our results reveal the importance of plant community composition to predict soil carbon and nutrient cycling in Mediterranean mixed forests. Indeed, soil carbon and nutrient cycling was explained by de-coupled local plants phylogenetic and morpho-functional traits across the studied forests, highlighting the role of plant community structure on the biogeochemical niche configuration.
The full study system showed that enzymatic activities related to soil carbon and nutrient cycling and enzyme ratios were mainly affected by the forest type (Jaén/Segura) and plant species composing each forest. The variations in the soil enzymatic activities were determined by the local habitat conditions, particularly soil properties such as pH, moisture and organic matter content, and the plant community attributes.
In fact, the phylogenetic approach revealed that soil activity was phylogenetically conserved across forests, indicating the importance of the evolutionary history of plant communities to understand soil carbon and nutrient cycling patterns in Mediterranean mixed forests. Together with plant community phylogeny, leaf traits and mycorrhizal association type of plant species predicted carbon and nutrient cycling in Mediterranean mixed forest soils. In addition, the shifts in soil enzymatic activity among AM and ECM plant species suggest that associated microbial communities have key implications to explain soil carbon and nutrient cycling variations in these forests. Our results open new insights to investigate the role of plant-soil-microbial interactions in explaining nutrient cycling in soils of Mediterranean forest ecosystems.
REFERENCES
Alcántara JM, Pulgar M, Trøjelsgaard K, Garrido JL, Rey PJ. 2018. Stochastic and deterministic effects on interactions between canopy and recruiting species in forest communities. Funct Ecol 32:2264–2274. https://doi.org/10.1111/1365-2435.13140.
Alcántara JM, Garrido JL, Rey PJ. 2019. Plant species abundance and phylogeny explain the structure of recruitment networks. New Phytol 223:366–376. https://doi.org/10.1111/nph.15774.
Allison SD, Vitousek PM. 2005. Responses of extracellular enzymes to simple and complex nutrient inputs. Soil Biol Biochem 37:937–944. https://doi.org/10.1016/j.soilbio.2004.09.014.
Alonso C, Medrano M, Perez R, Canto A, Parra-Tabla V, Herrera CM. 2019. Interspecific variation across angiosperms in global DNA methylation: phylogeny, ecology and plant features in tropical and Mediterranean communities. New Phytol 224:949–960. https://doi.org/10.1111/nph.16046.
Aponte C, García LV, Marañón T. 2013. Tree species effects on nutrient cycling and soil biota: a feedback mechanism favouring species coexistence. For Ecol Manage 309:36–46. https://doi.org/10.1016/j.foreco.2013.05.035.
Averill C, Bhatnagar JM, Dietze MC, Pearse WD, Kivlin SN. 2019. Global imprint of mycorrhizal fungi on whole-plant nutrient economics. Proc Natl Acad Sci USA 116:23163–23168. https://doi.org/10.1073/pnas.1906655116.
Bahram M, Netherway T, Hildebrand F, Pritsch K, Drenkhan R, Loit K, Anslan S, Bork P, Tedersoo L. 2020. Plant nutrient-acquisition strategies drive topsoil microbiome structure and function. New Phytol 227:1189–1199. https://doi.org/10.1111/nph.16598.
Baldrian P. 2017. Forest microbiome: diversity, complexity and dynamics. FEMS Microbiol Rev 41:109–130. https://doi.org/10.1093/femsre/fuw040.
Benítez-Benítez C, Escudero M, Rodríguez-Sánchez F, Martín-Bravo S, Jiménez-Mejías P. 2018. Pliocene-Pleistocene ecological niche evolution shapes the phylogeography of a Mediterranean plant group. Mol Ecol 27:1696–1713. https://doi.org/10.1111/mec.14567.
Brundrett MC, Tedersoo L. 2020. Resolving the mycorrhizal status of important northern hemisphere trees. Plant Soil 454:3–34. https://doi.org/10.1007/s11104-020-04627-9.
Brzostek ER, Dragoni D, Brown ZA, Phillips RP. 2015. Mycorrhizal type determines the magnitude and direction of root-induced changes in decomposition in a temperate forest. New Phytol 206:1274–1282. https://doi.org/10.1111/nph.13303.
Bueno CG, Moora M, Gerz M, Davison J, Öpik M, Pärtel M, Helm A, Ronk A, Kühn I, Zobel M. 2017. Plant mycorrhizal status, but not type, shifts with latitude and elevation in Europe. Glob Chang Biol 26:690–699. https://doi.org/10.1111/geb.12582.
Bukovská P, Bonkowski M, Konvalinková T, Beskid O, Hujslová M, Püschel D, Řezáčová V, Gutiérrez-Núñez MS, Gryndler M, Jansa J. 2018. Utilization of organic nitrogen by arbuscular mycorrhizal fungi: is there a specific role for protists and ammonia oxidizers? Mycorrhiza 28:269–283. https://doi.org/10.1007/s00572-018-0851-y.
Carteron A, Cichonski F, Laliberté E. 2021. Ectomycorrhizal stands accelerate decomposition to a greater extent than arbuscular mycorrhizal stands in a northern deciduous forest. Ecosystems 25(6):1234–1248. https://doi.org/10.1007/s10021-021-00712-x.
Cavender-Bares J, Kozak KH, Fine PVA, Kembel SW. 2009. The merging of community ecology and phylogenetic biology. Ecol Lett 12:693–715. https://doi.org/10.1111/j.1461-0248.2009.01314.x.
Cheeke TE, Phillips RP, Brzostek ER, Rosling A, Bever JD, Fransson P. 2017. Dominant mycorrhizal association of trees alters carbon and nutrient cycling by selecting for microbial groups with distinct enzyme function. New Phytol 214:432–442. https://doi.org/10.1111/nph.14343.
Chen J, Sinsabaugh RL. 2021. Linking microbial functional gene abundance and soil extracellular enzyme activity: Implications for soil carbon dynamics. Glob Change Biol 27:1322–1325. https://doi.org/10.1111/gcb.15506.
Chen J, Luo Y, Xia J, Wilcox KR, Cao J, Zhou X, Jiang L, Niu S, Estera KY, Huang R, Wu F, Hu T, Liang J, Shi Z, Guo J, Wang RW. 2017. Warming effects on ecosystem carbon fluxes are modulated by plant functional types. Ecosystems 20:515–526. https://doi.org/10.1007/s10021-016-0035-6.
Courty PE, Bréda N, Garbaye J. 2007. Relation between oak tree phenology and the secretion of organic matter degrading enzymes by Lactarius quietus ectomycorrhizas before and during bud break. Soil Biol Biochem 39:1655–1663. https://doi.org/10.1016/j.soilbio.2007.01.017.
Davies TJ, Wolkovich EM, Kraft NJ, Salamin N, Allen JM, Ault TR, Betancourt JL, Bolmgren K, Cleland EE, Cook BI, Mazer SJ, McCabe GJ, Pau S, Regetz J, SchwartsTravers MDSE. 2013. Phylogenetic conservatism in plant phenology. J Ecol 101:1520–1530. https://doi.org/10.1111/1365-2745.12154.
De Bello F, Šmilauer P, Diniz-Filho JAF, Carmona CP, Lososová Z, Herben T, Götzenberger L. 2017. Decoupling phylogenetic and functional diversity to reveal hidden signals in community assembly. Methods Ecol Evol 8:1200–1211. https://doi.org/10.1111/2041-210X.12735.
De Groote SR, Vanhellemont M, Baeten L, de Schrijver A, Martel A, Bonte D, Lens L, Verheyen K. 2018. Tree species diversity indirectly affects nutrient cycling through the shrub layer and its high-quality litter. Plant Soil 427:335–350. https://doi.org/10.1007/s11104-018-3654-1.
Domínguez MT, Aponte C, Pérez-Ramos IM, García LV, Villar R, Marañón T. 2012. Relationships between leaf morphological traits, nutrient concentrations and isotopic signatures for Mediterranean woody plant species and communities. Plant Soil 357:407–424. https://doi.org/10.1007/s11104-012-1214-7.
Fernandez CW, Kennedy PG. 2016. Revisiting the “Gadgil effect”: Do interguild fungal interactions control carbon cycling in forest soils? New Phytol 209:1382–1394. https://doi.org/10.1111/nph.13648.
Fernandez M, Vernay A, Henneron L, Adamik L, Malagoli P, Balandier P. 2022. Plant N economics and the extended phenotype: Integrating the functional traits of plants and associated soil biota into plant-plant interactions. J Ecol 110:2015–2032. https://doi.org/10.1111/1365-2745.13934.
Frey SD. 2019. Mycorrhizal Fungi as mediators of soil organic matter dynamics. Annu Rev Ecol Syst 50:237–259. https://doi.org/10.1146/annurev-ecolsys-110617-062331.
Grivet D, Climent J, Zabal-Aguirre M, Neale DB, Vendramin GG, González-Martínez SC. 2013. Adaptive evolution of Mediterranean pines. Mol Phylogenet Evol 68:555–566. https://doi.org/10.1016/j.ympev.2013.03.032.
Haghverdi K, Kooch Y. 2019. Effects of diversity of tree species on nutrient cycling and soil-related processes. Catena 178:335–344. https://doi.org/10.1016/j.catena.2019.03.041.
Hodge A, Fitter AH. 2010. Substantial nitrogen acquisition by arbuscular mycorrhizal fungi from organic material has implications for N cycling. Proc Natl Acad Sci USA 107:13754–13759. https://doi.org/10.1073/pnas.1005874107.
Keck F, Rimet F, Bouchez A, Franc A. 2016. Phylosignal: an R package to measure, test, and explore the phylogenetic signal. Ecol Evol 6:2774–2780. https://doi.org/10.1002/ece3.2051.
Keller AB, Phillips RP. 2019. Leaf litter decay rates differ between mycorrhizal groups in temperate, but not tropical, forests. New Phytol 222:556–564. https://doi.org/10.1111/nph.15524.
Kuzyakov Y. 2010. Priming effects: interactions between living and dead organic matter. Soil Biol Biochem 42:1363–1371. https://doi.org/10.1016/j.soilbio.2010.04.003.
Lebreton A, Zeng Q, Miyauchi S, Kohler A, Dai YC, Martin FM. 2021. Evolution of the mode of nutrition in symbiotic and saprotrophic fungi in forest ecosystems. Annu Rev Ecol Evol Syst 52:385–404. https://doi.org/10.1146/annurev-ecolsys-012021-114902.
LeRoy CJ, Hipp AL, Lueders K, Follstad Shah JJ, Kominoski JS, Ardón M, Dodds WK, Gessner MO, Griffiths NA, Lecerf A, Manning DWP, Sinsabaugh RL, Webster JR. 2020. Plant phylogenetic history explains in-stream decomposition at a global scale. J Ecol 108:17–35. https://doi.org/10.1111/1365-2745.13262.
Liese R, Alings K, Meier IC. 2017. Root branching is a leading root trait of the plant economics spectrum in temperate trees. Front Plant Sci 8:1–12. https://doi.org/10.3389/fpls.2017.00315.
López-Angulo J, Swenson NG, Cavieres LA, Escudero A. 2018. Interactions between abiotic gradients determine functional and phylogenetic diversity patterns in Mediterranean-type climate mountains in the Andes. J Veg Sci 29:245–254. https://doi.org/10.1111/jvs.12607.
Martina Š, Baldrian P. 2011. Effects of soil properties and management on the activity of soil organic matter transforming enzymes and the quantification of soil-bound and free activity. Plant Soil 338:99–110. https://doi.org/10.1007/s11104-010-0296-3.
Meeds JA, Marty Kranabetter J, Zigg I, Dunn D, Miros F, Shipley P, Jones MD. 2021. Phosphorus deficiencies invoke optimal allocation of exoenzymes by ectomycorrhizas. ISME J 15:1478–1489. https://doi.org/10.1038/s41396-020-00864-z.
Midgley MG, Sims RS. 2020. Mycorrhizal association better predicts tree effects on soil than leaf habit. Front for Glob Chang 3:1–14. https://doi.org/10.3389/ffgc.2020.00074.
Morin E, Miyauchi S, San Clemente H, Chen EC, Pelin A, de la Providencia I, Ndikumana S, Beaudet D, Hainaut M, Drula E, Kuo A, Tang N, Roy S, Viala J, Henrissat B, Grigorev IV, Corradi N, Roux C, Martin FM. 2019. Comparative genomics of Rhizophagusirregularis, R. cerebriforme, R. diaphanus and Gigasporarosea highlights specific genetic features in Glomeromycotina. New Phytol 222:1584–1598. https://doi.org/10.1111/nph.15687.
Münkemüller T, Lavergne S, Bzeznik B, Dray S, Jombart T, Schiffers K, Thuiller W. 2012. How to measure and test phylogenetic signal. Methods Ecol Evol 3:743–756. https://doi.org/10.1111/j.2041-210X.2012.00196.x.
Perea AJ, Garrido JL, Alcántara JM. 2021. Plant functional traits involved in the assembly of canopy–recruit interactions. J Veg Sci 32:1–12. https://doi.org/10.1111/jvs.12991.
Pérez-Izquierdo L, Zabal-Aguirre M, Flores-Rentería D, González-Martínez SC, Buée M, Rincón A. 2017. Functional outcomes of fungal community shifts driven by tree genotype and spatial-temporal factors in Mediterranean pine forests. Environ Microbiol 19:1639–1652. https://doi.org/10.1111/1462-2920.13690.
Pérez-Izquierdo L, Saint-André L, Santenoise P, Buée M, Rincón A. 2018. Tree genotype and seasonal effects on soil properties and biogeochemical functioning in Mediterranean pine forests. Eur J Soil Sci 69:1087–1097. https://doi.org/10.1111/ejss.12712.
Pérez-Izquierdo L, Zabal-Aguirre M, González-Martínez SC, Buée M, Verdú M, Rincón A, Goberna M. 2019. Plant intraspecific variation modulates nutrient cycling through its below ground rhizospheric microbiome. J Ecol 107:1594–1605. https://doi.org/10.1111/1365-2745.13202.
Pérez-Valera E, Verdú M, Navarro-Cano JA, Goberna M. 2018. Resilience to fire of phylogenetic diversity across biological domains. Mol Ecol 27:2896–2908. https://doi.org/10.1111/mec.14729.
Phillips RP, Brzostek E, Midgley MG. 2013. The mycorrhizal-associated nutrient economy: a new framework for predicting carbon–nutrient couplings in temperate forests. New Phytol 199:41–51. https://doi.org/10.1111/nph.12221.
Pierre-Emmanuel C, François M, Marc-André S, Myriam D, Stéven C, Fabio Z, Marc B, Claude P, Adrien T, Jean G, Franck R. 2016. Into the functional ecology of ectomycorrhizal communities: environmental filtering of enzymatic activities. J Ecol 104:1585–1598. https://doi.org/10.1111/1365-2745.12633.
Poorter L, Bongers F. 2006. Leaf traits are good predictors of plant performance across 53 rain forest species. Ecology 87:1733–1743. https://doi.org/10.1890/0012-9658(2006)87[1733:LTAGPO]2.0.CO;2.
Pulgar M, Alcántara JM, Rey PJ. 2017. Effects of sampling effort on estimates of the structure of replacement networks. J Veg Sci 28:445–457. https://doi.org/10.1111/jvs.12492.
Purahong W, Durka W, Fischer M, Dommert S, Schöps R, Buscot F, Wubet T. 2016. Tree species, tree genotypes and tree genotypic diversity levels affect microbe-mediated soil ecosystem functions in a subtropical forest. Sci Rep 6:1–11. https://doi.org/10.1038/srep36672.
R Core Team. 2021. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/.
Rundel PW, Arroyo MT, Cowling RM, Keeley JE, Lamont BB, Vargas P. 2016. Mediterranean biomes: evolution of their vegetation, floras, and climate. Annu Rev Ecol Evol Syst 47:383–407. https://doi.org/10.1146/annurev-ecolsys-121415-032330.
Segnitz RM, Russo SE, Davies SJ, Peay KG. 2020. Ectomycorrhizal fungi drive positive phylogenetic plant–soil feedbacks in a regionally dominant tropical plant family. Ecology 101:1–15. https://doi.org/10.1002/ecy.3083.
Senior JK, Potts BM, O’Reilly-Wapstra JM, Bissett A, Wooliver RC, Bailey JK, Glen M, Schweitzer JA. 2018. Phylogenetic trait conservatism predicts patterns of plant-soil feedback. Ecosphere 9(10):e02409. https://doi.org/10.1002/ecs2.2409.
Sinsabaugh RL, Lauber CL, Weintraub MN, Ahmed B, Allison SD, Crenshaw C, Contosta AR, Cusack D, Frey S, Gallo ME, Gartner TB, Hobbie SE, Holland K, Keeler BL, Powers JS, Stursova M, Takacs-Vesbach C, Waldrop MP, Wallenstein MD, Zak DR, Zeglin LH. 2008. Stoichiometry of soil enzyme activity at global scale. Ecol Lett 11:1252–1264. https://doi.org/10.1111/j.1461-0248.2008.01245.x.
Sinsabaugh RL, Hill BH, Follstad Shah JJ. 2009. Ecoenzymatic stoichiometry of microbial organic nutrient acquisition in soil and sediment. Nature 462:795–798. https://doi.org/10.1038/nature08632.
Smith GR, Wan J. 2019. Resource-ratio theory predicts mycorrhizal control of litter decomposition. New Phytol 223:1595–1606. https://doi.org/10.1111/nph.15884.
Stock SC, Köster M, Dippold MA, Nájera F, Matus F, Merino C, Spielvogel S, Gorbushina A, Boy J, Kuzyakov Y. 2019. Environmental drivers and stoichiometric constraints on enzyme activities in soils from rhizosphere to continental scale. Geoderma 337:973–982. https://doi.org/10.1016/j.geoderma.2018.10.030.
Sulman BN, Brzostek ER, Medici C, Shevliakova E, Menge DN, Phillips RP. 2017. Feedbacks between plant N demand and rhizosphere priming depend on type of mycorrhizal association. Ecol Lett 20:1043–1053. https://doi.org/10.1111/ele.12802.
Tedersoo L, Bahram M, Cajthaml T, Põlme S, Hiiesalu I, Anslan S, Harend H, Buegger F, Pritsch K, Koricheva J, Abarenkov K. 2016. Tree diversity and species identity effects on soil fungi, protists and animals are context dependent. ISME J 10:346–362. https://doi.org/10.1038/ismej.2015.116.
Tedersoo L, Bahram M, Zobel M. 2020. How mycorrhizal associations drive plant population and community biology. Science 367:1–9. https://doi.org/10.1126/science.aba1223.
Theuerl S, Buscot F. 2010. Laccases: Toward disentangling their diversity and functions in relation to soil organic matter cycling. Biol Fertil Soils 46:215–225. https://doi.org/10.1007/s00374-010-0440-5.
Tian J, Pausch J, Yu G, Blagodatskaya E, Kuzyakov Y. 2016. Aggregate size and glucose level affect priming sources: a three-source-partitioning study. Soil Biol Biochem 97:199–210. https://doi.org/10.1016/j.soilbio.2016.03.013.
Tisserant E, Kohler A, Dozolme-Seddas P, Balestrini R, Benabdellah K, Colard A, Croll D, da Silva C, Gomez SK, Koul R, Ferrol N, Fiorilli V, Formey D, Franken PH, Helber N, Hijri M, Lanfranco L, Lindquist E, Liu Y, Malbreil M, Morin E, Poulain J, Shapiro H, van Tuinen D, Waschke A, Azcón-Aguilar C, Bécard G, Bonfante P, Harrison MJ, Küster H, Lammers P, Paszkowski U, Requena N, Rensing SA, Roux C, Sanders IR, Shachar-Hill Y, Tuskan G, Young JPW, Gianinazzi-Pearson V, Martin F. 2012. The transcriptome of the arbuscular mycorrhizal fungus Glomus intraradices (DAOM 197198) reveals functional tradeoffs in an obligate symbiont. New Phytol 193:755–769. https://doi.org/10.1111/j.1469-8137.2011.03948.x.
Urbina I, Sardans J, Grau O, Beierkuhnlein C, Jentsch A, Kreyling J, Peñuelas J. 2017. Plant community composition affects the species biogeochemical niche. Ecosphere 8(5):e01801. https://doi.org/10.1002/ecs2.1801.
Wang D, Chen J, Felton AJ, Xia L, Zhang Y, Luo Y, Cheng X, Cao J. 2021. Post-fire co-stimulation of gross primary production and ecosystem respiration in a meadow grassland on the Tibetan Plateau. Agric for Meteorol 303:108388. https://doi.org/10.1016/j.agrformet.2021.108388.
Waring BG, Weintraub SR, Sinsabaugh RL. 2014. Ecoenzymatic stoichiometry of microbial nutrient acquisition in tropical soils. Biogeochemistry 117:101–113. https://doi.org/10.1007/s10533-013-9849-x.
Webb CO, Ackerly DD, McPeek MA, Donoghue MJ. 2002. Phylogenies and community ecology. Annu Rev Ecol Syst 33:475–505. https://doi.org/10.1146/annurev.ecolsys.33.010802.150448.
Weintraub SR, Wieder WR, Cleveland CC, Townsend AR. 2013. Organic matter inputs shift soil enzyme activity and allocation patterns in a wet tropical forest. Biogeochemistry 114:313–326. https://doi.org/10.1007/s10533-012-9812-2.
Wooliver R, Pellegrini AF, Waring B, Houlton BZ, Averill C, Schimel J, Hedin LO, Bailey JK, Schweitzer JA. 2019. Changing perspectives on terrestrial nitrogen cycling: the importance of weathering and evolved resource-use traits for understanding ecosystem responses to global change. Funct Ecol 33:1818–1829. https://doi.org/10.1111/1365-2435.13377.
ACKNOWLEDGEMENTS
We gratefully acknowledge L. Pérez-Izquierdo, L. Pomarède, R. Gómez and L. López for their help. This work was supported by the project COEXMED-II (CGL2015-69118-C2-1P and CGL2015-69118-C2-2P) founded by the Spanish State Research Agency (AEI). J.P.R. and A.P. held pre-doctoral fellowships awarded by AEI (BES-2016-078055 and BES-2016-077688, respectively).
Funding
Open Access funding provided thanks to the CRUE-CSIC agreement with Springer Nature.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Author Contributions
JMA, CA, AR designed the research; JPR, AP, JLG, JMA, CA, AR performed sampling, JPR did laboratory work, JPR, ALG, AR analysed data; JPR wrote the manuscript. All authors revised the manuscript.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Prieto-Rubio, J., Perea, A., Garrido, J.L. et al. Plant Traits and Phylogeny Predict Soil Carbon and Nutrient Cycling in Mediterranean Mixed Forests. Ecosystems 26, 1047–1060 (2023). https://doi.org/10.1007/s10021-022-00815-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10021-022-00815-z