Abstract
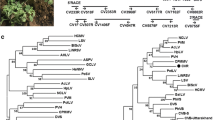
The complete genome sequence of a putative novel member of the genus Sadwavirus was determined by high-throughput sequencing of a chrysanthemum from an orchard of the Tongxiang Agricultural Science Institute in Tongxiang, Zhejiang province. The complete genome sequence was confirmed using RT-PCR and the rapid amplification of cDNA ends (RACE) method. The predicted genome of the putative virus is composed of two RNA molecules, 7016 and 6772 nucleotides in length, excluding their poly-A tails. The new virus was tentatively named “chrysanthemum sadwavirus” (ChSV). The Pro-Pol region of RNA1 and the CP region of RNA2 of ChSV shared the highest amino acid sequence identity (53.01% and 36.40%, respectively) with the corresponding sequences of lettuce secovirus 1 (LSV-1). Phylogenetic analysis showed that ChSV clustered with members of the subgenus Stramovirus (genus Sadwavirus). Taken together, these results suggest that ChSV is a new member of the genus Sadwavirus.


Similar content being viewed by others
Data availability
All data included in this study are available upon request by contact with the corresponding author.
References
Liu H, Luo C, Chen D et al (2021) Whole-transcriptome analysis of differentially expressed genes in the mutant and normal capitula of Chrysanthemum morifolium. BMC Genom Data 22:2
Qu L, Ruan JY, Jin LJ et al (2017) Xanthine oxidase inhibitory effects of the constituents of Chrysanthemum morifolium stems. Phytochem Lett 19:39–45
Zhong X, Yang L, Li J et al (2022) Integrated next-generation sequencing and comparative transcriptomic analysis of leaves provides novel insights into the ethylene pathway of Chrysanthemum morifolium in response to a Chinese isolate of chrysanthemum virus B. Virol J 19:182
Li J, Wu X, Liu H et al (2023) Identification and molecular characterization of a novel carlavirus infecting chrysanthemum morifolium in China. Viruses 15:1029
Gong J, Chu B, Gong L et al (2019) Comparison of phenolic compounds and the antioxidant activities of fifteen Chrysanthemum morifolium Ramat cv. ‘Hangbaiju’ in China. Antioxidants 8: 325
Fuchs M, Hily JM, Petrzik K et al (2022) ICTV Report Consortium (2022): ICTV Virus Taxonomy Profile: Secoviridae 2022. J Gen Virol 103:001807
Funding
This work was supported by the Regional Demonstration Project of the Alliance of Municipal Academies of Agricultural Sciences in Zhejiang Province, grant number 2022SJLM12.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
This article does not contain any experiments involving humans or animals, and no ethical approval was required.
Additional information
Handling Editor: Jesús Navas-Castillo.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, J., Dong, Y., Wang, H. et al. Identification and complete genome sequence of a novel sadwavirus discovered in chrysanthemum (Chrysanthemum morifolium Ramat.). Arch Virol 168, 295 (2023). https://doi.org/10.1007/s00705-023-05916-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00705-023-05916-1