Abstract
Immunoassays are a powerful diagnostic tool and are widely used for the quantification of proteins and biomolecules in medical diagnosis and research. Enzyme-linked immunosorbent assay (ELISA) is the most commonly used immunoassay format and allows the detection of biomarkers at a very low concentration. The diagnostic platforms such as enzyme immunoassay (EIA), chemiluminescence (CL) assay, polymerase chain reaction (PCR), flow cytometry (FC), and mass spectrometry (MS) have been used to identify molecular biomarkers. However, these diagnostic tools requiring expensive equipment, long testing time, and qualified personnel that is not always available in small local hospitals with limited resources. The lateral flow immunoassay (LFIA) platform was developed for rapidly obtaining laboratory results and to make urgent decisions in emergency medicine, as well as the recently introduced concept of testing at the site of care (point-of-care, POC). The simultaneous measurement of different substances from a single sample called multiplex assays have become increasingly significant for in vitro quantification of multiple analytes in a single sample, thereby minimising cost, time, and volume. In multiplex immunoassays, the ligands are immobilized either in planar format (flat surface) or on microspheres in suspension that binds to target analytes in sample. The multiplex technology has established itself in proteomic networks and pathways, validation of genomic discoveries, and in the development of clinical biomarkers. In the present review article, various types of monoplex/simplex and complex/multiplex immunoassays have been analysed that are increasingly being applied in laboratory medicine. Also, some advantages and disadvantages of these multiplex assays have also been included such as experimental animals, in vitro tests using cell lines and tissue samples, 3-dimensional modelling and bioprinting, in silico tests, organ-on-chip, and computer modelling.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Laboratory measurements have become an integral part of medical diagnosis and therapy which allows the evaluation of pathology and treatment of a patient. The laboratory must validate the technical standard and reproducibility of measurement through standard methodology and adequate quality management. The doctor/physician should select the right pathological parameters to be analysed and interpretation of laboratory results in patient care. During the last two decades, laboratory diagnostics and bioassays have undergone a technological revolution (Elmlinger 2011). An early and accurate diagnosis of a disease plays an important role in its effective treatment, especially in emergency where an immediate decision needs to be made and the rapid and precise identification of the pathology is vital. However, the clinical evidence based on a single analyte or biomarker is not adequate for an appropriate diagnosis of a disease or monitoring its treatment. The potential biomarkers have a pathophysiological significance and the clinical applications may have a profound impact on disease diagnosis and therapy (Ahsan and Ahmad 2020; Ahmad and Ahsan 2014; Ahsan 2013). The various biomarkers are an integral part of biomedical research and clinical practice. Many clinical tests and assays such as the lipid profile, blood pressure, body temperature, or pulmonary function test (PFT) serve as clinical biomarkers and the body mass index (BMI) measurement as a risk biomarker which is used to assess the risk for metabolic diseases such as diabetes mellitus, hypertension, and dyslipidemia, and may lead to preventative health measures (Bays et al. 2007). Biomarkers can also be safety indicators for determining health status, such as liver and kidney function (LFT, KFT) by measuring serum creatinine, uric acid, urea nitrogen, cystatin C, etc. (Kar et al. 2018; Bonventre et al. 2010; Babrak et al. 2019).
In clinical trials, the well-defined traditional biomarkers are important and accepted metrics to assess the clinical status and therapeutic effects for clinical endpoints and adverse events (Agency and Committee for Medicinal Products Human Use (CHMP), 2018). A biomarker is defined as “a characteristic that is measured as an indicator of normal biological processes, pathogenic processes, or responses to an exposure or intervention including therapeutic interventions” (Babrak et al. 2019; FDA-NIH Biomarker Working Group 2016). The FDA-NIH Joint Leadership Council has put together the Biomarkers, Endpoints and other Tools Resource (BEST) for coordinating translational science by regularizing medical terms and improve scientific understanding regarding biomarkers in clinical sciences and practice (Babrak et al. 2019; FDA-NIH Biomarker Working Group 2016). The FDA-NIH Biomarker Working Group defined two categories of biomarkers: (i) disease-associated biomarkers (susceptibility/risk, diagnostic, prognostic, monitoring biomarkers) and (ii) drug-related biomarkers (predictive, pharmacodynamics/response, safety biomarkers).
The conventional laboratory-based methods like high-performance liquid chromatography (HPLC), gas chromatography (GC), mass spectrometry (MS), enzyme-linked immunosorbent assay (ELISA), and polymerase chain reaction (PCR) require complex procedures to obtain results, but several situations often require fast and on-site analyte detection. As a consequence, the scientific research has focused on the development and optimization of portable, affordable, and user-friendly rapid methods of analysis for the point-of-care (POC) testing (Nardo et al. 2021). The microbiological, immunological, and diagnostic methods such as enzyme immunoassay (EIA), chemiluminescence assay (CIA), polymerase chain reaction (PCR), flow cytometry (FC), and mass spectrometry (MS) have been used to accurately identify molecular markers (Andryukov 2020). The contemporary monoplex/simplex techniques such as EIA and ELISA are able to characterize a single analyte for the accurate diagnosis of diseases. However, the monitoring of more complex, multifactorial diseases such as cancer, autoimmunity/immunodeficiency, and neurodegenerative diseases requires the analyses of multiple biomarkers in order to implement optimized therapeutic regimens (Tighe et al. 2015). In addition, it is advantageous to screen different analytes simultaneously, enabling a rapid, low-cost, and reliable quantification of samples (Ahsan 2019; Dincer et al. 2017). The development of technologies for the analysis of genome (genomic), transcriptome (transciptomic), proteins (proteomic), and metabolome (metabolomic) has ushered in a new era of analysis and discovery, which has yielded novel biomarkers (Ahsan 2019).
The immunochemical bioanalytical methods represent one of the most versatile strategies for point-of-need (PON) applications due to their ability to provide rapid and specific results. Therefore, most screening and rapid laboratory assays and methods are based on immunoassays. An immunoassay is a biochemical test that is commonly used to measure the concentration of a target molecule. This method is based on the reaction of an analyte/antigen (Ag) with a selective antibody (Ab) forming an Ab-Ag complex. The efficacy of immunoassays is mainly based on the efficiency of Ab-Ag complex formation and the ability to detect the rate of immunocomplex formation (Nardo et al. 2021).
Monoplex immunoassays
Immunoassays (IAs) allow the sensitive and specific detection of various analytes in complex biological samples and are widely used in hospitals, laboratories and research for the diagnosis of diseases and drug development. Since their introduction in 1960s, the radioimmunoassay (RIA) or in the 1970s, ELISA have become an indispensable analytical tools in a wide range of clinical diagnostic applications leading to better therapeutic choices. It is also widely used in industrial and analytical quality control such as the detection of contaminants in food and water and monitoring specific molecules during the processing of foods (Pal 2015). IAs have many applications in diverse scientific fields, e.g., agriculture and environment, veterinary science, clinical medicine, food and nutrition, and molecular biology. It is useful for the analysis of compounds such as proteins, peptides, microorganisms, toxins, hormones, antibiotics, vitamins, pesticides, metal ions, and nucleotides. IAs have been widely used for the quantification of proteins and small molecules in medical diagnostics, proteomics, drug discovery, and biomedical research (Table 1). The IAs are supported by improved labelling and detection techniques (radionuclides, enzymes, biotin/streptavidine, dyes, fluorophores, chromophores, etc.) and methods for different phase separation (adsorption techniques and support material e.g., microplates, coated tubes, beads) (Elmlinger 2011; Boguszewska et al. 2019).
The enzyme immunoassays (EIAs) are the most common immunoanalytical assays that determine the concentration of antigens or antibodies in a complex mixture in which the activity of the enzyme is detected. The use of enzymes is advantageous since it uses a variety of substrates such as hydrogen peroxide (H2O2) that form the coloured compound of chromogen leading to the generation of signals (Boguszewska et al. 2019). The enzymes horse radish peroxidase (HRP) and alkaline phosphatase (ALP) are the most commonly used enzymes in IAs (Grange et al. 2014). Enzymes are attached to the antibody or antigen by means of glutaraldehyde, sodium periodate, or enzyme-antienzyme complex (e.g., peroxidase-antiperoxidase, PAP; alkaline phosphatase-alkaline antiphosphatase, APAAP). The IA methods involve labelling the antigen or antibody, depending on the assay format, through direct conjugation with an enzyme, fluorophore, or chromophore or indirect conjugation with biotin (vitamin B7) or avidin/streptavidin (proteins). Small antigens often undergo steric changes through conjugation with enzymes; therefore, these antigens can be conjugated with a spacer such as biotin, which couples the enzyme by using the linkers, avidin (egg protein) and streptavidin (Staphylococcus aureus protein). The egg protein avidin and the bacterial streptavidin have a very high affinity for biotin (Elmlinger 2011). The bound enzyme metabolizes the substrate depending upon the reaction time resulting in the amplification of signal (Elmlinger 2011).
Enzyme-linked immunosorbent assay
Eva Engvall and Peter Perlmann (Engvall and Perlmann 1971) described a technique in where the antigens were immobilized onto a microplate well, incubated with diluted antiserum, and then, the concentration of antibody in the antiserum was quantified using an enzyme-linked anti-immunoglobulin antibody (Engvall and Perlmann 1971) known as the enzyme-linked immunosorbent assay (ELISA). The enzyme immunoassay (EIA) was developed by Bauke Klass van Weemen and Antonius Hermanus Wilhelmus Maria Schuurs (Weemen and Schuurs 1971) for the quantification of antigen rather than antibody and is similar to ELISA (Grange et al. 2014). The EIA allows the detection of very small quantities of antigens such as protein, peptide, hormone, and antibody in a sample. They utilize the enzyme-labelled (alkaline phosphatase, glucose oxidase) antigens and antibodies to detect the biological molecules (biomolecules). The antigen is allowed to bind to a specific antibody, which is subsequently detected by a secondary enzyme-coupled antibody. A chromogenic substrate for the enzyme yields a visible colour or fluorescence indicating the presence of an antigen using the method of colorimetry, also known as filter photometry. The fluorogenic substrates have higher sensitivity and can accurately measure the antigen concentration in the sample. The key step in ELISA is the direct or indirect detection of an antigen by immobilizing the antigen or antigen-specific capture antibody, respectively (Fig. 1). The antigen can be specifically selected from a sample mixture through a “capture” antibody. The antigen is thus “sandwiched” between the capture and a detection antibody. If the antigen is small in size or has only one epitope (antigenic determinant) for antibody binding, a competitive method is used for labelling either the antigen which competes for the unlabeled antigen–antibody complex or the antibody that competes for the free or bound antigen in a sample and can be used for the qualitative and quantitative estimation (Gan and Patel 2013).
The different types of enzyme-linked immunosorbent assays (ELISAs): a direct, b indirect, c sandwich, and d competitive (reproduced from (Boguszewska et al. 2019))
ELISA is a solid-phase test that uses a colour reaction of the enzyme coupled to the antibody and is performed on microtitre plates where the antigen or antibody is adsorbed onto the walls of the plate. The technique is based on the immobilization of the antigen on the surface of the solid phase and the addition of sample that contains antibodies specific for the antigen, covalently linked to the enzyme. The antigen forms an immune complex with the antibody in which the antibody is also bound to the substrate. After adding the substrate, the enzyme catalyses the reaction resulting in a coloured product. The concentration of the product is determined spectrophotometrically through an ELISA reader allowing quantitative analysis. The amount of product formed corresponds to the concentration of the antibody-antigen complex which is calculated based on a standard curve determined using specific standard. The quantity of a product, equivalent to the concentration of the antibody-antigen complex, is depended on the amount of the given substance in the sample. Therefore, there are several types of methods which are known as direct, indirect, sandwich, and competitive ELISAs (Boguszewska et al. 2019; González-Martínez et al. 2018; Slage and Immunoassays 1996; Wu 2006; McCarthy 2003; David 2005; Li et al. 2018; Cox et al. 2004; Davies 2013) (Fig. 1).
In laboratory diagnostics, an indirect ELISA is most commonly employed in which two types of antibodies are used, a primary antibody, which recognizes a given antigen and a secondary antibody, which is labelled and recognizes the primary antibody. The standard ELISA technique does not show high sensitivity and, therefore, a “sandwich”-type ELISA was introduced in which the antigen is not immobilized directly onto the substrate but is bound to the coat antibody on the microplate (Fig. 1). This assay is used to detect the amount of the test protein in a sample and diagnostic tests e.g., to determine the titre of antibodies in the blood. The oxidative DNA damage products e.g., thymine glycol (Tg, 5,6-dihydro-5,6-dihydroxythymine), and 8-hydroxyguanine (8-OHGuo, 8-oxo-Gua) may also be detected through the commercially available kits (Boguszewska et al. 2019). The ELISA test is one of the most popular and common techniques due to a number of advantages such as versatility, high sensitivity and selectivity, specificity of reaction, relatively short analysis time, simplicity of execution, and low cost making it practicable and economical. However, there are also some disadvantages e.g., non-availability of specific antibodies and the reduced specificity of antibodies labelled with enzymes which have led to the improvements and introduction of more efficient methods and techniques (Boguszewska et al. 2019).
Radio immunoassay
The first radioactive IA was developed by Rosalyn Yalow and Soloman Berson in 1959 (Yalow and Berson 1959). They used radiolabelled insulin to measure the concentration of insulin in human plasma which led to the development of RIA. Therefore, RIA replaced the existing bioassays and precipitation tests (radial double diffusion/radial immunodiffusion, RID; immunoelectrophoresis, IEP; electroimmunoassay/electroimmunodiffusion/rocket immunoelectrophoresis, RIEP). RIA is a quantitative method that uses radioactive isotopes for the detection of antigen–antibody interaction through direct or indirect measurement of the unlabeled substance binding to a specific antibody. The higher the radioactivity of the sample, the higher the concentration of the substance present. There are a number of isotopes that are employed in RIA such as 125I (radioactive iodine), 32P (radioactive phosphorus), 14C (radioactive carbon), and 3H (radioactive tritium). Depending on the radioactive compound, β (low penetration) or γ (high penetration) radiation is emitted (Annesley 2010; Chard 1996; Goldsmith 1975), which is detected by a gamma counter. RIA is employed in many diagnostic tests due to its sensitivity and simplicity and consists of three stages: immunoreaction (antigen/antibody binding), competitive binding and competitive displacement reaction, which gives specificity and measurement of radioemission providing high sensitivity (0.0006–0.006 µg antibody/ml). The RIA is a robust and inexpensive technique but is limited by the half-life of the radioactive material, hazardous waste disposal, specialized instruments, and trained laboratory personnel. For decades, the technique was mainly applied for diagnostic purposes to analyse an antibody or antigen for the diagnosis of a disease or pathological condition (Haschek et al. 2010; Preedy et al. 2009; Berton and Mitchell 2012). Nowadays, it is also used in screening for immunity, allergens in the food industry, or molecular biology laboratories. A few variants of the RIA method are also employed that enlarge the range of applications such as immunoradiometric assay (IRMA), radioallergosorbent test (RAST), and Farr assay (dsDNA-Ab RIA) which are employed for the detection of anti-double stranded DNA (anti-dsDNA) antibodies in autoimmune diseases (Boguszewska et al. 2019; Smeenk et al. 1991, 1990; Derksen et al. 2002; Fadal 1992; Miles 1975).
Another commonly used IA is electrochemiluminescence (ECL) which is widely used in large automated immunoassay platforms. It is based on the oxidation of a tripropylamine (TPA) and ruthenium-II (Rhu(II)) chelate label of the antigen at the anode. An excited state of Ru(II) intermediate product is generated, which emits light that is detected and quantified. The IA is carried out by causing the ruthenium label to bind to antibodies on suspended or magnetic beads (Elmlinger 2011). Moreover, IAs that does not require the use of enzymes and radionuclides are now being developed. These assays include competition assays using fluorescent peptides and a variety of labelled streptavidin compounds for use with biotinylated antibodies or peptides. The enzyme multiplied immunoassay technique (EMIT) requires an enzyme-linked antigen that competes with the sample antigen for antibody binding, and the enzyme is deactivated by antibody binding. In the fluorescence polarization immunoassay (FPIA), the bound and unbound fluorescein-conjugated antigens emit fluorescence that can be detected (Grange et al. 2014).
Lateral flow immunoassay
Among the immunoassay-based analytical platforms, the lateral flow immunoassay (LFIA), also known as immunochromatographic tests, has been successfully used in diagnosis of many diseases and conditions as they allow rapid detection of molecular ligands in bioanalytical samples (Table 2). The popularity of these diagnostic platforms is constantly increasing in healthcare facilities, particularly those facing limited budgets and time, as well as in household use for individual health monitoring. The advantages of these low-cost devices over modern laboratory-based analysers is their availability, rapid detection, and ease of use. The effectiveness of these portable diagnostic tools is primarily associated with their high analytical sensitivity and specificity and ease of visual analysis of results. The LFIA is becoming increasingly popular in developing countries, when applied at small hospitals, in emergency situations where screening and monitoring health is crucial and for individual self-testing (Andryukov 2020).
The principle of diagnostics based on paper chromatography (PC) was first proposed by R. Yalow and S. Berson (Yalow and Berson 1960) using paraffin/wax paper to detect insulin in human blood and was named as sol particle immunoassay (SPIA) or lateral flow immunoassay (LFIA). The PC method was later replaced by nitrocellulose, and the range of laboratory diagnostics was extended for determining other blood analytes (hormones, enzymes, vitamins, and markers of infectious diseases). As the technology developed, the range of its applications expanded to infectious diseases, cardiovascular diseases, cancer biomarkers, food pathogens, and veterinary diagnostics. Several variants of LFIA have been proposed that are more sensitive, affordable, and user friendly allowing their use not only by laboratory staff, but also by other medical specialists and patients for self-monitoring of their health (Andryukov 2020). The current generation of LFIAs have high sensitivity and selectivity which is achieved by combining thin-layer chromatography (TLC), analyte-specific antibodies, DNA/RNA-specific sequences, and analyte labelling (Posthuma-Trumpie et al. 2009). Therefore, when antibodies are exclusively used as recognition elements, it is known as LFIA. However, a combination of antigen–antibody interaction and the detection of specific doubled-stranded amplicon (ds-amplicon) after PCR is known as nucleic acid lateral flow immunoassay (NALFIA) and that of specific nucleic acid hybridization of amplicons with immobilized complementary probes is known as nucleic acid lateral flow assay (NALF) (Posthuma-Trumpie et al. 2009).
The LFIA, also known as immunochromatographic strip test (ICST), or rapid diagnostic test (RDT), has become one of the most successful analytical platforms for PON testing requiring little or minimum infrastructure. The LFIA is a paper-based bioanalytical technique for the on-site detection of targets in a few minutes. LFIAs satisfied all the criteria of an ideal POC testing that is required to be “ASSURED” (Affordable, Sensitive, Specific, User-Friendly, Rapid and Robust, Equipment-Free and Delivered) (Kettler et al. 2004), and has become the benchmark for POC/PON testing (Nardo et al. 2021). The LFIA can be considered as a lab-in-a-hand and together with other PON tests is an example of sample-to-lab to lab-to-sample for improving the decision making and turnaround time. The LFIA application has been rapidly extended to the detection of hormones, parasites, bacteria, cells, viruses, biological markers for clinical purposes to include food and feed safety, veterinary medicine, environmental control, forensic analysis, etc. Usually, for both the competitive and the noncompetitive format, the detection limit of LFIA is between 0.1 and 10 ng/mL. The analytical sensitivity and versatility have made LFIA the most commercially available POC diagnostic platform (Nardo et al. 2021; Andryukov 2020). The advancement in LFIA technology is the development of multiplex systems or format which allows the detection of various bacterial or viral antigens in a single assay (Nardo et al. 2021; Andryukov 2020; Jørgensen et al. 2015) (Table 2). The wide acceptance of LFIA platform has also played a prominent role in the detection of the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) (Espejo et al. 2020; Ruhan et al. 2020; Zhu and Woong 2020; Deeks et al. 2020; Dinnes et al. 2020; Vashist 2020; Mekonnen et al. 2020; Kailasa et al. 2021; Antiochia 2021; Ernst et al. 2021) (Fig. 2).
The different diagnostic assays for the detection of coronavirus, SARS-CoV-2. Abbreviations: NAAT nucleic acid amplification test, ELISA enzyme-linked immunosorbent assay, CT computed tomography, LFIA lateral flow immunochromatographic assays, NGS next-generation sequencing (reproduced from (Oishee et al. 2021))
Multiplex immunoassays
There is an increasing demand for technologies that are capable of extracting large amount of bioinformation from a limited sample volume for better diagnosis, prognosis, and treatment of diseases. This has led to the development of low-cost, flexible, and high throughput methods for simultaneous detection of multiple proteins. The development of multiplex immunoassays allow the simultaneous measurement of multiple analytes in a single biological sample with minimal assay time, cost, and sample volume (Ahsan and Ahmad 2020). After overcoming the technical hurdles, the multiplex platform-based technologies have been applied in biomedical research and clinical diagnostics (Zheng and He 2012). The so called multiplex technology could be useful in the biopharmaceutical industry to rapidly identify and screen various preparations which may be helpful in the identification and quantification of biomarkers in pharmaceutical product and epidemiological studies (Ahsan 2019).
The concept of multiplex immunoassay was described by Feinberg in 1961 for “microspot” test of antibody-antigen reaction in thin agar films and first demonstrated by Roger Ekins (Ekins 1989; Feinberg 1961). In 1989, Ekins described the principles of microarray technology and proposed that a tiny spot of a purified antibody provides better sensitivity rather than a larger quantity. This technique has immense potential applications in biomedical research and clinical diagnostics and a large number of platforms have been developed with the advancement in technology (e.g., fluidics, optics, automated sample handling device) and bioinformatics leading to a high-throughput multiplex immunoassays (Hsu et al. 2009; Krishhan et al. 2009; Nolan and Mandy 2006). Therefore, multiplex technology provides high analytical accuracy and saves time, material, and labour cost allowing efficient handling of a large number of samples over conventional ELISAs (Tozzoli et al. 2015). Although there are still some challenges (e.g., complexity, cost, validation), multiplex protein test panels are now being increasingly used in clinical diagnostics (Rajagopal et al. 2013). Several commercial multiplex immunoassay platforms are available in the market, including Luminex bead–based platforms, Multiarray Technology from Meso Scale Discovery, and protein microarray platforms, etc. (Zheng and He 2012) (Table 3).
An ideal device for emergency testing should offer high performance, sensitivity multiplexing capability, short turnaround time, less system complexity including low cost for analysis and maintenance and minimum user intervention (Dincer et al. 2017). The multiplex diagnostic device capable of high-throughput analysis of several parameters has become increasing significant in the last few decade that are able to characterize various biomarkers simultaneously e.g., RNAs, metabolites, proteins, and cells (Dincer et al. 2017). Multiplex immunoassay confers several advantages over simple IAs including increased efficiency, greater output per sample volume, and higher throughput and detailed diagnostics. Most of the diagnostic methods rely on IAs or enzymatic reactions and depend on the sample (e.g., matrix effects, patient-to-patient variations) and environment (e.g., temperature, humidity). The emerging requirements for novel biomarkers (e.g., aptamers) or targets (e.g., RNAs, tumour cells, exosomes, miRNA) and their applications for diagnostic, prognostic, and therapeutic implications will form the basis of multiplex formats in future (Dincer et al. 2017; Rajagopal et al. 2013).
Multiplex technology may also be useful for the simultaneous detection of different autoantibodies in various autoimmune diseases (AID) (Tozzoli et al. 2015). Multiplex assays can detect specific autoantibodies in a single run, whereas a traditional ELISA uses a single antigen to detect only a single specificity of autoantibodies. Thus, in multiplex assays, a combination of recombinant or native antigens or antigenic peptide is used to detect autoantibodies. The classical IAs such as double immunodiffusion (DID) or immunoprecipitation (IP) can detect specific autoantibodies in a single run (Satoh et al. 2015). The different types of multiplex assays include line immunoassay (LIA), multiplex bead assay (MBA) and solid-phase antigen microarray. LIA is similar to the dot/slot blot or western blot immunoassay in which the diluted serum is incubated with several antigens bound onto a strip. In MBA, beads of different sizes and/or fluorochromes are coated with specific antigen and mixed to allow detection of a particular autoantibody. In antigen microarrays, the antigens are coated onto a slide or membrane. The strips, beads, slide/membrane, antigen mixture are incubated with a dilute serum, and many autoantibodies can be detected (Satoh et al. 2015).
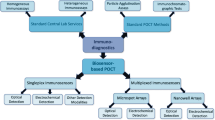
The multiplex immunoassays employ traditional IA methods in which either antibodies or proteins/peptides are used as binder molecules to capture circulating proteins or antibodies, respectively. Basically, there are two types of multiplex immunoassays: (a) planar microarray (protein chips) and (b) suspension array (microparticle or bead microarrays) (Boguszewska et al. 2019; Zheng and He 2012) (Fig. 3). The protein chip assays are miniaturized and contain small amounts of purified proteins in a high-density format and allow the simultaneous determination of a large number of analytes from a small amount of sample in a single experiment. These assays are prepared by immobilizing proteins onto a microscope slide and then probed for a variety of functions and the resulting signals are detected using a fluorescent or radioisotopic label (Pal 2015). The planar arrays are of two types, either the slide or microtitre-based format. In planar microassays, the capture ligands are immobilized on a rigid two-dimensional support, probed with sample and the fluorescent or chemiluminescent signals detected. In the slide-based format, the repeated or individual assays composed of specific sets of antibodies are printed robotically on the activated slide surface. The sample matrix is applied, and discrete assays are spatially separated and considered as individual microtitre wells. The microtitre-based assay consists of antibodies bound onto the wells of a standard and conventional protein-binding ELISA plate (Ellington et al. 2010). The application of a specific secondary antibody coupled to a chemiluminescent or fluorescent reporter results in the detection of signal.
The multiplex immunoassays: A planar microarray (protein chips) and B suspension array (microparticle or bead microarrays) (reprinted from (Tighe et al. 2015)). These multiplex immunoassays use serum samples for the analysis of various biomarkers. A Planar arrays can be produced in two formats - slide or microtitre. (i) The slide‐based format support numerous layouts whereby repeated or individual assays composed of specific sets of antibodies are printed robotically upon the activated slide surface. (ii) The sample matrix is applied and discrete assays are spatially separated and treated as individual microtitre wells. (iii) The detection is through the application of specific secondary antibodies coupled with a chemiluminescent/fluorescent reporter systems. (iv) Microtitre‐based immunoassay harbour regularly printed antibody sets within the wells of a standard (SBS format) protein‐binding plate. (v) The plate may thus be treated as in a conventional ELISA (i.e. blocking, incubation and washing followed by detection with a set of reporter‐conjugated detection antibodies). B (i) The suspension assay employs thousands of micrometre‐sized plastic microbeads infused with a single (or several) chemiluminescent/fluorescent dyes and a functionally activated surface, prior to linking with a specific capture antibody. (ii) Numerous sets of such beads are prepared, each maintaining separate capture antibodies according to the analyte and a unique fluorescent signature enabling identification. (iii) The detection antibodies, which are individually labelled with a single chemiluminescent/fluorescent reporter (separate from those contained within the beads) are added upon completion of incubation and washing stages. (iv) Each bead thus accommodates a ‘sandwich’ consisting of the captured target analyte and the reporter‐conjugated detection antibody. (v) The bead analyte reporter constructs are analyzed in a flow chamber enabling bead separation in which lasers excite the chemiluminescent/fluorescent reporters and emitted light is collected by a series of detectors for quantitative analysis
In suspension assays, the capture ligands are immobilized onto colour or size-coded microspheres and specific fluorescent signals are detected using flow cytometry (Ellington et al. 2010). The suspension assay employs plastic microbeads infused with a chemiluminescent/fluorescent dye and an activated surface linking it to a specific capture antibody. The detection antibodies with chemiluminescent/fluorescent reporter are added upon completion of incubation and washing stages. Several beads are prepared, each with a separate capture antibody according to the analyte and a unique fluorescent signature that enables identification. Each bead accommodates a “sandwich” consisting of a captured target analyte and similar reporter-conjugated detection antibody. The bead analyte reporter constructs are analysed in a flow chamber where lasers excite the reporters and the emitted light is collected by a series of detectors for quantitative analysis (Tighe et al. 2015).
Multiplex assays as an alternative to animal research
Alternatives to animal testing include in vitro tests using cell lines, tissue samples, 3-dimensional modelling and bioprinting, in silico tests, organ-on-chip technologies such as 3-dimensional organoids, and computer modelling (Groell et al. 2018; Mittal et al. 2019; Benam et al. 2019; Doke and Dhawale 2015; Burt et al. 2018; Herrmann et al. 2019). The advancements in bio-informatics and in silico computational methods along with cell and tissue engineering have been developed as alternatives to animal studies (Swaminathan et al. 2019 May; Danchin et al. 1991; Sieburg 1990). The two important advancements are (i) organ (animal/human)-on-a-chip models, which employ multi-channel 3D-microfluidic cell culture chips to simulate the activity, mechanics, and physiological responses of organs and organ systems, acting as a type of artificial organ. In this model, a tandem connection to other organs using flow dynamics can be used to create more relevant in vivo systems (Souza et al. 2018); (ii) disease-in-a-dish model, where patient-derived cells are grown into organoids for prediction of individual drug responses, and successful phase III clinical trials (Orwant 2006; Hall 2011). The ability to grow cells in 3D in controlled environment has lead to the development of cultured organs (organoids) that show functional properties of organs in the human body. These organoids are useful tools for testing drug, pathology, and regenerative medicine, as they replicate human physiology, diseases, and drug response (Fig. 4). The 3D-culture systems provide cost-effective and relevant information and a better translational model compared to experimental animal studies. Thus, the conventional clinical diagnostics tools that measure plasma creatinine, cholesterol, and glucose are being expanded to include multiplexing technology utilizing omics tools (Swaminathan et al. 2019 May; Low et al. 2018). The goal for reducing the time and cost of drug development, together with reduced public support for animal research, is driving attempts to find alternatives to animal testing such as multiplexing and microarray platforms. In addition, accurately identifying the agents that are human safe but fail animal testing will more likely increase potential effective therapeutics in human diseases.
The concept of organ-on-a-chip e.g., multiorgan chips using biosensors, organoids derived from stem cells, and patient-derived diseased organs (Source: www.reagenebiosciences.com. Reproduced from (Swaminathan et al. 2019 May))
Conclusion
The monoplex/simplex immunoanalytical techniques are widely used as diagnostic or analytical tools in biomedical research for the detection and quantification of specific antigens or antibodies in a given sample through the method of ELISA. However, some diagnostic laboratories and organizations employ alternative screening analytical platforms such as the microarray and multiplex immunoassays. The multiplex assay is the simultaneous on-site detection of different analytes from a single specimen and has recently gained importance in clinical diagnostics or directly at POC. These techniques are based on protein or antibody microarrays or chips, coated beads, glass fibres, or microcapillary discs using immobilized RNA, DNA, cDNA, and aptamers. The multiplex reactions are sometimes advantageous over conventional immunoassays by performing many reactions and the ability to extract more information from the same sample in a fast and efficient manner. However, multiplex assays are not available in most diagnostic laboratories and requires expensive equipment and trained personal. The multiplex assays require advanced methodology or/and technology such as fluorescence or chemiluminescence (PCR, ELISA), assays (microarrays, gel electrophoresis), and signals (capillary electrophoresis). The LFIA is one of the most successful analytical platforms for the on-site detection of target substances. The features of LFIA have made them a very important and significant tool in clinical diagnostic where they can improve patient care by enabling prompt diagnosis and treatment. The goal of reducing the time and cost of drug development are the incentives and driving forces to find alternatives to animal testing that may bring about an improvement in identifying human safety and toxicity for therapeutics. In addition, accurate identification of agents that are human safe will increase potential effective therapeutics in human diseases. The alternatives to animal research such as cell and tissue platforms, computational in silico modelling, 3D tissue platforms, and organ-on-chip research have shown great promise in diagnostics and pharmacology and may have a significant impact on advancements and technological developments.
Abbreviations
- EIA:
-
Enzyme immunoassay
- CIA:
-
Chemiluminescence assay
- PCR:
-
Polymerase chain reaction
- FC:
-
Flow cytometry
- MS:
-
Mass spectrometry
- ELISA:
-
Enzyme-linked immunosorbent assay
- RIA:
-
Radio immunoassay
- EMIT:
-
The enzyme multiplied immunoassay technique
- FPIA:
-
Fluorescence polarization immunoassays assay
- SPIA:
-
Sol particle immunoassay
- LFIA:
-
Lateral flow immunoassay
- NALFIA:
-
Nucleic acid lateral flow immunoassay
- NALF:
-
Nucleic acid lateral flow assay
- POC:
-
Point-of-care
- PON:
-
Point-of-need
References
Ahmad R, Ahsan H (2014) Role of peroxynitrite-modified biomolecules in the etiopathogenesis of systemic lupus erythematosus. Clin Exp Med 14(1):1–11
Ahsan H (2013) 3-Nitrotyrosine: a biomarker of nitrogen free radical species modified proteins in systemic autoimmunogenic conditions. Hum Immunol 74(10):1392–1399
Ahsan H (2019) The biomolecules of beauty: biochemical pharmacology and immunotoxicology of cosmeceuticals. J Immunoassay Immunochem 40(1):91–108
Ahsan H, Ahmad R (2020) Multiplex technology for biomarker immunoassays. 1–10. https://doi.org/10.5772/intechopen.91730
Andryukov BG (2020) Six decades of lateral flow immunoassay: from determining metabolic markers to diagnosing COVID-19. AIMS Microbiol 6(3):280–304
Annesley TM (2010) It’s about the journey, not the destination: the birth of radioimmunoassay. Clin Chem 56(4):671–672
Antiochia R (2021) Paper-based biosensors: frontiers in point-of-care detection of COVID-19 disease. Biosensors 11:110
Babrak LM, Menetski J, Rebhan M, Nisato G, Zinggeler M, Brasier N, Baerenfaller K, Brenzikofer T, Baltzer L, Vogler C, Gschwind L, Schneider C, Streiff F, Groenen PMA, Miho E (2019) Traditional and digital biomarkers: two worlds apart? Digit Biomark 3(2):92–102
Bays HE, Chapman RH, Grandy S (2007) SHIELD Investigators’ Group. The relationship of body mass index to diabetes mellitus, hypertension and dyslipidaemia: comparison of data from two national surveys. Int J Clin Pract 61(5):737–47
Benam KH, Gilchrist S, Kleensang A, Satz AB, Willet C, Zhang Q (2019) Exploring new technologies in biomedical research. Drug Discov Today 24:1242–1247
Berton TR, Mitchell DL (2012) Quantification of DNA photoproducts in mammalian cell DNA using radioimmunoassay. Methods Mol Biol 920:177–187
Boguszewska K, Szewczuk M, Urbaniak S, Karwowski BT (2019) Review: immunoassays in DNA damage and instability detection. Cell Mol Life Sci 76(23):4689–4704
Bonventre JV, Vaidya VS, Schmouder R, Feig P, Dieterle F (2010) Next-generation biomarkers for detecting kidney toxicity. Nat Biotechnol 28(5):436–440
Burt T, Vuong LT, Baker E et al (2018) Phase 0, including microdosing approaches: applying the three Rs and increasing the efficiency of human drug development. Altern Lab Anim 46:335–4
Chard T (1996) Immunoassay. Elsevier
Cox KL, Devanarayan V, Kriauciunas A, Manetta J, Montrose C, and Sittampalam S (2004) Immunoassay methods. Eli Lilly & Company and the National Center for Advancing Translational Sciences
Danchin A, Médigue C, Gascuel O, Soldano H, Hénaut A (1991) From data banks to data bases. Res Microbiol 142:913–916
David W (2005) The immunoassay handbook, 3rd edn. Elsevier Ltd, Amsterdam
Davies C (2013) Principles of competitive and immunometric assays (including ELISA). Immunoass Handb 29–59
de Souza AC, Sardela VF, de Sousa VP, Pereira HMG (2018) Zebrafish (Danio rerio): a valuable tool for predicting the metabolism of xenobiotics in humans? Comp Biochem Physiol C Toxicol Pharmacol 212:34–46
Deeks JJ, Dinnes J, Takwoingi Y, Davenport C, Spijker R, Taylor-Phillips S, Adriano A, Beese S, Dretzke J, Ferrante di Ruffano L et al (2020) Cochrane COVID-19 Diagnostic Test Accuracy Group. Antibody tests for identification of current and past infection with SARS-CoV-2. Cochrane Database Syst Rev 6:CD013652
Derksen RH, Bast EJ, Strooisma T, Jacobs JW (2002) A comparison between the Farr radioimmunoassay and a new automated fluorescence immunoassay for the detection of antibodies against double stranded DNA in serum. Ann Rheum Dis 61(12):1099–1102
Di Nardo F, Chiarello M, Cavalera S, Baggiani C, Anfossi LT (2021) years of lateral flow immunoassay technique applications: trends, challenges and future perspectives. Sensors 21:5185
Dincer C, Bruch R, Kling A, Dittrich PS, Urban GA (2017) Multiplexed point-of-care testing - xPOCT. Trends Biotechnol 35(8):728–742
Dinnes J, Deeks JJ, Adriano A, Berhane S, Davenport C, Dittrich S, Emperador D, Takwoingi Y, Cunningham J, Beese S et al (202) Cochrane COVID-19 Diagnostic Test Accuracy Group. Rapid, point-of-care antigen and molecular-based tests for diagnosis of SARS-CoV-2 infection. Cochrane Database Syst Rev 8:CD013705
Doke SK, Dhawale SC (2015) Alternatives to animal testing: a review. Saudi Pharm J 23:223–229
Ekins RP (1989) Multi-analyte immunoassay. J Pharm Biomed Anal 7:155–168
Ellington AA, Kullo IJ, Bailey KR, Klee GG (2010) Antibody-based protein multiplex platforms: technical and operational challenges. Clin Chem 56(2):186–193
Elmlinger MW (2011) Laboratory measurements of hormones and related biomarkers: technologies, quality management and validation. Ranke and Mullis. P-E (eds): Diagnostics of Endocrine Function in Children and Adolescents, ed 4. Basel, Karger 1–31
Engvall E, Perlmann P (1971) Enzyme-linked immunosorbent assay (ELISA). Quantitative Assay of Immunoglobulin g Immunochemistry 8:871–874
Ernst E, Wolfe P, Stahura C, Edwards KA (2021) Technical considerations to development of serological tests for SARS-CoV-2. Talanta 224:121883
Espejo AP, Akgun Y, Al Mana AF, Tjendra Y, Millan NC, Gomez-Fernandez C, Cray C (2020) Review of current advances in serologic testing for COVID-19. Am J Clin Pathol 154:293–304
European Medicines Agency (2018) Committee for Medicinal Products Human Use (CHMP) European Medicines Agency. https://www.ema.europa.eu/en/committees/committee-medicinalproducts-human-use-chmp
Fadal RG (1992) Experience with RAST-based immunotherapy. Otolaryngol Clin N Am 25(1):43–60
FDA-NIH Biomarker Working Group (2016) BEST (biomarkers, endpoints, and other tools) resource. Silver Spring: FDA. Available from: http://www.ncbi.nlm.nih.gov/books/NBK326791/
Feinberg JG (1961) A ‘Microspot’ test for antigens and antibodies. Nature 192:985–986
Gan SD, Patel KR (2013) Enzyme immunoassay and enzyme-linked immunosorbent assay. J Invest Dermatol 133(9):e12
Goldsmith SJ (1975) Radioimmunoassay: review of basic principles. Semin Nucl Med 5(2):125–152
González-Martínez MÁ, Puchades R, Maquieira Á (2018) Immunoanalytical technique: enzyme-linked immunosorbent assay (ELISA). Modern Tech Food Authent Elsevier 617–57
Grange RD, Thompson JP, Lambert DG (2014) Radioimmunoassay, enzyme and non-enzyme-based immunoassays. Br J Anaesth 112(2):213–216
Groell F, Jordan O, Borchard G (2018) In vitro models for immunogenicity prediction of therapeutic proteins. Eur J Pharm Biopharm 130:128–142
Hall SS (2011) Diseases in a dish. Sci Am 304:40–45
Haschek WM, Rousseaux CG, Wallig MA (2010) Fundamentals of toxicologic pathology. Boston
Hsu H-Y, Joos TO, Koga H (2009) Multiplex microsphere based flow cytometric platforms for protein analysis and their application in clinical proteomics — from assays to results. Electrophoresis 30:4008–4019
Herrmann K, Pistollato F, Stephens ML (2019) Beyond the 3 Rs: expanding the use of humanrelevant replacement methods in biomedical research. Altex 36:343–352
Jørgensen CS, Uldum SA, Sørensen JF, Skovsted IC, Otte S, Elverdal PL (2015) Evaluation of a new lateral flow test for detection of Streptococcus pneumoniae and Legionella pneumophila urinary antigen. J Microbiol Methods 116:33–36
Kailasa SK, Mehta VN, Koduru JR, Basu H, Singhal RK, Murthy ZVP, Park T-J (2021) An overview of molecular biology and nanotechnology based analytical methods for the detection of SARS-CoV-2: promising biotools for the rapid diagnosis of COVID-19. Analyst 146:1489–1513
Kar S, Paglialunga S, Islam R (2018) Cystatin C is a more reliable biomarker for determining eGFR to support drug development studies. J Clin Pharmacol 58(10):1239–1247
Kettler H, White K, Hawkes SJ (2004) UNDP/World Bank/WHO Special Programme for Research and Training in Tropical Diseases. Mapping the Landscape of Diagnostics for Sexually Transmitted Infections: Key Findings and Recommendations. Available online: https://apps.who.int/iris/handle/10665/68990
Krishhan VV, Khan IH, Luciw PA (2009) Multiplexed microbead immunoassays by flow cytometry for molecular profiling: basic concepts and proteomics applications. Crit Rev Biotechnol 29(1):29–43
Li Y, Zhang G, Mao X, Yang S, De Ruyck K, Wu Y (2018) High sensitivity immunoassays for small molecule compounds detection — novel noncompetitive immunoassay designs. TrAC Trends Anal Chem 103:198–208
Low SK, Zembutsu H, Nakamura Y (2018) Breast cancer: the translation of big genomic data to cancer precision medicine. Cancer Sci 109:497–506
McCarthy J (2003) Immunological techniques: ELISA. Detect Path Food 241–58
Mekonnen D, Mengist HM, Derbie A, Nibret E, Munshea A, He H, Li B, Jin T (2020) Diagnostic accuracy of serological tests and kinetics of severe acute respiratory syndrome coronavirus 2 antibody: a systematic review and meta-analysis. Rev Med Virol 31:e2181
Miles LE (1975) Properties, variants, and applications of the immunoradiometric assay method. Ric Clin Lab 5(1):59–72
Mittal R, Woo FW, Castro CS et al (2019) Organ-onchip models: implications in drug discovery and clinical applications. J Cell Physiol 234:8352–8380
Nolan JP, Mandy F (2006) Multiplexed and microparticle-based analyses: quantitative tools for the large-scale analysis of biological systems. Cytometry A 69(5):318–325
Oishee MJ, Ali T, Jahan N, Khandker SS, Haq MA, Khondoker MU, Sil BK, Lugova H, Krishnapillai A, Abubakar AR, Kumar S, Haque M, Jamiruddin MR, Adnan N (2021) COVID-19 pandemic: review of contemporary and forthcoming detection tools. Infect Drug Resist 14:1049–1082
Orwant R (2006) Dawn of the zombies. New Sci 255:40–43
Pal AK (2015) Overview of multiplex immunoassays. Res Trend Biochem, Mol Biol Microbiol. https://smjournals.com/ebooks/research-trends-biochemistry-molecular-biology-microbiology/chapters/RTBMBM-15–01.pdf
Posthuma-Trumpie GA, Korf J, van Amerongen A (2009) Lateral flow (immuno)assay: its strengths, weaknesses, opportunities and threats. A Literature Survey Anal Bioanal Chem 393(2):569–582
Preedy VR, Burrow GN, Watson R (2009) Comprehensive handbook of iodine: nutritional, biochemical, pathological and therapeutic aspects. Elsevier, Amsterdam
Rajagopal A, Scherer A, Homyk A, Kartalov E (2013) Supercolor coding methods for large-scale multiplexing of biochemical assays. Anal Chem 85(16):7629–7636
Ruhan A, Wang H, Wang W, Tan W (2020) Summary of the detection kits for SARS-CoV-2 approved by the national medical products administration of China and their application for diagnosis of COVID-19. Virol Sin 35:699–712
Satoh M, Tanaka S, Chan EK (2015) The uses and misuses of multiplex autoantibody assays in systemic autoimmune rheumatic diseases. Front Immunol 6:181
Sieburg HB (1990) Physiological studies in silico. Stud Sci Complex 12:321–342
Slage KM, Ghosn SJ (1996) Immunoassays. tools for sensitive, specific, and accurate test results. Lab Med 27(3):117–83
Smeenk R, Brinkman K, van den Brink H, Swaak T (1990) A comparison of assays used for the detection of antibodies to DNA. Clin Rheumatol 9(1 Suppl 1):63–72
Smeenk RJ, van den Brink HG, Brinkman K, Termaat RM, Berden JH, Swaak AJ (1991) Anti-dsDNA: choice of assay in relation to clinical value. Rheumatol Int 11(3):101–107
Swaminathan S, Kumar V, Kaul R (2019) Need for alternatives to animals in experimentation: An Indian perspective. Indian J Med Res 149(5):584–592
Tighe PJ, Ryder RR, Todd I, Fairclough LC (2015) ELISA in the multiplex era: potentials and pitfalls. Proteomics Clin Appl 9(3–4):406–422
Tozzoli R, D’Aurizio F, Villalta D, Bizzaro N (2015) Automation, consolidation, and integration in autoimmune diagnostics. Autoimmun Highlights 6(1–2):1–6
Van Weemen BK, Schuurs AH (1971) Immunoassay using antigen-enzyme conjugates. FEBS Lett 15:232–236
Vashist SK (2020) In vitro diagnostic assays for COVID-19: recent advances and emerging trends. Diagnostics 10:202
Wu AH (2006) A selected history and future of immunoassay development and applications in clinical chemistry. Clin Chim Acta 369(2):119–124
Yalow RS, Berson SA (1959) Assay of plasma insulin in human subjects by immunological methods. Nature 184(Suppl. 21):1648–1649
Yalow RS, Berson SA (1960) Immunoassay of endogenous plasma insulin in man. J Clin Invest 39:1157–1175
Zheng W, He L (2012) Multiplexed immunoassays. Advances in immunoassay technology. Edited by Norman H. L. Chiu and Theodore K. Christopoulos
Zhu N, Woong PK (2020) Advances in viral diagnostic technologies for combating COVID-19 and future pandemics. SLAS Technol 25:513–521
Acknowledgements
The review article is dedicated in the memory of my beloved father, one of the most generous and compassionate human beings who lived and passed this side of the world. I would like to thank my family for their endless, everlasting, and eternal support in life and wellbeing. I also wish to acknowledge the inspiration, motivation, and encouragement of Prof. Fahim H. Khan and Prof. Waseem A. Siddiqui, Aligarh Muslim University, India. I wish to thank the anonymous reviewers for their appreciation, suggestions, and recommendation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author declares no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ahsan, H. Monoplex and multiplex immunoassays: approval, advancements, and alternatives. Comp Clin Pathol 31, 333–345 (2022). https://doi.org/10.1007/s00580-021-03302-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00580-021-03302-4