Abstract
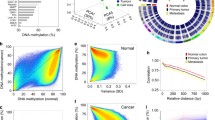
We describe a newly developed technique for rapid identification of positions of genomic DNA breaks, preexisting or introduced by specific digestion, in particular, by restriction endonucleases (RIDGES). We applied RIDGES in analyzing unmethylated CCGG sites distribution along a 1-Mb long genome region (D19S208–COX7A1 on chromosome 19) in cancerous and normal lung tissues. Both tissues were characterized by a profoundly uneven density of unmethylated sites along the fragment. Interestingly, the distribution of hypomethylated regions did not correlate with gene locations within the fragment, and one of the most hypomethylated areas contained practically no genes. We also demonstrated that the methylation pattern of a long genome DNA fragment was rather stable and practically unchanged in human lung cancer tissue as compared with its normal counterpart, in accordance with the suggestion (Ross et al. in Nat Genet 24:227–235, 2000) that cell lines of common origin have typically similar transcription profiles. An analogous suggestion might probably be made for global methylation patterns of genomic DNA.



Similar content being viewed by others
References
Adorjan P, Distler J, Lipscher E, Model F, Muller J, Pelet C, Braun A, Florl AR, Gutig D, Grabs G, Howe A, Kursar M, Lesche R, Leu E, Lewin A, Maier S, Muller V, Otto T, Scholz C, Schulz WA, Seifert HH, Schwope I, Ziebarth H, Berlin K, Piepenbrock C, Olek A (2002) Tumour class prediction and discovery by microarray-based DNA methylation analysis. Nucleic Acids Res 30:e21
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Antequera F (2003) Structure, function and evolution of CpG island promoters. Cell Mol Life Sci 60:1647–1658
Azhikina T, Gainetdinov I, Skvortsova Y, Batrak A, Dmitrieva N, Sverdlov E (2004) Non-methylated Genomic Sites Coincidence Cloning (NGSCC): an approach to large scale analysis of hypomethylated CpG patterns at predetermined genomic loci. Mol Genet Genomics 271:22–32
Baylin SB, Herman JG (2000) DNA hypermethylation in tumorigenesis: epigenetics joins genetics. Trends Genet 16:168–174
Chernov IP, Akopov SB, Nikolaev LG, Sverdlov ED (2002) Identification and mapping of nuclear matrix-attachment regions in a one megabase locus of human chromosome 19q13.12: long-range correlation of S/MARs and gene positions. J Cell Biochem 84:590–600
Chong S, Whitelaw E (2004) Epigenetic germline inheritance. Curr Opin Genet Dev 14:692–696
Ehrlich M (2002) DNA methylation in cancer: too much, but also too little. Oncogene 21:5400–5413
Fraga MF, Esteller M (2002) DNA methylation: a profile of methods and applications. Biotechniques 33:632, 634, 636–649
Futscher BW, Oshiro MM, Wozniak RJ, Holtan N, Hanigan CL, Duan H, Domann FE (2002) Role for DNA methylation in the control of cell type specific maspin expression. Nat Genet 31:175–179
International Human Genome Sequencing Consortium (2001) Initial sequencing and analysis of the human genome. Nature 409:860–921
Kusui C, Kimura T, Ogita K, Nakamura H, Matsumura Y, Koyama M, Azuma C, Murata Y (2001) DNA methylation of the human oxytocin receptor gene promoter regulates tissue-specific gene suppression. Biochem Biophys Res Commun 289:681–686
Lavrentieva I, Broude NE, Lebedev Y, Gottesman II, Lukyanov SA, Smith CL, Sverdlov ED (1999) High polymorphism level of genomic sequences flanking insertion sites of human endogenous retroviral long terminal repeats. FEBS Lett 443:341–347
Leung YF, Cavalieri D (2003) Fundamentals of cDNA microarray data analysis. Trends Genet 19:649–659
Olek A, Oswald J, Walter J (1996) A modified and improved method for bisulphite based cytosine methylation analysis. Nucleic Acids Res 24:5064–5066
Olsen AS, Georgescu A, Johnson S, Carrano AV (1996) Assembly of a 1-Mb restriction-mapped cosmid contig spanning the candidate region for Finnish congenital nephrosis (NPHS1) in 19q13.1. Genomics 34:223–225
Robinson PN, Bohme U, Lopez R, Mundlos S, Nurnberg P (2004) Gene-ontology analysis reveals association of tissue-specific 5′ CpG-island genes with development and embryogenesis. Hum Mol Genet 13:1969–1978
Ross DT, Scherf U, Eisen MB, Perou CM, Rees C, Spellman P, Iyer V, Jeffrey SS, Van de Rijn M, Waltham M, Pergamenschikov A, Lee JC, Lashkari D, Shalon D, Myers TG, Weinstein JN, Botstein D, Brown PO (2000) Systematic variation in gene expression patterns in human cancer cell lines. Nat Genet 24:227–235
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Velculescu VE, Zhang L, Vogelstein B, Kinzler KW (1995) Serial analysis of gene expression. Science 270:484–487
Weber M, Davies JJ, Wittig D, Oakeley EJ, Haase M, Lam WL, Schubeler D (2005) Chromosome-wide and promoter-specific analyses identify sites of differential DNA methylation in normal and transformed human cells. Nat Genet 37:853–862
Acknowledgments
The authors thank V.K. Potapov and N.V. Skaptsova for oligonucleotide synthesis and B.O. Glotov for critical reading of the manuscript. The work was financially supported by grants N 2006.2003.4 (grant of the President of the Russian Federation) and a Russian Academy of Sciences grant of the Program “Physico-chemical biology. Structural, functional, and evolutional analysis of genomic cis-regulatory systems.”
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by G. Georgiev
Tatyana Azhikina and Ildar Gainetdinov have equally contributed to the work
Rights and permissions
About this article
Cite this article
Azhikina, T., Gainetdinov, I., Skvortsova, Y. et al. Methylation-free site patterns along a 1-Mb locus on Chr19 in cancerous and normal cells are similar. A new fast approach for analyzing unmethylated CCGG sites distribution. Mol Genet Genomics 275, 615–622 (2006). https://doi.org/10.1007/s00438-006-0111-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-006-0111-2