Abstract
Purpose
To explore whether targeted next generation sequencing (NGS) of liquid biopsy in advanced non-small cell lung cancer (NSCLC) could potentially overcome the innate problems that arise with standard tissue biopsy, like intratumoral heterogeneity and the inability to obtain adequate samples for analysis.
Methods
The Scopus, Cochrane Library, and MEDLINE (via PubMed) databases were searched for studies with matched tissue and liquid biopsies from advanced NSCLC patients, analyzed with targeted NGS. The number of mutations detected in tissue biopsy only, liquid biopsy only, or both was assessed and the positive percent agreement (PPA) of the two methods was calculated for every clinically relevant gene.
Results
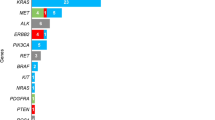
A total of 644 unique relevant articles were retrieved and data were extracted from 38 studies fulfilling the inclusion criteria. The sample size was composed of 2000 mutations tested in matched tissue and liquid biopsies derived from 1141 patients. No studies analyzed circulating tumor cells. The calculated PPA rates were 53.6% (45/84) for ALK, 53.9% (14/26) for BRAF, 56.5% (13/23) for ERBB2, 67.8% (428/631) for EGFR, 64.2% (122/190) for KRAS, 58.6% (17/29) for MET, 54.6% (12/22) for RET, and 53.3% (8/15) for ROS1. We additionally recorded data for 65 genes that are not recommended by current guidelines for mutational testing. An extra category containing results of unspecified genes was added, with a PPA rate of 55.7% (122/219).
Conclusion
Despite many advantages, liquid biopsy might be unable to fully substitute its tissue counterpart in detecting clinically relevant mutations in advanced NSCLC patients. However, it may serve as a helpful tool when making therapeutic decisions. More studies are needed to evaluate its role in everyday clinical practice.




Similar content being viewed by others
Data availability
Raw data available upon request.
References
Aggarwal C, Thompson JC, Black TA et al (2019) Clinical Implications of Plasma-Based Genotyping with the Delivery of Personalized Therapy in Metastatic Non-Small Cell Lung Cancer. JAMA Oncol 5:173–180. https://doi.org/10.1001/jamaoncol.2018.4305
Ai B, Liu H, Huang Y, Peng P (2016) Circulating cell-free DNA as a prognostic and predictive biomarker in non-small cell lung cancer. Oncotarget 7:44583–44595. https://doi.org/10.18632/oncotarget.10069
Chae YK, Oh MS (2019) Detection of minimal residual disease using ctDNA in lung cancer: current evidence and future directions. J Thorac Oncol 14:16–24. https://doi.org/10.1016/j.jtho.2018.09.022
Chen Y, Han T, Zhou Y et al (2019) Comparing the efficacy of targeted next-generation sequencing in the identification of somatic mutations in circulating tumor DNA from different stages of lung cancer. Neoplasma 66:652–660. https://doi.org/10.4149/neo_2018_181130N910
Chinniah C, Aguarin L, Cheng P et al (2019) Early detection of recurrence in patients with locally advanced non–small-cell lung cancer via circulating tumor cell analysis. Clin Lung Cancer. https://doi.org/10.1016/j.cllc.2019.04.011
Couraud S, Vaca-Paniagua F, Villar S et al (2014) Noninvasive diagnosis of actionable mutations by deep sequencing of circulating free DNA in lung cancer from never-smokers: a proof-of-concept study from BioCAST/IFCT-1002. Clin Cancer Res 20:4613–4624. https://doi.org/10.1158/1078-0432.CCR-13-3063
Dagogo-Jack I, Brannon AR, Ferris LA et al (2018) Tracking the evolution of resistance to ALK tyrosine kinase inhibitors through longitudinal analysis of circulating tumor DNA. JCO Precis Oncol 2018:1–14. https://doi.org/10.1200/po.17.00160
Dono M, De Luca G, Lastraioli S et al (2019) Tag-based next generation sequencing: a feasible and reliable assay for EGFR T790M mutation detection in circulating tumor DNA of non small cell lung cancer patients. Mol Med 25:15. https://doi.org/10.1186/s10020-019-0082-5
El Achi H, Khoury JD, Loghavi S et al (2019) Liquid biopsy by next-generation sequencing: a multimodality test for management of cancer. Curr Hematol Malig Rep 14:358–367. https://doi.org/10.1007/s11899-019-00532-w
Ettinger DS, Aisner DL, Wood DE et al (2018) NCCN guidelines ® insights non–small cell lung cancer, version 5.2018 featured updates to the NCCN guidelines. J Natl Compr Cancer Netw 16:807–821. https://doi.org/10.6004/jnccn.2018.0062
Fernandez-Marmiesse A, Gouveia S, Couce ML (2017) NGS technologies as a turning point in rare disease research, diagnosis and treatment. Curr Med Chem 25:404–432. https://doi.org/10.2174/0929867324666170718101946
Gai W, Sun K (2019) Epigenetic biomarkers in cell-free DNA and applications in liquid biopsy. Genes (Basel). https://doi.org/10.3390/genes10010032
Gallo M, De Luca A, Frezzetti D et al (2019) The potential of monitoring treatment response in non-small cell lung cancer using circulating tumour cells. Expert Rev Mol Diagn 19:1–12. https://doi.org/10.1080/14737159.2019.1640606
Garcia J, Forestier J, Dusserre E et al (2018) Cross-platform comparison for the detection of RAS mutations in cfDNA (ddPCR Biorad detection assay, BEAMing assay, and NGS strategy). Oncotarget 9:21122–21131. https://doi.org/10.18632/oncotarget.24950
Ge M, Zhan Q, Zhang Z et al (2019) Different next-generation sequencing pipelines based detection of tumor DNA in cerebrospinal fluid of lung adenocarcinoma cancer patients with leptomeningeal metastases. BMC Cancer 19:1–8. https://doi.org/10.1186/s12885-019-5348-3
Giroux Leprieur E, Herbretau G, Dumenil C et al (2018) Circulating tumor DNA evaluated by Next-Generation Sequencing is predictive of tumor response and prolonged clinical benefit with nivolumab in advanced non-small cell lung cancer. Oncoimmunology 7:e1424675. https://doi.org/10.1080/2162402X.2018.1424675
Guo Q, Wang J, Xiao J et al (2018) Heterogeneous mutation pattern in tumor tissue and circulating tumor DNA warrants parallel NGS panel testing. Mol Cancer 17:4–8. https://doi.org/10.1186/s12943-018-0875-0
Hagemann IS, Devarakonda S, Lockwood CM et al (2015) Clinical next-generation sequencing in patients with non-small cell lung cancer. Cancer 121:631–639. https://doi.org/10.1002/cncr.29089
Hanna N, Johnson D, Temin S et al (2017) Systemic therapy for stage IV non–small-cell lung cancer: American Society of clinical oncology clinical practice guideline update. J Clin Oncol 35:3484–3515. https://doi.org/10.1200/JCO.2017.74.6065
Heerink WJ, de Bock GH, de Jonge GJ et al (2017) Complication rates of CT-guided transthoracic lung biopsy: meta-analysis. Eur Radiol 27:138–148. https://doi.org/10.1007/s00330-016-4357-8
Horn L, Whisenant JG, Wakelee H et al (2019) Monitoring therapeutic response and resistance: analysis of circulating tumor DNA in patients With ALK+ lung cancer. J Thorac Oncol 14:1901–1911. https://doi.org/10.1016/j.jtho.2019.08.003
Hu Y, Ulrich BC, Supplee J et al (2018) False-positive plasma genotyping due to clonal hematopoiesis. Clin Cancer Res 24:4437–4443. https://doi.org/10.1158/1078-0432.CCR-18-0143
Iwama E, Sakai K, Azuma K et al (2017) Monitoring of somatic mutations in circulating cell-free DNA by digital PCR and next-generation sequencing during afatinib treatment in patients with lung adenocarcinoma positive for EGFR activating mutations. Ann Oncol Off J Eur Soc Med Oncol 28:136–141. https://doi.org/10.1093/annonc/mdw531
Iwama E, Sakai K, Azuma K et al (2018) Exploration of resistance mechanisms for epidermal growth factor receptor-tyrosine kinase inhibitors based on plasma analysis by digital polymerase chain reaction and next-generation sequencing. Cancer Sci 109:3921–3933. https://doi.org/10.1111/cas.13820
Jennings LJ, Arcila ME, Corless C et al (2017) Guidelines for validation of next-generation sequencing-based oncology panels: a joint consensus recommendation of the Association for Molecular Pathology and College of American Pathologists. J Mol Diagnostics 19:341–365. https://doi.org/10.1016/j.jmoldx.2017.01.011.Guidelines
Jin Y, Shi X, Zhao J et al (2018) Mechanisms of primary resistance to EGFR targeted therapy in advanced lung adenocarcinomas. Lung Cancer 124:110–116. https://doi.org/10.1016/j.lungcan.2018.07.039
Kaisaki PJ, Cutts A, Popitsch N et al (2016) Targeted next-generation sequencing of plasma DNA from cancer patients: Factors influencing consistency with tumour DNA and prospective investigation of its utility for diagnosis. PLoS ONE 11:1–13. https://doi.org/10.1371/journal.pone.0162809
Kalemkerian GP, Narula N, Kennedy EB et al (2018) Molecular testing guideline for the selection of patients with lung cancer for treatment with targeted tyrosine kinase inhibitors: American society of clinical oncology endorsement of the college of American pathologists/ international association for the. J Clin Oncol 36:911–919. https://doi.org/10.1200/JCO.2017.76.7293
Karachaliou N, Mayo-De Las Casas C, Queralt C et al (2015) Association of EGFR L858R mutation in circulating free DNA with survival in the EURTAC trial. JAMA Oncol 1:149–157. https://doi.org/10.1001/jamaoncol.2014.257
Kerr KM, Bubendorf L, Edelman MJ et al (2014) Second ESMO consensus conference on lung cancer: pathology and molecular biomarkers for non-small-cell lung cancer. Ann Oncol 25:1681–1690. https://doi.org/10.1093/annonc/mdu145
Lam VK, Tran HT, Banks KC et al (2019) Targeted tissue and cell-free tumor DNA sequencing of advanced lung squamous-cell carcinoma reveals clinically significant prevalence of actionable alterations. Clin Lung Cancer 20:30–36.e3. https://doi.org/10.1016/j.cllc.2018.08.020
Laufer-Geva S, Rozenblum AB, Twito T et al (2018) The clinical impact of comprehensive genomic testing of circulating cell-free DNA in advanced lung cancer. J Thorac Oncol 13:1705–1716. https://doi.org/10.1016/j.jtho.2018.07.101
Lee JY, Qing X, Xiumin W et al (2016) Longitudinal monitoring of EGFR mutations in plasma predicts outcomes of NSCLC patients treated with EGFR TKIs: Korean Lung Cancer Consortium (KLCC-12–02). Oncotarget 7:6984–6993. https://doi.org/10.18632/oncotarget.6874
Leighl NB, Page RD, Raymond VM et al (2019) Clinical utility of comprehensive cell-free DNA analysis to identify genomic biomarkers in patients with newly diagnosed metastatic non–small cell lung cancer. Clin Cancer Res 25:4691–4700. https://doi.org/10.1158/1078-0432.ccr-19-0624
Levy MA, Lovly CM, Pao W (2012) Translating genomic information into clinical medicine: Lung cancer as a paradigm. Genome Res 22:2101–2108. https://doi.org/10.1101/gr.131128.111
Li YS, Jiang BY, Yang JJ et al (2018) Unique genetic profiles from cerebrospinal fluid cell-free DNA in leptomeningeal metastases of EGFR-mutant non-small-cell lung cancer: a new medium of liquid biopsy. Ann Oncol Off J Eur Soc Med Oncol 29:945–952. https://doi.org/10.1093/annonc/mdy009
Li BT, Janku F, Jung B et al (2019) Ultra-deep next-generation sequencing of plasma cell-free DNA in patients with advanced lung cancers: results from the Actionable Genome Consortium. Ann Oncol Off J Eur Soc Med Oncol 30:597–603. https://doi.org/10.1093/annonc/mdz046
Lindeman NI, Cagle PT, Aisner DL et al (2018) Updated molecular testing guideline for the selection of lung cancer patients for treatment with targeted tyrosine kinase inhibitors guideline from the college of American pathologists, the international association for the study of lung cancer, and the a. Arch Pathol Lab Med 142:321–346. https://doi.org/10.5858/arpa.2017-0388-CP
Liu L, Liu H, Shao D et al (2018) Development and clinical validation of a circulating tumor DNA test for the identification of clinically actionable mutations in nonsmall cell lung cancer. Genes Chromosom Cancer 57:211–220. https://doi.org/10.1002/gcc.22522
Mayo-de-Las-Casas C, Garzon Ibanez M, Jordana-Ariza N et al (2018) An update on liquid biopsy analysis for diagnostic and monitoring applications in non-small cell lung cancer. Expert Rev Mol Diagn 18:35–45. https://doi.org/10.1080/14737159.2018.1407243
McCoach CE, Blakely CM, Banks KC et al (2018) Clinical utility of cell-free DNA for the detection of ALK fusions and genomic mechanisms of ALK inhibitor resistance in non–small cell lung cancer. Clin Cancer Res 24:2758–2770. https://doi.org/10.1158/1078-0432.CCR-17-2588
Merker JD, Oxnard GR, Compton C et al (2018) Circulating tumor DNA analysis in patients with cancer: American society of clinical oncology and college of American pathologists joint review. Arch Pathol Lab Med 142:1242–1253. https://doi.org/10.5858/arpa.2018-0901-SA
Mok TS, Wu Y-L, Ahn M-J et al (2016) Osimertinib or platinum-pemetrexed in EGFR T790M–positive lung cancer. N Engl J Med 376:629–640. https://doi.org/10.1056/nejmoa1612674
Overman MJ, Modak J, Kopetz S et al (2013) Use of research biopsies in clinical trials: are risks and benefits adequately discussed? J Clin Oncol 31:17–22. https://doi.org/10.1200/JCO.2012.43.1718
Papadopoulou E, Tsoulos N, Tsantikidi K et al (2019) Clinical feasibility of NGS liquid biopsy analysis in NSCLC patients. PLoS ONE 14:e0226853. https://doi.org/10.1371/journal.pone.0226853
Paweletz CP, Sacher AG, Raymond CK et al (2016) Bias-corrected targeted next-generation sequencing for rapid, multiplexed detection of actionable alterations in cell-free dna from advanced lung cancer patients. Clin Cancer Res 22:915–922. https://doi.org/10.1158/1078-0432.CCR-15-1627-T
Pécuchet N, Zonta E, Didelot A et al (2016) Base-position error rate analysis of next-generation sequencing applied to circulating tumor DNA in non-small cell lung cancer: a prospective study. PLoS Med 13:1–19. https://doi.org/10.1371/journal.pmed.1002199
Petrackova A, Vasinek M, Sedlarikova L et al (2019) Standardization of sequencing coverage depth in NGS: recommendation for detection of clonal and subclonal mutations in cancer diagnostics. Front Oncol 9:1–6. https://doi.org/10.3389/fonc.2019.00851
Phallen J, Leal A, Woodward BD et al (2019) Early noninvasive detection of response to targeted therapy in non–small cell lung cancer. Cancer Res 79:1204–1213. https://doi.org/10.1158/0008-5472.CAN-18-1082
Planchard D, Popat S, Kerr K et al (2018) Metastatic non-small cell lung cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann Oncol 29:192–237. https://doi.org/10.1093/annonc/mdy275
Postel M, Roosen A, Laurent-Puig P et al (2018) Droplet-based digital PCR and next generation sequencing for monitoring circulating tumor DNA: a cancer diagnostic perspective. Expert Rev Mol Diagn 18:7–17. https://doi.org/10.1080/14737159.2018.1400384
Pritchett MA, Camidge DR, Patel M et al (2019) Prospective clinical validation of the invisionfirst-lung circulating tumor dna assay for molecular profiling of patients with advanced nonsquamous non–small-cell lung cancer. JCO Precis Oncol. https://doi.org/10.1200/po.18.00299
Rachiglio AM, Esposito Abate R, Sacco A et al (2016) Limits and potential of targeted sequencing analysis of liquid biopsy in patients with lung and colon carcinoma. Oncotarget 7:66595–66605. https://doi.org/10.18632/oncotarget.10704
Rizvi H, Sanchez-Vega F, La K et al (2018) Molecular determinants of response to anti-programmed cell death (PD)-1 and anti-programmed death-ligand 1 (PD-L1) blockade in patients with non-small-cell lung cancer profiled with targeted next-generation sequencing. J Clin Oncol 36:633–641. https://doi.org/10.1200/JCO.2017.75.3384
Rolfo C, Mack PC, Scagliotti GV et al (2018) Liquid biopsy for advanced non-small cell lung cancer (NSCLC): a statement paper from the IASLC. J Thorac Oncol 13:1248–1268. https://doi.org/10.1016/j.jtho.2018.05.030
Sabari JK, Offin M, Stephens D et al (2018) A prospective study of circulating tumor dna to guide matched targeted therapy in lung cancers. J Natl Cancer Inst 111:575–583. https://doi.org/10.1093/jnci/djy156
Sabour L, Sabour M, Ghorbian S (2017) Clinical applications of next-generation sequencing in cancer diagnosis. Pathol Oncol Res 23:225–234. https://doi.org/10.1007/s12253-016-0124-z
Sacher AG, Paweletz C, Dahlberg SE et al (2016) Prospective validation of rapid plasma genotyping for the detection of EGFR and kras mutations in advanced lung cancer. JAMA Oncol 2:1014–1022. https://doi.org/10.1001/jamaoncol.2016.0173
Santarpia M, Karachaliou N, González-Cao M et al (2016) Feasibility of cell-free circulating tumor DNA testing for lung cancer. Biomark Med 10:417–430. https://doi.org/10.2217/bmm.16.6
Schram AM, Chang MT, Jonsson P, Drilon A (2017) Fusions in solid tumours: diagnostic strategies, targeted therapy, and acquired resistance. Nat Rev Clin Oncol 14:735–748. https://doi.org/10.1038/nrclinonc.2017.127
Schwaederlé MC, Patel SP, Husain H et al (2017) Utility of genomic assessment of blood-derived circulating tumor DNA (ctDNA) in patients with advanced lung adenocarcinoma. Clin Cancer Res 23:5101–5111. https://doi.org/10.1158/1078-0432.CCR-16-2497
Shiau CJ, Babwah JP, Da Cunha SG et al (2014) Sample features associated with success rates in population-based EGFR mutation testing. J Thorac Oncol 9:947–956. https://doi.org/10.1097/JTO.0000000000000196
Siravegna G, Marsoni S, Siena S, Bardelli A (2017) Integrating liquid biopsies into the management of cancer. Nat Rev Clin Oncol 14:531–548. https://doi.org/10.1038/nrclinonc.2017.14
Streubel A, Stenzinger A, Stephan-Falkenau S et al (2019) Comparison of different semi-automated cfDNA extraction methods in combination with UMI-based targeted sequencing. Oncotarget 10:5690–5702. https://doi.org/10.18632/oncotarget.27183
Sundaresan TK, Sequist LV, Heymach JV et al (2016) Detection of T790M, the acquired resistance EGFR mutation, by tumor biopsy versus noninvasive blood-based analyses. Clin Cancer Res 22:1103–1110. https://doi.org/10.1158/1078-0432.CCR-15-1031
Supplee JG, Milan MSDD, Lim LP et al (2019) Sensitivity of next-generation sequencing assays detecting oncogenic fusions in plasma cell-free DNA. Lung Cancer 134:96–99. https://doi.org/10.1016/j.lungcan.2019.06.004
Tamminga M, De Wit S, Hiltermann TJN et al (2019) Circulating tumor cells in advanced non-small cell lung cancer patients are associated with worse tumor response to checkpoint inhibitors. J Immunother Cancer 7:1–9. https://doi.org/10.1186/s40425-019-0649-2
Tan DSW, Camilleri-Broët S, Tan EH et al (2014) Intertumor heterogeneity of non-small-cell lung carcinomas revealed by multiplexed mutation profiling and integrative genomics. Int J Cancer 135:1092–1100. https://doi.org/10.1002/ijc.28750
Tang Y, Liu X, Ou Z et al (2019) Maximum allele frequency observed in plasma: a potential indicator of liquid biopsy sensitivity. Oncol Lett 18:2118–2124. https://doi.org/10.3892/ol.2019.10490
Thress KS, Paweletz CP, Felip E et al (2015) Acquired EGFR C797S mediates resistance to AZD9291 in advanced non-small cell lung cancer harboring EGFR T790M HHS Public Access Author manuscript. Nat Med 21:560–562. https://doi.org/10.1038/nm.3854
Tong L, Ding N, Tong X et al (2019) Tumor-derived DNA from pleural effusion supernatant as a promising alternative to tumor tissue in genomic profiling of advanced lung cancer. Theranostics 9:5532–5541. https://doi.org/10.7150/thno.34070
Toor OM, Ahmed Z, Bahaj W et al (2018) Correlation of somatic genomic alterations between tissue genomics and ctdna employing next-generation sequencing: analysis of lung and gastrointestinal cancers. Mol Cancer Ther 17:1123–1132. https://doi.org/10.1158/1535-7163.MCT-17-1015
Tran LS, Pham H-ATAT, Tran V-UU et al (2019) Ultra-deep massively parallel sequencing with unique molecular identifier tagging achieves comparable performance to droplet digital PCR for detection and quantification of circulating tumor DNA from lung cancer patients. PLoS ONE 14:e0226193. https://doi.org/10.1371/journal.pone.0226193
Travis WD, Brambilla E, Nicholson AG et al (2015) The 2015 World Health Organization classification of lung tumors: impact of genetic, clinical and radiologic advances since the 2004 classification. J Thorac Oncol 10:1243–1260. https://doi.org/10.1097/JTO.0000000000000630
VanderLaan PA, Yamaguchi N, Folch E et al (2014) Success and failure rates of tumor genotyping techniques in routine pathological samples with non-small-cell lung cancer. Lung Cancer 84:39–44. https://doi.org/10.1016/j.lungcan.2014.01.013
Vanni I, Coco S, Truini A et al (2015) Next-generation sequencing workflow for NSCLC critical samples using a targeted sequencing approach by ion torrent PGMTM platform. Int J Mol Sci 16:28765–28782. https://doi.org/10.3390/ijms161226129
Veldore VH, Choughule A, Routhu T et al (2018) Validation of liquid biopsy: plasma cell-free DNA testing in clinical management of advanced non-small cell lung cancer. Lung Cancer Targets Ther 9:1–11. https://doi.org/10.2147/LCTT.S147841
Vestergaard HH, Christensen MR, Lassen UN (2018) A systematic review of targeted agents for non-small cell lung cancer. Acta Oncol (Madr) 57:176–186
Villaflor V, Won B, Nagy R et al (2016) Biopsy-free circulating tumor DNA assay identifies actionable mutations in lung cancer. Oncotarget 7:66880–66891. https://doi.org/10.18632/oncotarget.11801
Vogelstein B, Papadopoulos N, Velculescu VE et al (2013) Cancer genome landscapes NIH public access. Science 339:1546–1558. https://doi.org/10.1126/science.1235122
Vollbrecht C, Lehmann A, Lenze D, Hummel M (2018) Validation and comparison of two NGS assays for the detection of EGFR T790M resistance mutation in liquid biopsies of NSCLC patients. Oncotarget 9:18529–18539. https://doi.org/10.18632/oncotarget.24908
Weber B, Meldgaard P, Hager H et al (2014) Detection of EGFR mutations in plasma and biopsies from non-small cell lung cancer patients by allele-specific PCR assays. BMC Cancer. https://doi.org/10.1186/1471-2407-14-294
Wu Z, Yang Z, Li CS et al (2019) Differences in the genomic profiles of cell-free DNA between plasma, sputum, urine, and tumor tissue in advanced NSCLC. Cancer Med 8:910–919. https://doi.org/10.1002/cam4.1935
Xu T, Kang X, You X et al (2017) Cross-platform comparison of four leading technologies for detecting EGFR mutations in circulating tumor DNA from non-small cell lung carcinoma patient plasma. Theranostics 7:1437–1446. https://doi.org/10.7150/thno.16558
Yang N, Li Y, Liu Z et al (2018) The characteristics of ctDNA reveal the high complexity in matching the corresponding tumor tissues. BMC Cancer 18:1–12. https://doi.org/10.1186/s12885-018-4199-7
Yao Y, Liu J, Li L et al (2017) Detection of circulating tumor DNA in patients with advanced non-small cell lung cancer. Oncotarget 8:2130–2140. https://doi.org/10.18632/oncotarget.12883
Zhu G, Ye X, Dong Z et al (2015) Highly sensitive droplet digital PCR method for detection of EGFR-activating mutations in plasma cell-free DNA from patients with advanced non-small cell lung cancer. J Mol Diagnostics 17:265–272. https://doi.org/10.1016/j.jmoldx.2015.01.004
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
SME: conceptualization, formal analysis, investigation, writing—original draft, and visualization. GIG: conceptualization, formal analysis, investigation, writing—original draft, and visualization. IPN: conceptualization, methodology, validation, writing—original draft, and supervision. VB: methodology, validation, formal analysis, data curation, writing—review and editing. PMS: methodology, writing—review and editing, and supervision. J-KW: methodology, writing—review and editing, and supervision. KPE: conceptualization, methodology, validation, resources, writing—review and editing, supervision, and project administration.
Corresponding author
Ethics declarations
Conflict of interest
The authors report no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Esagian, S.M., Grigoriadou, G.Ι., Nikas, I.P. et al. Comparison of liquid-based to tissue-based biopsy analysis by targeted next generation sequencing in advanced non-small cell lung cancer: a comprehensive systematic review. J Cancer Res Clin Oncol 146, 2051–2066 (2020). https://doi.org/10.1007/s00432-020-03267-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-020-03267-x