Abstract
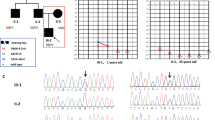
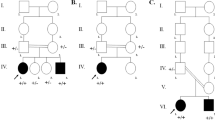
Nonsyndromic genetic deafness is highly heterogeneous in its clinical presentation, pattern of inheritance and underlying genetic causes. Mutations in TMPRSS3 gene encoding transmembrane serine protease account for <1 % of autosomal recessive nonsyndromic hearing loss (ARNSHL) in Caucasians. Targeted next generation sequencing in the index family with profound deaf parents and a son, and Sanger sequencing of selected TMPRSS3 gene regions in a cohort of thirty-five patients with suspected ARNSHL was adopted. A son and his mother in the index family were homozygous for TMPRSS3 c.208delC (p.His70Thrfs*19) variant. Father was digenic compound heterozygote for the same variant and common GJB2 c.35delG variant. Three additional patients from the ARNSHL cohort were homozygous for TMPRSS3 c.208delC. TMPRSS3 defects seem to be an important cause of ARNSHL in Slovenia resulting in uniform phenotype with profound congenital hearing loss, and satisfactory hearing and speech recognition outcome after cochlear implantation. Consequently, TMPRSS3 gene analysis should be included in the first tier of genetic investigations of ARNSHL along with GJB2 and GJB6 genes.
Similar content being viewed by others
References
Morton CW, Nance E (2006) Newborn hearing screening––a silent revolution. N Engl J Med 354:2151–2164
Kenna MA, Feldman HA, Neault MW, Frangulov A, Wu BL, Fligor B, Rehm HL (2010) Audiologic phenotype and progression in GJB2 (Connexin 26) hearing loss. Arch Otolaryngol Head Neck Surg 136:81–87
Beck C, Pérez-Álvarez JC, Sigruener A, Haubner F, Seidler T, Aslanidis C et al (2014) Identification and genotype/phenotype correlation of mutations in a large German cohort with hearing loss. Eur Arch Otorhinolaryngol. doi:10.1007/s00405-014-3157-5
Petersen MB, Willems PJ (2006) Non-syndromic, autosomal–recessive deafness. Clin Genet 69:371–392
Battellino S, Rudolf G, Zargi M, Podkrajsek KT, Peterlin B (2011) Connexin 26 (GJB2) and connexin 30 del(GJB6-D13S1830) mutations in Slovenians with prelingual non-syndromic deafness. Int Adv Otol 7:372–378
Battelino S, Repič Lampret B, Zargi M, Podkrajšek KT (2012) Novel connexin 30 and connexin 26 mutational spectrum in patients with progressive sensorineural hearing loss. J Laryngol Otol 126:763–769
Scott HS, Kudoh J, Wattenhofer M, Shibuya K, Berry A, Chrast R et al (2001) Insertion of beta-satellite repeats identifies a transmembrane protease causing both congenital and childhood onset autosomal recessive deafness. Nat Genet 27:59–63
Wattenhofer M, Di Iorio MV, Rabionet R, Dougherty L, Pampanos A, Schwede T et al (2002) Mutations in the TMPRSS3 gene are a rare cause of childhood nonsyndromic deafness in Caucasian patients. J Mol Med (Berl) 80:124–131
Walsh T, Abu Rayan A, Abu Sa’ed J, Shahin H, Shepshelovich J, Lee MK et al (2006) Genomic analysis of a heterogeneous Mendelian phenotype: multiple novel alleles for inherited hearing loss in the Palestinian population. Hum Genomics 2:203–211
Masmoudi S, Antonarakis SE, Schwede T, Ghorbel AM, Gratri M, Pappasavas MP et al (2001) Novel missense mutations of TMPRSS3 in two consanguineous Tunisian families with non-syndromic autosomal recessive deafness. Hum Mutat 18:101–108
Chung J, Park SM, Chang SO, Chung T, Lee KY, Kim AR et al (2014) A novel mutation of TMPRSS3 related to milder auditory phenotype in Korean postlingual deafness: a possible future implication for a personalized auditory rehabilitation. J Mol Med (Berl) 92:651–663
Wattenhofer M, Sahin-Calapoglu N, Andreasen D, Kalay E, Caylan R, Braillard B et al (2005) A novel TMPRSS3 missense mutation in a DFNB8/10 family prevents proteolytic activation of the protein. Hum Genet 117:528–535
Fakim SA, Naderpoor M, Shahidi N, Basharhashemi F, Nejati N, Sakha SH, Alizadeh M (2010) Study of prevalence and causes of hearing loss in high risk neonates admitted to neonatal ward and neonatal intensive care unit. Int Adv Otol 6:365–370
Vozzi D, Morgan A, Vuckovic D, D’Eustacchio A, Abdulhadi K, Rubinato E et al (2014) Hereditary hearing loss: a 96 gene targeted sequencing protocol reveals novel alleles in a series of Italian and Qatari patients. Gene 542:209–216
Gu X, Guo L, Ji H, Sun S, Chai R, Wang L, Li H (2014) Genetic testing for sporadic hearing loss using targeted massively parallel sequencing identifies 10 novel mutations. Clin Genet. doi:10.1111/cge.12431
Weegerink NJ, Schraders M, Oostrik J, Huygen PL, Strom TM, Granneman S (2011) Genotype–phenotype correlation in DFNB8/10 families with TMPRSS3 mutations. J Assoc Res Otolaryngol 12:7537–7566
Ahmed ZM, Li XC, Powell SD, Riazuddin S, Young TL, Ramzan K (2004) Characterization of a new full length TMPRSS3 isoform and identification of mutant alleles responsible for nonsyndromic recessive deafness in Newfoundland and Pakistan. BMC Med Genet 5:24
Lee YJ, Park D, Kim SY, Park WJ (2003) Pathogenic mutations but not polymorphisms in congenital and childhood onset autosomal recessive deafness disrupt the proteolytic activity of TMPRSS3. J Med Genet 40:629–631
Acknowledgments
This work was supported in part by the Slovenian Research Agency grants P3-0343, J3-6798, J3-6800.
Conflict of interest
None declared.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Battelino, S., Klancar, G., Kovac, J. et al. TMPRSS3 mutations in autosomal recessive nonsyndromic hearing loss. Eur Arch Otorhinolaryngol 273, 1151–1154 (2016). https://doi.org/10.1007/s00405-015-3671-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00405-015-3671-0