Abstract
To supply abundant ribosomes, multiple copies of ribosomal RNA genes (rDNA) are conserved from bacterial to human cells. In eukaryotic genomes, clusters of tandemly repeated rDNA units are present, and their number is stably maintained. Due to high level of transcription of rRNA genes, the repetitive structure is prone to rearrangement. In budding yeast, rDNA homeostasis can compensate for this by the regulation of recombination events that will change the copy number. The histone deacetylase Sir2 plays a key role in rDNA copy maintenance and its expression level determines a state of “maintenance” or “amplification” of rDNA copy number. We recently showed that Upstream Activating Factors (UAF) for RNA polymerase I act as a RNA polymerase II repressor of SIR2 transcription in response to rDNA copy loss. Furthermore, the amount of UAF, which is limited in the cell, determines the stable copy number of rDNA and is a molecular switch for rDNA recovery. In this mini-review, we propose a “Musical Chair” model for rDNA copy counting as mediated by UAF and Sir2. The model describes how a straightforward molecular mechanism can account for the “cellular memory” of the proper rDNA copy number.

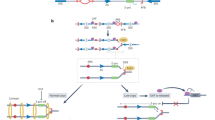
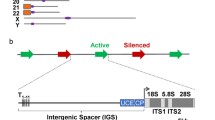
modified from Iida and Kobayashi 2019

Similar content being viewed by others
References
Bierhoff H, Postepska-Igielska A, Grummt I (2014) Noisy silence: non-coding RNA and heterochromatin formation at repetitive elements. Epigenetics 9:53–61. https://doi.org/10.4161/epi.26485
Brewer BJ, Lockshon D, Fangman WL (1992) The arrest of replication forks in the rDNA of yeast occurs independently of transcription. Cell 71:267–276
Burkhalter MD, Sogo JM (2004) rDNA enhancer affects replication initiation and mitotic recombination: Fob1 mediates nucleolytic processing independently of replication. Mol Cell 15:409–421. https://doi.org/10.1016/j.molcel.2004.06.024
Conconi A, Widmer RM, Koller T, Sogo JM (1989) Two different chromatin structures coexist in ribosomal RNA genes throughout the cell cycle. Cell 57:753–761
Dammann R, Lucchini R, Koller T, Sogo JM (1993) Chromatin structures and transcription of rDNA in yeast Saccharomyces cerevisiae. Nucleic Acids Res 21:2331–2338
Dammann R, Lucchini R, Koller T, Sogo JM (1995) Transcription in the yeast rRNA gene locus: distribution of the active gene copies and chromatin structure of their flanking regulatory sequences. Mol Cell Biol 15:5294–5303
French SL, Osheim YN, Cioci F et al (2003) In exponentially growing Saccharomyces cerevisiae cells, rRNA synthesis is determined by the summed RNA polymerase I loading rate rather than by the number of active genes. Mol Cell Biol 23:1558–1568. https://doi.org/10.1128/MCB.23.5.1558-1568.2003
Iida T, Kobayashi T (2019) RNA polymerase I activators count and adjust ribosomal RNA gene copy number. Mol Cell 73:645–654.e13. https://doi.org/10.1016/j.molcel.2018.11.029
Kakui Y, Uhlmann F (2018) SMC complexes orchestrate the mitotic chromatin interaction landscape. Curr Genet 64:335–339. https://doi.org/10.1007/s00294-017-0755-y
Keener J, Dodd JA, Lalo D, Nomura M (1997) Histones H3 and H4 are components of upstream activation factor required for the high-level transcription of yeast rDNA by RNA polymerase I. Proc Natl Acad Sci USA 94:13458–13462
Keys DA, Lee BS, Dodd JA et al (1996) Multiprotein transcription factor UAF interacts with the upstream element of the yeast RNA polymerase I promoter and forms a stable preinitiation complex. Genes Dev 10:887–903
Kobayashi T (2003) The replication fork barrier site forms a unique structure with Fob1p and inhibits the replication fork. Mol Cell Biol 23:9178–9188. https://doi.org/10.1128/MCB.23.24.9178-9188.2003
Kobayashi T, Ganley ARD (2005) Recombination regulation by transcription-induced cohesin dissociation in rDNA repeats. Science 309:1581–1584. https://doi.org/10.1126/science.1116102
Kobayashi T, Heck DJ, Nomura M, Horiuchi T (1998) Expansion and contraction of ribosomal DNA repeats in Saccharomyces cerevisiae: requirement of replication fork blocking (Fob1) protein and the role of RNA polymerase I. Genes Dev 12:3821–3830
Kobayashi T, Horiuchi T, Tongaonkar P et al (2004) SIR2 regulates recombination between different rDNA repeats, but not recombination within individual rRNA genes in yeast. Cell 117:441–453
Kulak NA, Pichler G, Paron I et al (2014) Minimal, encapsulated proteomic-sample processing applied to copy-number estimation in eukaryotic cells. Nat Methods 11:319–324. https://doi.org/10.1038/nmeth.2834
Long EO, Dawid IB (1980) Repeated genes in eukaryotes. Annu Rev Biochem 49:727–764. https://doi.org/10.1146/annurev.bi.49.070180.003455
Siddiqi IN, Dodd JA, Vu L et al (2001) Transcription of chromosomal rRNA genes by both RNA polymerase I and II in yeast uaf30 mutants lacking the 30 kDa subunit of transcription factor UAF. EMBO J 20:4512–4521. https://doi.org/10.1093/emboj/20.16.4512
Smith ML, Cui W, Jackobel AJ et al (2018) Reconstitution of RNA polymerase I Upstream activating factor and the roles of histones H3 and H4 in complex assembly. J Mol Biol 430:641–654. https://doi.org/10.1016/j.jmb.2018.01.003
Takeuchi Y, Horiuchi T, Kobayashi T (2003) Transcription-dependent recombination and the role of fork collision in yeast rDNA. Genes Dev 17:1497–1506. https://doi.org/10.1101/gad.1085403
Tongaonkar P, French SL, Oakes ML et al (2005) Histones are required for transcription of yeast rRNA genes by RNA polymerase I. Proc Natl Acad Sci USA 102:10129–10134. https://doi.org/10.1073/pnas.0504563102
Weitao T, Budd M, Hoopes LLM, Campbell JL (2003) Dna2 helicase/nuclease causes replicative fork stalling and double-strand breaks in the ribosomal DNA of Saccharomyces cerevisiae. J Biol Chem 278:22513–22522. https://doi.org/10.1074/jbc.M301610200
Yang J, Li F (2017) Are all repeats created equal? Understanding DNA repeats at an individual level. Curr Genet 63:57–63. https://doi.org/10.1007/s00294-016-0619-x
Acknowledgements
This work was supported in part by grants-in-aid for Scientific Research (17H01443 to TK and 26640117 to TI) from the Japan Society for the Promotion of Science (JSPS) and Takeda Science Foundation to TK. The authors declare that they have no competing interests.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by M. Kupiec.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Iida, T., Kobayashi, T. How do cells count multi-copy genes?: “Musical Chair” model for preserving the number of rDNA copies. Curr Genet 65, 883–885 (2019). https://doi.org/10.1007/s00294-019-00956-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-019-00956-0