Abstract
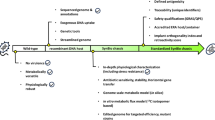
The genus Acinetobacter has been recognized to take up exogenous DNA from the environment. In this study, we conducted natural transformation with a novel diesel-degrading Acinetobacter sp. strain, designated strain DR1, using the broad host range plasmid pRK415. Many factors, including temperature, quantities of DNA, and aeration have proven critically important for efficient natural transformation. Interestingly, the Acinetobacter sp. strain DR1 (pRK415) differed both phenotypically and physiologically from the wild-type strain in several regards, including motility, biofilm formation ability, and responses to oxidative stress: the transformed cells were rendered more sensitive to hydrogen peroxide and cumene hydroperoxide, and their motilities and biofilm formation activity were also attenuated. Our data demonstrated that caution should be exercised when conducting genetic manipulation with plasmids, due to the possibility that phenotypic and physiological changes in the host might occur along with the uptake of plasmids.




Similar content being viewed by others
References
Gerischer U (2008) Acinetobacter molecular biology. Caister Academic Press, Norfolk
Prudhomme M, Attaiech L, Sanchez G et al (2006) Antibiotic stress induces genetic transformability in the human pathogen Streptococcus pneumonia. Science 313:89–92
Johnsborg O, Håvarstein LS (2009) Regulation of natural genetic transformation and acquisition of transforming DNA in Streptococcus pneumonia. FEMS Microbiol Rev 33:627–642
Nielsen KM, Bones AM, Van Elsas JD (1997) Induced natural transformation of Acinetobacter calcoaceticus in soil microcosms. Appl Environ Microbiol 63:3972–3977
Pontiroli A, Rizzi A, Simonet P et al (2009) Visual evidence of horizontal gene transfer between plants and bacteria in the phytosphere of transplastomic tobacco. Appl Environ Microbiol 75:3314–3322
Rizzi A, Pontiroli A, Brusetti L et al (2008) Strategy for in situ detection of natural transformation-based horizontal gene transfer events. Appl Environ Microbiol 74:1250–1254
de Vries J, Herzfeld T, Wackernagel W (2004) Transfer of plastid DNA from tobacco to the soil bacterium Acinetobacter sp. strain by natural transformation. Mol Microbiol 53:323–334
Vaneechoutte M, Young DM, Ornston LN et al (2006) Naturally transformable Acinetobacter sp. strain ADP1 belongs to the newly described species Acinetobacter baylyi. Appl Environ Microbiol 72:932–936
Kang YS, Park W (2010) Trade-off between antibiotic resistance and biological fitness in Acinetobacter sp. strain DR1. Environ Microbiol 12:1304–1318
Kang YS, Park W (2010) Protection against diesel oil toxicity by sodium chloride-induced exopolysaccharides in Acinetobacter sp. strain DR1. J Biosci Bioeng 109:118–123
Yeom J, Jeon CO, Madsen EL, Park W (2009) Ferredoxin-NADP+reductase from Pseudomonas putida functions as a ferric reductase. J Bacteriol 191:1472–1479
Cormack BP, Valdivia RH, Falkow S (1996) FACS-optimized mutants of the green fluorescent protein (GFP). Gene 173:33–38
Andersen JB, Sternberg C, Poulsen LK et al (1998) New unstable variants of green fluorescent protein for studies of transient gene expression in bacteria. Appl Environ Microbiol 64:2240–2246
Yin S, Fuangthong M, Laratta WP, Shapleigh JP (2003) Use of a green fluorescent protein-based reporter fusion for detection of nitric oxide produced by denitrifiers. Appl Environ Microbiol 699:3938–3944
Ray JL, Nielsen KM (2005) Experimental methods for assaying natural transformation and inferring horizontal gene transfer. Methods Enzymol 395:491–520
Kim YH, Lee Y, Kim S et al (2006) The role of periplasmic antioxidant enzymes (superoxide dismutase and thiol peroxidase) of the Shiga toxin-producing Escherichia coli O157:H7 in the formation of biofilms. Proteomics 6:6189–6193
Lee Y, Oh S, Park W (2009) Inactivation of the Pseudomonas putida KT2440 dsbA gene promotes extracellular matrix production and biofilm formation. FEMS Microbiol Lett 297:38–48
Kohanski MA, DePristo MA, Collins JJ (2010) Sublethal antibiotic treatment leads to multidrug resistance via radical-induced mutagenesis. Mol Cell 37:311–320
Baquero F, Martínez JL, Cantón R (2008) Antibiotics and antibiotic resistance in water environments. Curr Opin Biotechnol 19:260–265
Chee-Sanford JC, Mackie RI, Koike S et al (2009) Fate and transport of antibiotic residues and antibiotic resistance genes following land application of manure waste. J Environ Qual 38:1086–1108
Mackie RI, Koike S, Krapac I et al (2006) Tetracycline residues and tetracycline resistance genes in groundwater impacted by swine production facilities. Anim Biotechnol 17:157–176
Juhas M, van der Meer JR, Gaillard M et al (2009) Genomic islands: tools of bacterial horizontal gene transfer and evolution. FEMS Microbiol Rev 33:376–393
Laskaris P, Tolba S, Calvo-Bado L, Wellington L (2010) Coevolution of antibiotic production and counter-resistance in soil bacteria. Environ Microbiol 12:783–796
Eckert B, Beck CF (1989) Overproduction of transposon Tn10-encoded tetracycline resistance protein results in cell death and loss of membrane potential. J Bacteriol 171:3557–3559
Nguyen TN, Phan QG, Duong LP et al (1989) Effects of carriage and expression of the Tn10 tetracycline-resistance operon on the fitness of Escherichia coli K12. Mol Biol Evol 6:213–225
Acknowledgment
This study was supported by a grant from the MEST/NRF to the Environment Biotechnology National Core Research Center (grant #: 20090091491), Korea.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Park, J., Park, W. Phenotypic and Physiological Changes in Acinetobacter sp. Strain DR1 with Exogenous Plasmid. Curr Microbiol 62, 249–254 (2011). https://doi.org/10.1007/s00284-010-9698-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-010-9698-y