Abstract
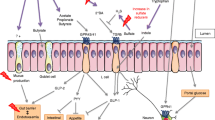
The human gut harbors a complex microbial community that performs a range of metabolic, physiological, and immunological functions. The host and its inhabiting microorganisms are often referred to as a “superorganism.” Dysbiosis of gut microflora has been associated with the pathogenesis of intestinal disorders including inflammatory bowel disease, colorectal cancer, and extra-intestinal disorders such as cardiovascular disease. Therefore, gut microbiome interventions are important for the prevention and treatment of diseases. However, ethical, economic, scientific, and time constraints limit the outcome of human intervention or animal studies targeting gut microbiota. We recently developed an in vitro batch fermentation model (the Kobe University Human Intestinal Microbiota Model, KUHIMM) that is capable of hosting a majority of gut microbial species in humans and also detects the metabolites produced by microorganisms in real time. In this mini review, we elucidated the characteristics of the KUHIMM and its applicability in analyzing the effect of diet, drugs, probiotics, and prebiotics on intestinal bacteria. In addition, we introduce as examples its application to disease models, such as ulcerative colitis, in which intestinal bacteria are intricately involved in the process of pathogenesis. We also discuss the potential of the KUHIMM in precision medicine.
Key points
• In vitro gut fermentation model to simulate human colonic microbiota
• Screening of potential prebiotics and probiotic candidates in healthy model
• Construction of disease models of ulcerative colitis and coronary artery disease
Graphical abstract



Similar content being viewed by others
Data Availability
All raw sequence data generated in this study have been deposited in MG-RAST as “Model Culture System of Human Colonic Microbiota_pH control” under the accession numbers mgm4921743.3–mgm4921745.3.
Code availability
Not applicable.
References
Aguirre M, Eck A, Koenen ME, Savelkoul PHM, Budding AE, Venema K (2016) Diet drives quick change in the metabolic activity and composition of human gut microbiota in a validated in vitro gut model. Res Microbiol 167:114–125. https://doi.org/10.1016/j.resmic.2015.09.006
Bambery B, Selgelid M, Weijer C, Savulescu J, Pollard AJ (2016) Ethical criteria for human challenge studies in infectious diseases. Public Health Ethics 9:92–103. https://doi.org/10.1093/phe/phv026
Carding S, Verbeke K, Vipond DT, Corfe BM, Owen LJ (2015) Dysbiosis of the gut microbiota in disease. Microb Ecol Health Dis 26:26191. https://doi.org/10.3402/mehd.v26.26191
Costello SP, Hughes PA, Waters O, Bryant RV, Vincent AD, Blatchford P, Katsikeros R, Makanyanga J, Campaniello MA, Mavrangelos C, Rosewarne CP, Bickley C, Peters C, Schoeman MN, Conlon MA, Roberts-Thomson IC, Andrews JM (2019) Effect of fecal microbiota transplantation on 8-week remission in patients with ulcerative colitis: a randomized clinical trial. JAMA 321:156–164. https://doi.org/10.1001/jama.2018.20046
Cummings JH, Pomare EW, Branch WJ, Naylor CP, Macfarlane GT (1987) Short chain fatty acids in human large intestine, portal, hepatic and venous blood. Gut 28:1221–1227. https://doi.org/10.1136/gut.28.10.1221
den Besten G, van Eunen K, Groen AK, Venema K, Reijngoud DJ, Bakker BM (2013) The role of short-chain fatty acids in the interplay between diet, gut microbiota, and host energy metabolism. J Lipid Res 54:2325–2340. https://doi.org/10.1194/jlr.R036012
Doré J, Simrén M, Buttler L, Guarner F (2013) Hot topics in gut microbiota. United European Gastroenterol J 1:311–318. https://doi.org/10.1177/2050640613502477
Fallingborg J (1999) Intraluminal pH of the human gastrointestinal tract. Dan Med Bull 46:183–196
Feagan BG, Rutgeerts P, Sands BE, Hanauer S, Colombel JF, Sandborn WJ, Van Assche G, Axler J, Kim HJ, Danese S, Fox I, Milch C, Sankoh S, Wyant T, Xu J, Parikh A, GEMINI 1 Study Group (2013) Vedolizumab as induction and maintenance therapy for ulcerative colitis. N Engl J Med 369:699–710. https://doi.org/10.1056/NEJMoa1215734
Frank DN, St Amand AL, Feldman RA, Boedeker EC, Harpaz N, Pace NR (2007) Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc Natl Acad Sci U S A 104:13780–13785. https://doi.org/10.1073/pnas.0706625104
Franke A, Balschun T, Sina C, Ellinghaus D, Häsler R, Mayr G, Albrecht M, Wittig M, Buchert E, Nikolaus S, Gieger C, Wichmann HE, Sventoraityte J, Kupcinskas L, Onnie CM, Gazouli M, Anagnou NP, Strachan D, McArdle WL, Mathew CG, Rutgeerts P, Vermeire S, Vatn MH, IBSEN study group, Krawczak M, Rosenstiel P, Karlsen TH, Schreiber S (2010) Genome-wide association study for ulcerative colitis identifies risk loci at 7q22 and 22q13 (IL17REL). Nat Genet 42:292–294. https://doi.org/10.1038/ng.553
Hatanaka M, Morita H, Aoyagi Y, Sasaki K, Sasaki D, Kondo A, Nakamura T (2020) Effective bifidogenic growth factors cyclo-Val-Leu and cyclo-Val-Ile produced by Bacillus subtilis C-3102 in the human colonic microbiota model. Sci Rep 10:7591. https://doi.org/10.1038/s41598-020-64374-w
Jahani-Sherafat S, Alebouyeh M, Moghim S, Amoli HA, Ghasemian-Safaei H (2018) Role of gut microbiota in the pathogenesis of colorectal cancer; a review article. Gastroenterol Hepatol Bed Bench 11:101–109 https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5990924
Javdan B, Lopez JG, Chankhamjon P, Lee YCJ, Hull R, Wu Q, Wang X, Chatterjee S, Donia MS (2020) Personalized mapping of drug metabolism by the human gut microbiome. Cell 181:1661–1679. https://doi.org/10.1016/j.cell.2020.05.001
Jostins L, Ripke S, Weersma RK, Duerr RH, McGovern DP, Hui KY, Lee JC, Schumm LP, Sharma Y, Anderson CA, Essers J, Mitrovic M, Ning K, Cleynen I, Theatre E, Spain SL, Raychaudhuri S, Goyette P, Wei Z, Abraham C, Achkar JP, Ahmad T, Amininejad L, Ananthakrishnan AN, Andersen V, Andrews JM, Baidoo L, Balschun T, Bampton PA, Bitton A, Boucher G, Brand S, Büning C, Cohain A, Cichon S, D'Amato M, De Jong D, Devaney KL, Dubinsky M, Edwards C, Ellinghaus D, Ferguson LR, Franchimont D, Fransen K, Gearry R, Georges M, Gieger C, Glas J, Haritunians T, Hart A, Hawkey C, Hedl M, Hu X, Karlsen TH, Kupcinskas L, Kugathasan S, Latiano A, Laukens D, Lawrance IC, Lees CW, Louis E, Mahy G, Mansfield J, Morgan AR, Mowat C, Newman W, Palmieri O, Ponsioen CY, Potocnik U, Prescott NJ, Regueiro M, Rotter JI, Russell RK, Sanderson JD, Sans M, Satsangi J, Schreiber S, Simms LA, Sventoraityte J, Targan SR, Taylor KD, Tremelling M, Verspaget HW, De Vos M, Wijmenga C, Wilson DC, Winkelmann J, Xavier RJ, Zeissig S, Zhang B, Zhang CK, Zhao H, International IBD Genetics Consortium (IIBDGC), Silverberg MS, Annese V, Hakonarson H, Brant SR, Radford-Smith G, Mathew CG, Rioux JD, Schadt EE, Daly MJ, Franke A, Parkes M, Vermeire S, Barrett JC, Cho JH (2012) Host-microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491:119–124. https://doi.org/10.1038/nature11582
Kazemian N, Mahmoudi M, Halperin F, Wu JC, Pakpour S (2020) Gut microbiota and cardiovascular disease: opportunities and challenges. Microbiome 8:36. https://doi.org/10.1186/s40168-020-00821-0
Kobayashi T, Siegmund B, Le Berre C, Wei SC, Ferrante M, Shen B, Bernstein CN, Danese S, Peyrin-Biroulet L, Hibi T (2020) Ulcerative colitis. Nat Rev Dis Primers 6:74. https://doi.org/10.1038/s41572-020-0205-x
Lin CS, Chang CJ, Lu CC, Martel J, Ojcius DM, Ko YF, Young JD, Lai HC (2014) Impact of the gut microbiota, prebiotics, and probiotics on human health and disease. Biomed J 37:259–268. https://doi.org/10.4103/2319-4170.138314
Litvak Y, Byndloss MX, Bäumler AJ (2018) Colonocyte metabolism shapes the gut mictobiota. Science 363:eaat9076. https://doi.org/10.1126/science.aat9076
Machiels K, Joossens M, Sabino J, De Preter V, Arijs I, Eeckhaut V, Ballet V, Claes K, Van Immerseel F, Verbeke K, Ferrante M, Verhaegen J, Rutgeerts P, Vermeire S (2014) A decrease of the butyrate-producing species Roseburia hominis and Faecalibacterium prausnitzii defines dysbiosis in patients with ulcerative colitis. Gut 63:1275–1283. https://doi.org/10.1136/gutjnl-2013-304833
Mak IWY, Evaniew N, Ghert M (2014) Lost in translation: animal models and clinical trials in cancer treatment. Am J Transl Res 6:114–118
Malakar AK, Choudhury D, Halder B, Paul P, Uddinf A, Chakraborty S (2019) A review on coronary artery disease, its risk factors, and therapeutics. J Cell Physiol 234:16812–16823. https://doi.org/10.1002/jcp.28350
Mussatto SI, Mancilha IM (2007) Non-digestible oligosaccharides: a review. Carbohydr Polym 68:587–597
Nihei N, Okamoto H, Furune T, Furune T, Ikuta N, Sasaki K, Rimbach G, Yoshikawa Y, Terao K (2018) Dietary α-cyclodextrin modifies gut microbiota and reduces fat accumulation in high-fat-diet-fed obese mice. Biofactors 336–347. https://doi.org/10.1002/biof.1429
Nishida A, Inoue R, Inatomi O, Bamba S, Naito Y, Andoh A (2018) Gut microbiota in the pathogenesis of inflammatory bowel disease. Clin J Gastroenterol 11:1–10. https://doi.org/10.1007/s12328-017-0813-5
Nissen L, Casciano F, Gianotti A (2020) Intestinal fermentation in vitro models to study food-induced gut microbiota shift: an updated review. FEMS Microbiol Lett 367:fnaa097. https://doi.org/10.1093/femsle/fnaa097
Oba S, Sunagawa T, Tanihiro R, Awashima K, Sugiyama H, Odani T, Nakamura Y, Kondo A, Sasaki D, Sasaki K (2020) Prebiotic effects of yeast mannan, which selectively promotes Bacteroides thetaiotaomicron and Bacteroides ovatus in a human colonic microbiota model. Sci Rep 10:17351. https://doi.org/10.1038/s41598-020-74379-0
Okahara K, Ishikawa D, Nomura K, Ito S, Haga K, Takahashi M, Shibuya T, Osada T, Nagahara A (2020) Matching between donors and ulcerative colitis patients is important for long-term maintenance after fecal microbiota transplantation. J Clin Med 9:1650. https://doi.org/10.3390/jcm9061650
Paramsothy S, Kamm MA, Kaakoush NO, Walsh AJ, van den Bogaerde J, Samuel D, Leong RWL, Connor S, Ng W, Paramsothy R, Xuan W, Lin E, Mitchell HM, Borody TJ (2017) Multidonor intensive faecal microbiota transplantation for active ulcerative colitis: a randomised placebo-controlled trial. Lancet 389:1218–1228. https://doi.org/10.1016/S0140-6736(17)30182-4
Payne AN, Zinhler A, Chassard C, Lacroix C (2012) Advances and perspectives in in vitro human gut fermentation modeling. Trends Biotechnol 30:17–25. https://doi.org/10.1016/j.tibtech.2011.06.011
Ran Z, Wu K, Matsuoka K, Jeen YT, Wei SC, Ahuja V, Chen M, Hu PJ, Andoh A, Kim HJ, Yang SK, Watanabe M, Ng SC, Hibi T, Hilmi IN, Suzuki Y, Han DS, Leung WK, Sollano J, Ooi CJ, Qian J (2020) Asian organization for Crohn’s and Colitis and Asia Pacific association of gastroenterology practice recommendations for medical management and monitoring of inflammatory bowel disease in Asia. J Gastroenterol Hapatol. https://doi.org/10.1111/jgh.15185
Reid G, Jass J, Sebulsky MT, McCormick JK (2003) Potential uses of probiotics in clinical practice. Clin Microbiol Rev 16:658–672
Reinisch W, Sandborn WJ, Hommes DW, D'Haens G, Hanauer S, Schreiber S, Panaccione R, Fedorak RN, Tighe MB, Huang B, Kampman W, Lazar A, Thakkar R (2011) Adalimumab for induction of clinical remission in moderately to severely active ulcerative colitis: results of a randomised controlled trial. Gut 60:780–787. https://doi.org/10.1136/gut.2010.221127
Rivière A, Selak M, Lantin D, Leroy F, Vuyst LD (2016) Bifidobacteria and butyrate-producing colon bacteria: importance and strategies for their stimulation in the human gut. Front Microbiol 7:979. https://doi.org/10.1126/science.aat9076
Rubtsov YP, Rasmussen JP, Chi EY, Fontenot J, Castelli L, Ye X, Treuting P, Siewe L, Roers A, Henderson WR Jr, Muller W, Rudensky AY (2008) Regulatory T cell-derived interleukin-10 limits inflammation at environmental interfaces. Immunity 28:546–558. https://doi.org/10.1016/j.immuni.2008.02.017
Rutgeerts P, Sandborn WJ, Feagan BG, Reinisch W, Olson A, Johanns J, Travers S, Rachmilewitz D, Hanauer SB, Lichtenstein GR, de Villiers WJ, Present D, Sands BE, Colombel JF (2005) Infliximab for induction and maintenance therapy for ulcerative colitis. N Engl J Med 353:2462–2476. https://doi.org/10.1056/NEJMoa050516
Sandborn WJ, Su C, Sands BE, D'Haens GR, Vermeire S, Schreiber S, Danese S, Feagan BG, Reinisch W, Niezychowski W, Friedman G, Lawendy N, Yu D, Woodworth D, Mukherjee A, Zhang H, Healey P, Panés J; OCTAVE Induction 1, OCTAVE Induction 2, and OCTAVE Sustain Investigator (2017) Tofacitinib as induction and maintenance therapy for ulcerative colitis. N Engl J Med 376:1723–1736. https://doi.org/10.1056/NEJMoa1606910
Sands BE, Sandborn WJ, Panaccione R, O'Brien CD, Zhang H, Johanns J, Adedokun OJ, Li K, Peyrin-Biroulet L, Van Assche G, Danese S, Targan S, Abreu MT, Hisamatsu T, Szapary P, Marano C, UNIFI Study Group (2019) Ustekinumab as induction and maintenance therapy for ulcerative colitis. N Engl J Med 381:1201–1214. https://doi.org/10.1056/NEJMoa1900750
Sasaki K, Sasaki D, Okai N, Tanaka K, Nomoto R, Fukuda I, Yoshida K, Kondo A, Osawa R (2017) Taurine does not affect the composition, diversity, or metabolism of human colonic microbiota simulated in a single-batch fermentation system. PLoS ONE 12:e0180991. https://doi.org/10.1371/journal.pone.0180991
Sasaki D, Sasaki K, Ikuta N, Yasuda T, Fukuda I, Kondo A, Osawa R (2018) Low amounts of dietary fibre increase in vitro production of short-chain fatty acids without changing human colonic microbiota structure. Sci Rep 8:435. https://doi.org/10.1038/s41598-017-18877-8
Sasaki D, Sasaki K, Kadowaki Y, Aotsuka Y, Kondo A (2019a) Bifidogenic and butyrogenic effects of young barley leaf extract in an in vitro human colonic microbiota model. AMB Expr 9:182. https://doi.org/10.1186/s13568-019-0911-5
Sasaki K, Inoue J, Sasaki D, Hoshi N, Shirai T, Fukuda I, Azuma T, Kondo A, Osawa R (2019b) Construction of a model culture system of human colonic microbiota to detect decreased Lachnospiraceae abundance and butyrogenesis in the feces of ulcerative colitis patients. Biotechnol J 14:e1800555. https://doi.org/10.1002/biot.201800555
Sasaki D, Sasaki K, Kondo A (2020a) Glycosidic linkage structures influence dietary fiber fermentability and propionate production by human colonic microbiota in vitro. Biotechnol J 1900523. https://doi.org/10.1002/biot.201900523
Sasaki K, Sasaki D, Inoue J, Hoshi N, Maeda T, Yamada R, Kondo A (2020b) Bacillus coagulans SANK 70258 suppresses Enterobacteriaceae in the microbiota of ulcerative colitis in vitro and enhances butyrogenesis in healthy microbiota. Appl Microbiol Biotechnol 104:3859–3867. https://doi.org/10.1007/s00253-020-10506-1
Scott KP, Antoine JM, Midtvedt T, van Hemert S (2015) Manipulating the gut microbiota to maintain health and treat disease. Microb Ecol Health Dis 26:25877. https://doi.org/10.3402/mehd.v26.25877
Sender R, Fuchs S, Miho R (2016) Revised estimates for the number of human and bacterial cells in the body. PLoS Biol 14:e1002533. https://doi.org/10.1371/journal.pbio.1002533
Silverberg MS, Cho JH, Rioux JD, McGovern DP, Wu J, Annese V, Achkar JP, Goyette P, Scott R, Xu W, Barmada MM, Klei L, Daly MJ, Abraham C, Bayless TM, Bossa F, Griffiths AM, Ippoliti AF, Lahaie RG, Latiano A, Paré P, Proctor DD, Regueiro MD, Steinhart AH, Targan SR, Schumm LP, Kistner EO, Lee AT, Gregersen PK, Rotter JI, Brant SR, Taylor KD, Roeder K, Duerr RH (2009) Ulcerative colitis-risk loci on chromosomes 1p36 and 12q15 found by genome-wide association study. Nat Genet 41:216–220. https://doi.org/10.1038/ng.275
Sood A, Mahajan R, Singh A, Midha V, Mehta V, Narang V, Singh T, Singh Pannu A (2019) Role of faecal microbiota transplantation for maintenance of remission in patients with ulcerative colitis: a pilot study. J Crohns Colitis 13:1311–1317. https://doi.org/10.1093/ecco-jcc/jjz060
Strober W, Fuss IJ, Blumberg RS (2002) The immunology of mucosal models of inflammation. Annu Rev Immunol 20:495–549. https://doi.org/10.1146/annurev.immunol.20.100301.064816
Sun L, Ma L, Ma Y, Zhang F, Zhao C, Nie Y (2018) Insights into the role of gut microbiota in obesity: pathogenesis, mechanisms, and therapeutic perspectives. Protein Cell 9:397–403. https://doi.org/10.1007/s13238-018-0546-3
Takagi R, Sasaki K, Sasaki D, Fukuda I, Tanaka K, Yoshida K, Kondo A, Osawa R (2016) A single-batch fermentation system to simulate human colonic microbiota for high-throughput evaluation of prebiotics. PLoS ONE 11:e0160533. https://doi.org/10.1371/journal.pone.0160533
Takimoto T, Hatanaka M, Hoshino T, Takara T, Tanaka K, Shimizu A, Motira H, Nakamura T (2018) Effect of Bacillus subtilis C-3102 on bone mineral density in healthy postmenopausal Japanese women: a randomized, placebo-controlled, double-blind clinical trial. Biosci Microbiota Food Health 37:87–96. https://doi.org/10.12938/bmfh.18-006
Thursby E, Juge N (2017) Introduction to the human gut microbiota. Biochem J 474:1823–1836. https://doi.org/10.1042/BCJ20160510
Tsitko I, Wiik-Miettinen F, Mattila O, Rosa-Sibakov N, Seppänen-Laakso T, Maukonen J, Nordlund E, Saarela M (2019) A small in vitro fermentation model for screening the gut microbiota effects of different fiber preparations. Int J Mol Sci 20:1925. https://doi.org/10.3390/ijms20081925
Van den Abbeele P, Grootaert C, Marzorati M, Possemiers S, Verstraete W, Gérard P, Rabot S, Bruneau A, Aidy SE, Derrien M, Zoetendal E, Kleerebezem M, Smidt H, Van de Wiele T (2010) Microbial community development in a dynamic gut model is reproducible, colon region specific, and selective for Bacteroidetes and Clostridium cluster IX. Appl Environ Microbiol 76:5237–5246. https://doi.org/10.1128/AEM.00759-10
Venema K, van den Abbeele P (2013) Experimental models of the gut microbiome. Best Prac Res Clin Gastroenterol 27:115–126. https://doi.org/10.1016/j.bpg.2013.03.002
Venema K, van Nuenen HMC, Smeets-Peeters M, Minekus M, Havenaar R (2000) TNO’s in vitro large intestinal model: an excellent screening tool for functional food and pharmaceutical research. Ernährung/Nutrition 24:558–564
Vermeiren J, Van den Abbeele P, Laukens D, Vigsnaes LK, De Vos M, Boon N, Van de Wiele T (2012) Decreased colonization of fecal Clostridium coccoides/Eubacterium rectale species from ulcerative colitis patients in an in vitro dynamic gut model with mucin environment. FEMS Microbiol Ecol 79:685–696. https://doi.org/10.1111/j.1574-6941.2011.01252.x
von Martels JZH, Sadabad MS, Bourgonje AR, Blokzijl T, Dijkstra G, Faber KN, Harmsen HJM (2017) The role of gut microbiota in health and disease: In vitro modeling of host-microbe interactions at the aerobe-anaerobe interphase of the human gut. Anaerobe 44:3–12. https://doi.org/10.1016/j.anaerobe.2017.01.001
Yoshida N, Emoto T, Yamashita T, Watanabe H, Hayashi T, Tabata T, Hoshi N, Hatano N, Ozawa G, Sasaki N, Mizoguchi T, Amin HZ, Hirota Y, Ogawa W, Yamada T, Hirata K (2018) Bacteroides vulgatus and Bacteroides dorei reduce gut microbial lipopolysaccharide production and inhibit atherosclerosis. Circulation 138:2486–2498. https://doi.org/10.1161/CIRCULATIONAHA.118.033714
Yoshida N, Sasaki K, Sasaki D, Yamashita T, Fukuda H, Hayashi T, Tabata T, Osawa R, Hirata K, Kondo A (2019) Effect of resistant starch on the gut microbiota and its metabolites in patients with coronary artery disease. J Artheroscler Thromb 26:705–719. https://doi.org/10.5551/jat.47415
Yoshida N, Yamashita T, Kishino S, Watanabe H, Sasaki K, Sasaki D, Tabata T, Sugiyama Y, Kitamura N, Saito Y, Emoto T, Hayashi T, Takahashi T, Sinohara M, Osawa R, Kondo A, Yamada T, Ogawa J, Hirata K (2020) A possible beneficial effect of Bacteroides on faecal lipopolysaccharide activity and cardiovascular diseases. Sci Rep 10:13009. https://doi.org/10.1038/s41598-020-69983-z
Acknowledgements
We are grateful to Ayami Fujino, Yasuko koura, Misa Yasuda, Miki Ishizaki, Yoko Sugita, and Kimiko Enda (Kobe University) for their assistance.
Funding
This study was partly funded by the Japan Society for the Promotion of Science (JSPS) (KAKENHI grant number 18K05487).
Author information
Authors and Affiliations
Contributions
NH and KS conceived and designed research. JI and DS conducted experiments and analyzed data. NH and KS wrote the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Ethics approval
Human subjects provided written informed consent prior to specimen collection. The study was performed in accordance with the guideline of Kobe University Hospital and was approved by the Institutional Ethics Review Board of Kobe University. All methods used in this study were in accordance with the Declaration of Helsinki.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Hoshi, N., Inoue, J., Sasaki, D. et al. The Kobe University Human Intestinal Microbiota Model for gut intervention studies. Appl Microbiol Biotechnol 105, 2625–2632 (2021). https://doi.org/10.1007/s00253-021-11217-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-021-11217-x