Abstract
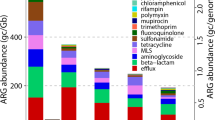
Studies on antibiotic production wastewater have shown that even a single antibiotic can select for multidrug resistant bacteria in aquatic environments. It is speculated that plasmids are an important mechanism of multidrug resistance (MDR) under high concentrations of antibiotics. Herein, two metagenomic libraries were constructed with plasmid DNA extracted from cultivable microbial communities in a biological wastewater treatment reactor supplemented with 0 (CONTROL) or 25 mg/L of oxytetracycline (OTC-25). The OTC-25 plasmidome reads were assigned to 72 antibiotic resistance genes (ARGs) conferring resistance to 13 types of antibiotics. Dominant ARGs, encoding resistance to tetracycline, aminoglycoside, sulfonamide, and multidrug resistance genes, were enriched in the plasmidome under 25 mg/L of oxytetracycline. Furthermore, 17 contiguous multiple-ARG carrying contigs (carrying ≥ 2 ARGs) were discovered in the OTC-25 plasmidome, whereas only nine were found in the CONTROL. Mapping of the OTC-25 plasmidome reads to completely sequenced plasmids revealed that the conjugative IncU resistance plasmid pFBAOT6 of Aeromonas caviae, carrying multidrug resistance transporter (pecM), tetracycline resistance genes (tetA, tetR), and transposase genes, might be a potential prevalent resistant plasmid in the OTC-25 plasmidome. Additionally, two novel resistant plasmids (containing contig C301682 carrying multidrug resistant operon mexCD-oprJ and contig C301632 carrying the tet36 and transposases genes) might also be potential prevalent resistant plasmids in the OTC-25 plasmidome. This study will be helpful to better understand the role of plasmids in the development of MDR in water environments under high antibiotic concentrations.


Similar content being viewed by others
References
Adams CA, Austin B, Meaden PG, McIntosh D (1998) Molecular characterization of plasmid-mediated Oxytetracycline resistance in Aeromonas salmonicida. Appl Environ Microbiol 64(11):4194–4201
Altman D (1991) Practical statistics for medical research. Chapman and Hall, London
Andersen SR, Sandaa RA (1994) Distribution of tetracycline resistance determinants among gram-negative bacteria isolated from polluted and unpolluted marine sediments. Appl Environ Microbiol 60(3):908–912
Barnett DW, Garrison EK, Quinlan AR, Strömberg MP, Marth GT (2011) BamTools: a C++ API and toolkit for analyzing and managing BAM files. Bioinformatics 27(12):1691–1692. https://doi.org/10.1093/bioinformatics/btr174
Bennett PM (2008) Plasmid encoded antibiotic resistance: acquisition and transfer of antibiotic resistance genes in bacteria. Br J Pharmacol 153(S1):S347–S357. https://doi.org/10.1038/sj.bjp.0707607
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30(15):2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Gordon JI (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7(5):335–336. https://doi.org/10.1038/nmeth.f.303
Chagas TP, Seki LM, Cury JC, Oliveira JA, Dávila AM, Silva DM, Asensi MD (2011) Multiresistance, beta-lactamase-encoding genes and bacterial diversity in hospital wastewater in Rio de Janeiro, Brazil. J Appl Microbiol 111(3):572–581. https://doi.org/10.1111/j.1365-2672.2011.05072.x
Chang G, Roth CB (2001) Structure of MsbA from E. coli: a homolog of the multidrug resistance ATP binding cassette (ABC) transporters. Science 293(5536):1793–1800. https://doi.org/10.1126/science.293.5536.1793
Chen Y-T, Shu H-Y, Li L-H, Liao T-L, Wu K-M, Shiau Y-R, Yan J-J, Su I-J, Tsai S-F, Lauderdale T-L (2006) Complete nucleotide sequence of pK245, a 98-kilobase plasmid conferring quinolone resistance and extended-spectrum-β-lactamase activity in a clinical Klebsiella pneumoniae isolate. Antimicrob Agents Chemother 50(11):3861–3866. https://doi.org/10.1128/aac.00456-06
Courvalin P (1994) Transfer of antibiotic resistance genes between gram-positive and gram-negative bacteria. Antimicrob Agents Chemother 38(7):1447–1451. https://doi.org/10.1128/AAC.38.7.1447
Dawson RJ, Locher KP (2006) Structure of a bacterial multidrug ABC transporter. Nature 443(7108):180–185. https://doi.org/10.1038/nature05155
DePaola A, Flynn PA, McPhearson RM, Levy SB (1988) Phenotypic and genotypic characterization of tetracycline- and oxytetracycline-resistant Aeromonas hydrophila from cultured channel catfish (Ictalurus punctatus) and their environments. Appl Environ Microbiol 54(7):1861–1863
Doggett NA, Stubben CJ, Chertkov O, Bruce DC, Detter JC, Johnson SL, Han CS (2013) Complete genome sequence of Bacillus thuringiensis Serovar Israelensis strain HD-789. Genome Announcements 1(6):e01023–e01013. https://doi.org/10.1128/genomeA.01023-13
Frost LS, Leplae R, Summers AO, Toussaint A (2005) Mobile genetic elements: the agents of open source evolution. Nat Rev Microbiol 3(9):722–732. https://doi.org/10.1038/nrmicro1235
Glavinas H, Krajcsi P, Cserepes J, Sarkadi B (2004) The role of ABC transporters in drug resistance, metabolism and toxicity. Current drug delivery 1(1):27–42. https://doi.org/10.2174/1567201043480036
Guerra B, Soto SM, Argüelles JM, Mendoza MC (2001) Multidrug resistance is mediated by large plasmids carrying a class 1 integron in the emergent Salmonella enterica serotype. Antimicrob Agents Chemother 45(4):1305–1308. https://doi.org/10.1128/aac.45.4.1305-1308.2001
Hinse D, Vollmer T, Rückert C, Blom J, Kalinowski J, Knabbe C, Dreier J (2011) Complete genome and comparative analysis of Streptococcus gallolyticus subsp. gallolyticus, an emerging pathogen of infective endocarditis. BMC Genomics 12(1):400. https://doi.org/10.1186/1471-2164-12-400
Kav AB, Sasson G, Jami E, Doron-Faigenboim A, Benhar I, Mizrahi I (2012) Insights into the bovine rumen plasmidome. Proc Natl Acad Sci U S A 109(14):5452–5457. https://doi.org/10.1073/pnas.1116410109
Kristiansson E, Fick J, Janzon A, Grabic R, Rutgersson C, Weijdegård B, Söderström H, Larsson DJ (2011) Pyrosequencing of antibiotic-contaminated river sediments reveals high levels of resistance and gene transfer elements. PLoS One 6(2):e17038. https://doi.org/10.1371/journal.pone.0017038
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. John Wiley, Chichester, pp 115–175
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9(4):357–359. https://doi.org/10.1038/nmeth.1923
Levy SB (1998) The challenge of antibiotic resistance. Sci Am 278(278):46–53. https://doi.org/10.1038/scientificamerican0398-46
Levy SB, Marshall B (2004) Antibacterial resistance worldwide: causes, challenges and responses. Nat Med 10(12):122–129
Levy SB, Fitzgerald GB, Macone AB (1976) Changes in intestinal flora of farm personnel after introduction of a tetracycline-supplemented feed on a farm. New Engl J Med 295(11):583–588. https://doi.org/10.1056/NEJM197609092951103
Li X-Z, Barré N, Poole K (2000) Influence of the MexA-MexB-OprM multidrug efflux system on expression of the MexC-MexD-OprJ and MexE-MexF-OprN multidrug efflux systems in Pseudomonas aeruginosa. J Antimicrob Chemother 46(6):885–893. https://doi.org/10.1093/jac/46.6.885
Li D, Yang M, Hu J, Zhang J, Liu R, Gu X, Zhang Y, Wang Z (2009a) Antibiotic-resistance profile in environmental bacteria isolated from penicillin production wastewater treatment plant and the receiving river. Environ Microbiol 11(6):1506–1517. https://doi.org/10.1111/j.1462-2920.2009.01878.x
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R (2009b) The sequence alignment/map format and SAMtools. Bioinformatics 25(16):2078–2079. https://doi.org/10.1093/bioinformatics/btp352
Li D, Yu T, Zhang Y, Yang M, Li Z, Liu M, Qi R (2010) Antibiotic resistance characteristics of environmental bacteria from an oxytetracycline production wastewater treatment plant and the receiving river. Appl Environ Microbiol 76(11):3444–3451. https://doi.org/10.1128/AEM.02964-09
Li A-D, Li L-G, Zhang T (2015) Exploring antibiotic resistance genes and metal resistance genes in plasmid metagenomes from wastewater treatment plants. Front Microbiol 6:1025. https://doi.org/10.3389/fmicb.2015.01025
Lin Y, Li J, Shen H, Zhang L, Papasian CJ (2011) Comparative studies of de novo assembly tools for next-generation sequencing technologies. Bioinformatics 27(15):2031–2037. https://doi.org/10.1093/bioinformatics/btr319
Liu M, Zhang Y, Yang M, Tian Z, Ren L, Zhang S (2012) Abundance and distribution of tetracycline resistance genes and mobile elements in an oxytetracycline production wastewater treatment system. Environ Sci Technol 46(14):7551–7557. https://doi.org/10.1021/es301145m
Lubelski J, Konings WN, Driessen AJ (2007) Distribution and physiology of ABC-type transporters contributing to multidrug resistance in bacteria. Microbiol Mol Biol Rev 71(3):463–476. https://doi.org/10.1128/MMBR.00001-07
Ma Y, Paulsen IT, Palenik B (2012) Analysis of two marine metagenomes reveals the diversity of plasmids in oceanic environments. Environ Microbiol 14(2):453–466. https://doi.org/10.1111/j.1462-2920.2011.02633.x
Ma L, Xia Y, Li B, Yang Y, Li L-G, Tiedje JM, Zhang T (2016) Metagenomic assembly reveals hosts of antibiotic resistance genes and the shared resistome in pig, chicken, and human feces. Environ Sci Technol 50(1):420–427. https://doi.org/10.1021/acs.est.5b03522
Mazel D (2006) Integrons: agents of bacterial evolution. Nat Rev Microbiol 4(8):608–620. https://doi.org/10.1038/nrmicro1462
Mazel D, Davies J (1999) Antibiotic resistance in microbes. Cell Mol Life Sci 56(9):742–754. https://doi.org/10.1007/s000180050021
Møller J, Bak AL, Stenderup A, Zachariae H, Afzelius H (1977) Changing patterns of plasmid-mediated drug resistance during tetracycline therapy. Antimicrob Agents Chemother 11(3):388–391. https://doi.org/10.1128/AAC.11.3.388
Morita Y, Murata T, Mima T, Shiota S, Kuroda T, Mizushima T, Gotoh N, Nishino T, Tsuchiya T (2003) Induction of mexCD-oprJ operon for a multidrug efflux pump by disinfectants in wild-type Pseudomonas aeruginosa PAO1. J Antimicrob Chemother 51(4):991–994. https://doi.org/10.1093/jac/dkg173
Poole K, Gotoh N, Tsujimoto H, Zhao Q, Wada A, Yamasaki T, Neshat S, Ji Y, Li XZ, Nishino T (1996) Overexpression of the mexC-mexD-oprJ efflux operon in nfxB-type multidrug-resistant strains of Pseudomonas aeruginosa. Mol Microbiol 21(4):713–725. https://doi.org/10.1046/j.1365-2958.1996.281397.x
Pruden A, Pei R, Storteboom H, Carlson KH (2006) Antibiotic resistance genes as emerging contaminants: studies in northern Colorado. Environ Sci Technol 40(23):7445–7450. https://doi.org/10.1021/es060413l
Purssell A, Fruci M, Mikalauskas A, Gilmour C, Poole K (2015) EsrC, an envelope stress-regulated repressor of the mexCD-oprJ multidrug efflux operon in Pseudomonas aeruginosa. Environ Microbiol 17(1):186–198. https://doi.org/10.1111/1462-2920.12602
Putman M, van Veen HW, Konings WN (2000) Molecular properties of bacterial multidrug transporters. Microbiol Mol Biol Rev 64(4):672–693. https://doi.org/10.1128/mmbr.64.4.672-693.2000
Rhodes G, Parkhill J, Bird C, Ambrose K, Jones MC, Huys G, Swings J, Pickup RW (2004) Complete nucleotide sequence of the conjugative tetracycline resistance plasmid pFBAOT6, a member of a group of IncU plasmids with global ubiquity. Appl Environ Microbiol 70(12):7497–7510. https://doi.org/10.1128/aem.70.12.7497-7510.2004
Robicsek A, Jacoby GA, Hooper DC (2006) The worldwide emergence of plasmid-mediated quinolone resistance. Lancet Infect Dis 6(10):629–640. https://doi.org/10.1016/S1473-3099(06)70599-0
Schmidt AS, Bruun MS, Dalsgaard I, Larsen JL (2001) Incidence, distribution, and spread of tetracycline resistance determinants and Integron-associated antibiotic resistance genes among motile Aeromonads from a fish farming environment. Appl Environ Microbiol 67(12):5675–5682. https://doi.org/10.1128/aem.67.12.5675-5682.2001
Sentchilo V, Mayer AP, Guy L, Miyazaki R, Green Tringe S, Barry K, Malfatti S, Goessmann A, Robinson-Rechavi M, van der Meer JR (2013) Community-wide plasmid gene mobilization and selection. ISME J 7(6):1173–1186. https://doi.org/10.1038/ismej.2013.13
Sun F, Zhou D, Wang Q, Feng J, Feng W, Luo W, Liu Y, Qiu X, Yin Z, Xia P (2016) Genetic characterization of a novel blaDIM-2-carrying megaplasmid p12969-DIM from clinical Pseudomonas putida. J Antimicrob Chemother 71(4):909–912. https://doi.org/10.1093/jac/dkv426
Szczepanowski R, Bekel T, Goesmann A, Krause L, Krömeke H, Kaiser O, Eichler W, Pühler A, Schlüter A (2008) Insight into the plasmid metagenome of wastewater treatment plant bacteria showing reduced susceptibility to antimicrobial drugs analysed by the 454-pyrosequencing technology. J Biotechnol 136(1):54–64. https://doi.org/10.1016/j.jbiotec.2008.03.020
Szczepanowski R, Linke B, Krahn I, Gartemann K-H, Gützkow T, Eichler W, Pühler A, Schlüter A (2009) Detection of 140 clinically relevant antibiotic-resistance genes in the plasmid metagenome of wastewater treatment plant bacteria showing reduced susceptibility to selected antibiotics. Microbiology 155(7):2306–2319. https://doi.org/10.1099/mic.0.028233-0
Tamaki H, Sekiguchi Y, Hanada S, Nakamura K, Nomura N, Matsumura M, Kamagata Y (2005) Comparative analysis of bacterial diversity in freshwater sediment of a shallow eutrophic lake by molecular and improved cultivation-based techniques. Appl Environ Microbiol 71(4):2162–2169. https://doi.org/10.1128/AEM.71.4.2162-2169.2005
Tang X, Lou C, Wang S, Lu Y, Liu M, Hashmi MZ, Liang X, Li Z, Liao Y, Qin W, Fan F, Xu J, Brookes PC (2015) Effects of long-term manure applications on the occurrence of antibiotics and antibiotic resistance genes (ARGs) in paddy soils: evidence from four field experiments in south of China. Soil Biol Biochem 90:179–187. https://doi.org/10.1016/j.soilbio.2015.07.027
Thomas CM, Nielsen KM (2005) Mechanisms of, and barriers to, horizontal gene transfer between bacteria. Nat Rev Microbiol 3(9):711–721. https://doi.org/10.1038/nrmicro1234
Tian Z, Zhang Y, Yu B, Yang M (2016) Changes of resistome, mobilome and potential hosts of antibiotic resistance genes during the transformation of anaerobic digestion from mesophilic to thermophilic. Water Res 98:261–269. https://doi.org/10.1016/j.watres.2016.04.031
Van't Veer LJ, Dai H, Van De Vijver MJ, He YD, Hart AA, Mao M, Peterse HL, Van Der Kooy K, Marton MJ, Witteveen AT (2002) Gene expression profiling predicts clinical outcome of breast cancer. Nature 415(6871):530–536
Wagner J, Coupland P, Browne HP, Lawley TD, Francis SC, Parkhill J (2016) Evaluation of PacBio sequencing for full-length bacterial 16S rRNA gene classification. BMC Microbiol 16(1):274. https://doi.org/10.1186/s12866-016-0891-4
Wu S, Wang G, Angert ER, Wang W, Li W, Zou H (2012) Composition, diversity, and origin of the bacterial community in grass carp intestine. PLoS One 7(2):e30440. https://doi.org/10.1371/journal.pone.0030440
Xiong W, Sun Y, Ding X, Wang M, Zeng Z (2015) Selective pressure of antibiotics on ARGs and bacterial communities in manure-polluted freshwater-sediment microcosms. Front Microbiol 6:194. https://doi.org/10.3389/fmicb.2015.00194
Yang J, Wang C, Shu C, Liu L, Geng J, Hu S, Feng J (2013a) Marine sediment bacteria harbor antibiotic resistance genes highly similar to those found in human pathogens. Microb Ecol 65(4):975–981. https://doi.org/10.1007/s00248-013-0187-2
Yang Y, Li B, Ju F, Zhang T (2013b) Exploring variation of antibiotic resistance genes in activated sludge over a four-year period through a metagenomic approach. Environ Sci Technol 47(18):10197–10205. https://doi.org/10.1021/es4017365
Zhang Y, Tian Z, Liu M, Shi ZJ, Hale L, Zhou J, Yang M (2015) High concentrations of the antibiotic spiramycin in wastewater lead to high abundance of ammonia-oxidizing archaea in nitrifying populations. Environ Sci Technol 49(15):9124–9132. https://doi.org/10.1021/acs.est.5b01293
Acknowledgments
We thank Dr. Ye Deng and Dr. Jie Feng for their help in statistical analysis.
Funding
This work was supported by the National Natural Scientific Foundation of China (21437005).
Author information
Authors and Affiliations
Contributions
Yu Zhang, Yanhong Shi, and Min Yang designed research; Yanhong Shi, Hong Zhang performed research; Yanhong Shi, Zhe Tian, Yu Zhang, and Min Yang analyzed data and wrote the paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
ESM 1
(PDF 1819 kb)
Rights and permissions
About this article
Cite this article
Shi, Y., Zhang, H., Tian, Z. et al. Characteristics of ARG-carrying plasmidome in the cultivable microbial community from wastewater treatment system under high oxytetracycline concentration. Appl Microbiol Biotechnol 102, 1847–1858 (2018). https://doi.org/10.1007/s00253-018-8738-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-8738-6