Abstract
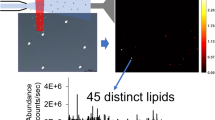
MALDI-TOF MS is traditionally used for “proteomics”, but is also a useful tool for lipid analysis. Depending on the applied matrix, however, some lipid classes are more sensitively detected than other ones and this may even lead to suppression effects if complex mixtures are analyzed. Therefore, a previous separation into the individual lipid classes is necessary. Using artificial lipid mixtures or easily available tissue extracts, it has been already shown that HPTLC-(High Performance Thin-Layer Chromatography)-separated lipids can be conveniently analyzed by MALDI-TOF MS directly on the TLC plate. Here we present an initial TLC-MALDI study of the lipid composition of ovine mesenchymal stem cells. Due to the complex composition of these cells, data are also compared to lipids extracted from human erythrocytes. It will be shown that even very minor lipid classes can be easily detected and with much higher sensitivity than by common staining protocols. Additionally, MS images of the developed TLC plates will be shown and potential applications, new methods of data analysis as well as problems discussed.









Similar content being viewed by others
References
Salgado AJ, Oliveira JT, Pedro AJ, Reis RL (2006) Curr Stem Cell Res Ther 1:345–364
Schiller J, Fuchs B, Arnold K (2006) Curr Org Chem 10:1771–1789
Fuchs B, Schiller J, Cross MA (2007) Chem Phys Lipids 150:229–238
Peterson BL, Cummings BS (2006) Biomed Chromatogr 20:227–243
Watson AD (2006) J Lipid Res 47:2101–2111
Wenk MR (2005) Nat Rev Drug Discov 4:594–610
Pulfer M, Murphy RC (2003) Mass Spectrom Rev 22:332–364
Lee SH, Williams MV, DuBois RN, Blair IA (2003) Rapid Commun Mass Spectrom 17:2168–2176
Schiller J, Süß R, Arnhold J, Fuchs B, Leßig J, Müller M, Petković M, Spalteholz H, Zschörnig O, Arnold K (2004) Prog Lipid Res 43:449–488
Schiller J, Süß R, Fuchs B, Müller M, Zschörnig O, Arnold K (2007) Front Biosci 12:2568–2579
Hillenkamp F, Peter-Katalinić J (2007) MALDI MS - A Practical Guide to Instrumentation, Methods and Application. Wiley-VCH, Weinheim
Gellermann GP, Appel TR, Davies P, Diekmann S (2006) Biol Chem 387:1267–1274
Petković M, Schiller J, Müller M, Benard S, Reichl S, Arnold K, Arnhold J (2001) Anal Biochem 289:202–216
Stubiger G, Belgacem O (2007) Anal Chem 79:3206–3213
Estrada R, Yappert MC (2004) J Mass Spectrom 39:412–422
Schiller J, Süß R, Fuchs B, Müller M, Petković M, Zschörnig O, Waschipky H (2007) Eur Biophys J 36:517–527
Touchstone JC (1995) J Chromatogr B Biomed Appl 671:169–195
Schiller J, Süß R, Fuchs B, Müller M, Zschörnig O, Arnold K (2003) Chromatographia 57:297–302
Fuchs B, Süß R, Nimptsch A, Schiller J (2008) Chromatographia, in press
Nakamura K, Suzuki Y, Goto-Inoue N, Yoshida-Noro C, Suzuki A (2006) Anal Chem 78:5736–5743
Dreisewerd K, Muthing J, Rohlfing A, Meisen I, Vukelic Z, Peter-Katalinić J, Hillenkamp F, Berkenkamp S (2005) Anal Chem 77:4098–4107
Rohlfing A, Müthing J, Pohlentz G, Distler U, Peter-Katalinić J, Berkenkamp S, Dreisewerd K (2007) Anal Chem 79:5793–808
Fuchs B, Schiller J, Süß R, Schürenberg M, Suckau D (2007) Anal Bioanal Chem 389:827–834
Nouri-Sorkhabi MH, Wright LC, Sullivan DR, Kuchel PW (1996) Lipids 31:765–770
Tannert A, Kurz A, Erlemann KR, Müller K, Herrmann A, Schiller J, Töpfer-Petersen E, Manjunath P, Müller P (2007) Eur Biophys J 36:461–475
Schulz R, Zscharnack M, Hanisch I, Geiling M, Hepp P, Bader A (2008) Biomed Mater Eng 18:55–70
Folch J, Lees M, Stanley GHS (1957) J Biol Chem 226:497–509
Schiller J, Arnold K (2002) Med Sci Monit 8:MT205–MT222
Schiller J, Müller M, Fuchs B, Arnold K, Huster D (2007) Curr Anal Chem 3:283–301
White T, Bursten S, Frederighi D, Lewis RA, Nudelman E (1998) Anal Biochem 10:109–117
Suckau D, Resemann A, Schürenberg M, Hufnagel P, Franzen J, Holle A (2003) Anal Bioanal Chem 376:952–965
Fuchs B, Schober C, Richter G, Süß R, Schiller J (2007) J Biochem Biophys Meth 70:689–692
Puppato A, DuPré DB, Stolowich N, Yappert MC (2007) Chem Phys Lipids 150:176–185
Petković M, Schiller J, Müller J, Müller M, Arnold K, Arnhold J (2001) Analyst 126:1042–1050
Al-Saad KA, Siems WF, Hill HH, Zabrouskov V, Knowles NR (2003) J Am Soc Mass Spectrom 14:373–382
Mehl JT, Gusev AI, Hercules DM (1997) Chromatographia 46:358–364
Schiller J, Süß R, Petković M, Zschörnig O, Arnold K (2002) Anal Biochem 309:311–314
Sugiyama E, Hara A, Uemura K, Taketomi T (1997) Glycobiology 7:719–724
He H, Conrad CA, Nilsson CL, Ji Y, Schaub TM, Marshall AG, Emmett MR (2007) Anal Chem 79:8423–8430
Schiller J, Müller K, Süß R, Arnhold J, Gey C, Herrmann A, Leßig J, Arnold K, Müller P (2003) Chem Phys Lipids 126:85–94
Hidaka H, Hanyu N, Sugano M, Kawasaki K, Yamauchi K, Katsuyama T (2007) Ann Clin Lab Sci 37:213–221
Stoeckli M, Chaurand P, Hallahan DE, Caprioli RM (2001) Nature Med 7:493–496
Acknowledgements
This work was supported by the German Research Council (DFG Schi 476/5–1) and the Federal Ministry of Education and Research (Grant BMBF 0313836).
Author information
Authors and Affiliations
Corresponding author
Additional information
Beate Fuchs, Jürgen Schiller and Rosmarie Süß contributed equally to this work.
Rights and permissions
About this article
Cite this article
Fuchs, B., Schiller, J., Süß, R. et al. Analysis of stem cell lipids by offline HPTLC-MALDI-TOF MS. Anal Bioanal Chem 392, 849–860 (2008). https://doi.org/10.1007/s00216-008-2301-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-008-2301-8