Abstract
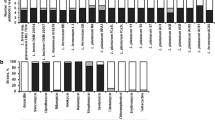
Wild rats are known to carry different microorganisms and are considered a reservoir of zoonotic pathogens worldwide. The urban rats were collected from five districts of Tehran and Gram-negative bacteria (GNB) were isolated from fecal samples and were identified using classical biochemical tests. The antibiotic susceptibility patterns of isolated bacteria were determined by Kirby–Bauer disk diffusion method, the results of which were interpreted in line with CLSI guideline. The frequency of antibiotic-resistant genes was identified using multiplex-PCR. Moreover, PCR method was used to identify the frequency of Escherichia coli O157:H7 and main categories of diarrheagenic E. coli including EPEC, ETEC, EIEC, EAEC, and STEC pathotypes. A total of 100 Rattus norvegicus were trapped and fecal samples were collected. Overall, 72 fecal samples were positive for GNB. E. coli (n = 46/72) had the highest frequency among the isolated GNB. Among E. coli isolates, the highest and lowest resistance rates belonged to ampicillin (56.5%) and ceftriaxone (0%), respectively. Klebsiella spp. was 100% resistant to imipenem, and streptomycin (0%) was the most effective antimicrobial agent on Klebsiella spp. Among surveyed genes, blaTEM (95.8%) and blaaadA-1 (58.3%) had the highest frequency, while blaKPC, and blaCMY-2 were not detected among Enterobacteriaceae. Herein, O157: H7 serotype was not detected and aEPEC (87%) was the most common pathotype detected. Results suggested that rodents might be a reservoir of antimicrobial-resistant pathogens and rodent control along with implementation of surveillance programs should be considered as a critical priority for urban health.

Similar content being viewed by others
Data availability statement
All data generated or analyzed during this study are included in this published article.
References
Aktan I, Sprigings K, La Ragione R, Faulkner L, Paiba G, Woodward MJJ (2004) Characterisation of attaching–effacing Escherichia coli isolated from animals at slaughter in England and Wales. Vet Microbiol 102:43–53
Ayral F et al (2015) The relationship between socioeconomic indices and potentially zoonotic pathogens carried by wild Norway rats: a survey in Rhône, France (2010–2012). Epidemiol Infect 143:586–599
Azimi T, Maham S, Fallah F, Azimi L, Gholinejad ZJ (2019) Evaluating the antimicrobial resistance patterns among major bacterial pathogens isolated from clinical specimens taken from patients in Mofid Children’s Hospital, Tehran, Iran: 2013–2018. Infect Drug Resist 12:2089
Bedenić B, Plečko V, Sardelić S, Uzunović S, Godič Torkar KJ (2014) Carbapenemases in gram-negative bacteria: laboratory detection and clinical significance. Biomed Res Int 2014:841951
Bok E, Mazurek J, Stosik M, Wojciech M, Baldy-Chudzik K (2015) Prevalence of virulence determinants and antimicrobial resistance among commensal Escherichia coli derived from dairy and beef cattle. Int J Environ Res Public Health 12:970–985
Burriel AR, Kritas SK, Kontos V (2008) Some microbiological aspects of rats captured alive at the port city of Piraeus, Greece. Int J Environ Health Res 18:159–164
Corzo-Ariyama HA, García-Heredia A, Heredia N, García S, León J, Jaykus L, Solís-Soto L (2019) Phylogroups, pathotypes, biofilm formation and antimicrobial resistance of Escherichia coli isolates in farms and packing facilities of tomato, jalapeño pepper and cantaloupe from Northern Mexico. Int J Food Microbiol 290:96–104
Deborah Chen H, Frankel G (2005) Enteropathogenic Escherichia coli: unravelling pathogenesis. FEMS Microbiol Rev 29:83–98
Desvars-Larrive A et al (2019) Urban brown rats (Rattus norvegicus) as possible source of multidrug-resistant Enterobacteriaceae and meticillin-resistant Staphylococcus spp., Vienna, Austria, 2016 and 2017. Euro Surveill. 24:1900149
Dhanji H, Murphy NM, Akhigbe C, Doumith M, Hope R, Livermore DM, Woodford NJ (2011) Isolation of fluoroquinolone-resistant O25b: H4-ST131 Escherichia coli with CTX-M-14 extended-spectrum β-lactamase from UK river water. J Antimicrob Chemother 66:512–516
Disease Control and Prevention (2013) Antibiotic resistance threats in the United States, 2013. Centres for Disease Control and Prevention.
ECDC (2017) Antimicrobial resistance surveillance in Europe 2012. Annual Report of the European antimicrobial resistance surveillance network (EARS-Net). ECDC, Stockholm
Firth C et al (2014) Detection of zoonotic pathogens and characterization of novel viruses carried by commensal Rattus norvegicus in New York City. mBio. 5:e1933-e01914
Gakuya F, Kyule M, Gathura P, Kariuki S (2001) Antimicrobial susceptibility and plasmids from Escherichia coli isolated from rats. East Afr Med J 78:518–522
Galán-Puchades MT et al (2018) First survey on zoonotic helminthosis in urban brown rats (Rattus norvegicus) in Spain and associated public health considerations. Vet Parasitol 259:49–52
Guenther S, Bethe A, Fruth A, Semmler T, Ulrich RG, Wieler LH, Ewers C (2012) Frequent combination of antimicrobial multiresistance and extraintestinal pathogenicity in Escherichia coliisolates from urban rats (Rattus norvegicus) in Berlin, Germany. PLoS ONE 7:e50331
Guenther S, Ewers C, Wieler LH (2011) Extended-spectrum beta-lactamases producing E.coli in wildlife, yet another form of environmental pollution. Front Microbiol 2:246
Himsworth CG et al (2015) Prevalence and characteristics of Escherichia coli and Salmonella spp. in the feces of wild urban Norway and black rats (Rattus norvegicus and Rattus rattus) from an inner-city neighborhood of Vancouver, Canada. J Wildl Dis 51:589–600
Ho P-L, Lo W-U, Lai EL, Law PY, Leung SM, Wang Y, Chow K-HJ (2015) Clonal diversity of CTX-M-producing, multidrug-resistant Escherichia coli from rodents. J Med Microbiol 64:185–190
Holland J, Louie L, Simor A, Louie MJ (2000) PCR detection of Escherichia coli O157: H7 directly from stools: evaluation of commercial extraction methods for purifying fecal DNA. J Clin Microbiol 38:4108–4113
Maas M et al (2018) Prevalence of Leptospira spp. and Seoul hantavirus in brown rats (Rattusnorvegicus) in four regions in the Netherlands, 2011–2015. Infect Ecol Epidemiol 8:1490135
Meerburg BG, Singleton GR, Kijlstra AJ (2009) Rodent-borne diseases and their risks for public health. Crit Rev Microbiol 35:221–270
Mohammadzadeh M, Goudarzi H, Dabiri H, Fallah F (2017) Characterization of enteropathogenic Escherichia coli associated with diarrhea among Iranian infants. J Pediatr Infect Dis 5:e36920
Moshtagian F, Alipour M, Yahyapour YJ (2016) Prevalence of Escherichia coli pathotypes among children with diarrhea in Babol, Northern Iran. Int J Enter Pathog 4:1–4
Nataro JP, Kaper JB (1998) Diarrheagenic Escherichia coli. Clin Microbiol Rev 11:142–201
Nielsen EM, Skov MN, Madsen JJ, Lodal J, Jespersen JB, Baggesen DL (2004) Verocytotoxin-producing Escherichia coli in wild birds and rodents in close proximity to farms. Appl Environ Microbiol 70:6944–6947
Nkogwe C, Raletobana J, Stewart-Johnson A, Suepaul S, Adesiyun A (2011) Frequency of detection of Escherichiacoli, Salmonella spp., and Campylobacter spp. in the faeces of wild rats (Rattus spp.) in Trinidad and Tobago. Vet Med Int 2011:686923
Pormohammad A, Nasiri MJ, Azimi T (2019) Prevalence of antibiotic resistance in Escherichiacoli strains simultaneously isolated from humans, animals, food, and the environment: a systematic review and meta-analysis. Infect Drug Resist 12:118
Rothenburger JL, Himsworth CH, Nemeth NM, Pearl DL, Jardine CM (2017) Environmental factors and zoonotic pathogen ecology in urban exploiter species. EcoHealth 14:630–641
Runge M et al (2013) Distribution of rodenticide resistance and zoonotic pathogens in Norway rats in Lower Saxony and Hamburg, Germany. Pest Manag Sci 69:403–408
Schaufler K et al (2018) Clinically relevant ESBL-producing K.pneumoniae ST307 and E.coli ST38 in an Urban West African rat population. Front Microbiol 9:150
Swedan S, Abu Alrub H (2019) Antimicrobial resistance, virulence factors, and pathotypes of Escherichia coli isolated from drinking water sources in Jordan. Pathogens 8:86
Taghadosi R, Shakibaie MR, Hosseini-Nave H (2019) Antibiotic resistance, ESBL genes, integrons, phylogenetic groups and MLVA profiles of Escherichia coli pathotypes isolated from patients with diarrhea and farm animals in south-east of Iran. Comp Immunol Microbiol Infect Dis 63:117–126
World Health Organization (2009) Diarrhoea: why children are still dying and what can be done. World Health Organization, Geneva
Acknowledgements
This article was extracted from the Ph.D. thesis written by Mr. Taher Azimi in Department of Pathobiology, School of Public Health, Tehran University of Medical Sciences, Tehran, Iran (Registration No. 240/316). This study was supported by research committee of Tehran University of Medical Sciences, Tehran, Iran (Grant No. 42072).
Funding
This research did not receive any specific grant from funding agencies in public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
TA, MRP, and MRA: conceptualization; data curation; formal analysis; and writing—original draft. TA, FF, LA, and MRP: conceptualization; methodology; project administration; and writing—original draft. AO, MRA, and ARF: data curation; formal analysis; writing—original draft; and writing—review and editing. TA and LA: language editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethics approval and consent to participate
The present study was approved by the Ethics Committee of School of Public Health and Allied Medical Sciences, Tehran University of Medical Sciences, with reference number IR.TUMS.SPH.REC.1398.035. All authors of this research paper have directly participated in the planning, execution, or analysis of this study.
Consent for publication
All authors made substantial contributions to the conception and design, acquisition of data, or analysis and interpretation of data. They played an active role in drafting the article or revising it critically to achieve important intellectual content, gave the final approval of the version to be published, and agreed to be accountable for all aspects of the work.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Azimi, T., Azimi, L., Fallah, F. et al. Detection and characterization of Enterobacteriaceae family members carried by commensal Rattus norvegicus from Tehran, Iran. Arch Microbiol 203, 1321–1334 (2021). https://doi.org/10.1007/s00203-020-02126-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-020-02126-0