Abstract
Arabidopsis thaliana has, in conjunction with A. arenosa, developed into a system for the molecular analysis of alloplolyploidy. However, there are very few Arabidopsis lines available to study autopolyploidy. In order to investigate polyploidy on a reliable basis, we have optimised conventional methodologies and developed a novel strategy for the rapid generation and identification of polyploids based on trichome branching patterns. The analysis of more than two dozen independently induced Arabidopsis lines has led to interesting observations concerning the relationship between cell size and ploidy levels and on the relative stability of tetraploidy in Arabidopsis over at least three consecutive generations. The most important finding of this work is that neo-tetraploid lines exhibit considerable stability through all the generations tested. The systematic generation of tetraploid collections through this strategy as well as the lines generated in this work will help to unravel the consequences of polyploidy, particularly tetraploidy, on the genome, on gene expression and on natural diversity in Arabidopsis.







Similar content being viewed by others
References
Adams KL, Wendel JF (2005a) Novel patterns of gene expression in polyploidy plants. Tends Genet 21:539–543
Adams KL, Wendel JF (2005b) Polyploidy and genome evolution in plants. Curr Opin Plant Biol 8:135–141
Albertin W, Balliau T, Brabant P, Chevre AM, Eber F, Malosse C, Thiellement H (2006) Numerous and rapid nonstochastic modifications of gene products in newly synthesized Brassica napus allotetraploids. Genetics 73:1101–1113
Altmann T, Damm B, Frommer WB, Martin T, Morris PC, Schweizer D, Willmitzer L, Schmidt R (1994) Easy determination of ploidy level in Arabidopsis thaliana plants by means of pollen size measurement. Plant Cell Rep 13:652–656
Bennett MD (2004) Perspectives on polyploidy in plants—ancient and neo. Biol J Lin Soc 82:411–423
Bouharmont J (1965) Fertility studies in polyploid Arabidopsis thaliana. In: Röbbelen G (ed) Arabidopsis research. Wasmund, Gelsenkirchen, pp 31–36
Comai L (2005) The advantages and disadvantages of being polyploid. Nat Rev Genet 6:836–846
Dawe RK, Freeling M (1991) Cell lineage and its consequences in higher plants. Plant J 1:3–8
Duckett CM, Oparka KJ, Prior DAM, Dolan L, Roberts K (1994) Dye-coupling in the root epidermis of Arabidopsis is progressively reduced during development. Development 120:3247–3255
Friml J, Vieten A, Sauer M, Weijers D, Schwarz H, Hamann T, Offringa R, Jurgens G (2003) Efflux-dependent auxin gradients establish the apical-basal axis of Arabidopsis. Nature 426:147–153
Gaeta RT, Pires JC, Iniguez-Luy F, Leon E, Osborn TC (2007) Genomic changes in resynthesized Brassica napus and their effect on gene expression and phenotype. Plant Cell 19:3403–3417
Galbraith DW, Harkins KR, Knapp S (1991) Systemic endopolyploidy in Arabidopsis thaliana. Plant Physiol 96:985–989
Hauser MT, Bauer E (2000) Histochemical analysis of root meristem activity in Arabidopsis thaliana using a cyclin: GUS (ß-glucuronidase) marker line. Plant Soil 226:1–10
Henry IM, Dilkes BP, Young K, Watson B, Wu H, Comai L (2005) Aneuploidy and genetic variation in the Arabidopsis thaliana triploid response. Genetics 170:1979–1988
Heslop-Harrison JS, Maluszynska J (1994) Molecular cytogenetics of Arabidopsis. In: Meyerowitz EM, Somerville CR (eds) Arabidopsis. Cold Spring Harbor Laboratory Press, New York, pp 63–88
Jürgens G, Mayer U, Torres Ruiz RA, Berleth T, Misera S (1991) Genetic analysis of pattern formation in Arabidopsis embryo. Dev Suppl 1:27–38
Kashkush K, Feldman M, Levy AA (2002) Gene loss, silencing and activation in a newly synthesized wheat allotetraploid. Genetics 160:1651–1659
Koornneef M (1994) Arabidopsis genetics. In: Meyerowitz EM, Somerville CR (eds) Arabidopsis. Cold Spring Harbor Laboratory Press, New York, pp 89–120
Larkins BA, Dilkes BP, Dante RA, Coelho CM, Woo YM, Liu Y (2001) Investigating the hows and whys of DNA endoreduplication. J Exp Bot 52:183–192
Lister C, Dean C (1993) Recombinant inbred lines for mapping RFLP and phenotypic markers in Arabidopsis thaliana. Plant J 4:745–750
Liu B, Brubaker CL, Mergeai G, Cronn RC, Wendel JF (2001) Polyploid formation in cotton is not accompanied by rapid genomic changes. Genome 44:321–330
Lukens LN, Pires JC, Leon E, Vogelzang R, Oslach L, Osborn T (2006) Patterns of sequence loss and cytosine methylation within a population of newly resynthesized Brassica napus allopolyploids. Plant Physiol 140:336–348
Madlung A, Masuelli RW, Watson B, Reynolds SH, Davison J, Comai L (2002) Remodeling of DNA methylation and phenotypic and transcriptional changes in synthetic Arabidopsis allotetraploids. Plant Physiol 129:733–746
Maluszynska J, Heslop-Harrison JS (1991) Localization of tandemly repeated DNA sequences in Arabidopsis thaliana. Plant J 1:159–166
Mayer U, Torres Ruiz RA, Berleth T, Misera S, Jürgens G (1991) Mutations affecting body organization in the Arabidopsis embryo. Nature 353:402–407
Melaragno JE, Mehrotra B, Coleman AW (1993) Relationship between endopolyploidy and cell size in epidermal tissue of Arabidopsis. Plant Cell 5:1661–1668
Mittelsten-Scheid O, Afsar K, Paszkowski J (2003) Formation of stable epialleles and their paramutation-like interaction in tetraploid Arabidopsis thaliana. Nat Genet 34:450–454
Osborn TC, Pires JC, Birchler JA, Auger DL, Chen ZJ, Lee HS, Comai L, Madlung A, Doerge RW, Colot V, Martienssen RA (2003) Understanding mechanisms of novel gene expression in polyploids. Trends Genet 19:141–147
Otto SP, Whitton J (2000) Polyploid incidence and evolution. Annu Rev Genet 34:401–437
Perazza D, Herzog M, Hülskamp M, Brown S, Dorne AM, Bonneville JM (1999) Trichome cell growth in Arabidopsis thaliana can be derepressed by mutations in at least five genes. Genetics 152:461–476
Redei GP (1964) Crossing experiences with polyploids. Arabidopsis Inf Serv 1:13
Santos JL, Alfaro D, Sanchez-Moran E, Armstrong SJ, Franklin FC, Jones GH (2003) Partial diploidization of meiosis in autotetraploid Arabidopsis thaliana. Genetics 165:1533–1540
Schmuths H, Meister A, Horres R, Bachmann K (2004) Genome size variation among accessions of Arabidopsis thaliana. Ann Bot 93:317–321
Soltis PS, Soltis DE (2000) The role of genetic and genomic attributes in the success of polyploids. Proc Natl Acad Sci USA 97:7051–7057
Song K, Lu P, Tang K, Osborn TC (1995) Rapid genome change in synthetic polyploids of Brassica and its implications for polyploid evolution. Proc Natl Acad Sci USA 92:7719–7723
Storchova Z, Breneman A, Cande J, Dunn J, Burbank K, O’Toole E, Pellman D (2006) Genome-wide genetic analysis of polyploidy in yeast. Nature 443:541–547
Takada S, Jürgens G (2007) Transcriptional regulation of epidermal cell fate in the Arabidopsis embryo. Development 134:1141–1150
Tilney-Bassett RA (1986) Plant chimeras. Edward Arnold Ltd, Baltimore
Wang J, Tian L, Madlung A, Lee HS, Chen M, Lee JJ, Watson B, Kagochi T, Comai L, Chen ZJ (2004) Stochastic and epigenetic changes of gene expression in Arabidopsis polyploids. Genetics 167:1961–1973
Wang J, Tian L, Lee HS, Wie NE, Jiang H, Watson B, Madlung A, Osborn TC, Doerge RW, Comai L, Chen ZJ (2006) Genomewide nonadditive gene regulation in Arabidopsis tetraploids. Genetics 172:507–517
Weiss H, Maluszynska J (2000) Chromosomal rearrangement in autotetraploid plants of Arabidopsis thaliana. Hereditas 133:255–261
Wolters AM, Visser RG (2000) Gene silencing in potato: allelic differences and effect of ploidy. Plant Mol Biol 43:377–386
Zhong XB, de Jong JH, Zabel P (1996) Preparation of tomato meioticpachytene and mitotic metaphase chromosomes suitable for fluorescence in situ hybridization (FISH). Chromosom Res 4:24–28
Acknowledgements
We thank Dr. Daniel and his group (Bayerische Landesanstalt für Landwirtschaft) for generous provision of the Partec II Flow Cytometer and help, Dr. Stephan Haug (Lehrstuhl für Mathematische Statistik, TU München) for helpful advice in statistics and an anonymous reviewer for useful hints. We owe very special thanks to Farhah Assaad for critically reading the manuscript and useful discussions. We thank the Nottingham Arabidopsis Stock Centre (NASC) for lines. We are particularly indebted to the following colleagues for kindly providing lines: Ortrun Mittelsten-Scheid (Wien), Jolanta Maluszynska (Katowice), Marie-Theres Hauser (Wien), Thomas Debener (Hannover) and Jiri Friml (Ghent). The financial support by the Deutsche Forschungsgemeinschaft (Grant GI 140/12-1 to A.G. and R.A.T.R.) is gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Kearsey.
Electronic supplementary material
Below is the link to the electronic supplementary material.
122_2009_966_MOESM1_ESM.tif
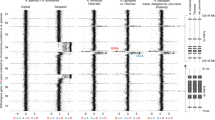
SFig. 1 Additional size features in polyploids. DAPI stained nuclei in stomata (a-c) and trichomes (d+e): a) diploid Col-0, b) tetraploid Col-0, c) octoploid Col-0, d) diploid Ler-0 and e) aneuploid Col-0 (black arrows). Comparison of organs of diploid and tetraploid transgenic lines, respectively (f-i). Root tips in f+g (CYCAt1::GUS transgene); whole embryo and root tips in h-i (DR5rev::GFP transgene). Note that the expression domains of the reporter constructs remain stable in the tetraploids. The GUS reporter indicates the actively dividing region in the root tip. The DR5rev::GFP reporter indicates the auxin maximum of the root tip (white arrow; inset in i) shows magnification of root tip). Sizes of seeds from different polyploids as indicated (j-m). Scale bars: in c) 10μm is the same in a+b); in e) 100μm is the same in d); in g) 80μm is the same in f); in h+i) is 80μm; in inset of i) 30μm; in m) 50μm is the same in j-l) (TIFF 7006 kb)
Rights and permissions
About this article
Cite this article
Yu, Z., Haage, K., Streit, V.E. et al. A large number of tetraploid Arabidopsis thaliana lines, generated by a rapid strategy, reveal high stability of neo-tetraploids during consecutive generations. Theor Appl Genet 118, 1107–1119 (2009). https://doi.org/10.1007/s00122-009-0966-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-009-0966-9