Abstract
Background and objective
Acute lung injury (ALI)/ acute respiratory distress syndrome (ARDS) was increasingly recognized as one of the most severe acute hyperimmune response of coronavirus disease 2019 (COVID-19). Clofazimine (CFZ) has attracted attention due to its anti-inflammatory property in immune diseases as well as infectious diseases. However, the role and potential molecular mechanism of CFZ in anti-inflammatory responses remain unclear.
Methods
We analyze the protein expression profiles of CFZ and LPS from Raw264.7 macrophages using quantitative proteomics. Next, the protective effect of CFZ on LPS-induced inflammatory model is assessed, and its underlying mechanism is validated by molecular biology analysis.
Results
LC–MS/MS-based shotgun proteomics analysis identified 4746 (LPS) and 4766 (CFZ) proteins with quantitative information. The key proteins and their critical signal transduction pathways including TLR4/NF-κB/HIF-1α signaling was highlighted, which was involved in multiple inflammatory processes. A further analysis of molecular biology revealed that CFZ could significantly inhibit the proliferation of Raw264.7 macrophages, decrease the levels of TNF-α and IL-1β, alleviate lung histological changes and pulmonary edema, improve the survival rate, and down-regulate TLR4/NF-κB/HIF-1α signaling in LPS model.
Conclusion
This study can provide significant insight into the proteomics-guided pharmacological mechanism study of CFZ and suggest potential therapeutic strategies for infectious disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), a highly infectious RNA virus, is responsible for coronavirus disease 2019 (COVID-19) [1], which has caused more than 5.8 million deaths (February 2022, Johns Hopkins University Coronavirus Resource Center) in modern history so far. The target of SARS-CoV-2 primarily infects the respiratory tract epithelial cells and activates the immune system, in turn, leading to cytokine storm (CS) and causing fever, cough, breathlessness, etc. [2]. In some cases, 10–20% of patients with COVID-19 might also develop acute lung injury (ALI) or progress to acute respiratory distress syndrome (ARDS) and fatal multi-organ dysfunction (MODS) and even death [3, 4].
Current management of severe COVID-19 focus on symptomatic and supportive treatment [3, 4]. Aside from combination therapy, there is no widely available antiviral therapies for severe pneumonia. Thus, it is in critical need to develop additional therapeutic strategies for COVID-19. Recently, clofazimine (CFZ) has raised some interest, which simultaneously inhibits viral activity and alleviates the symptoms in COVID-19 patients [5]. Previously, it has been used in treating leprosy, and more recently in multidrug-resistant tuberculosis [6, 7]. Accumulated evidence points out that CFZ displays an important immunomodulatory activity and anti-inflammatory property in immune disease, including discoid lupus erythematosus, chronic lymphocytic leukemia, Crohn disease, ulcerative colitis, etc. [5, 8,9,10,11]. In addition, it is anti-bacteria by generating reactive oxygen species (ROS), particularly superoxide and hydrogen peroxide (H2O2) [12, 13]. Up to now, despite the aforementioned interesting findings in the literature, there is still much to be understood about the role of CFZ in ALI/ARDS.
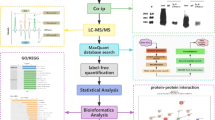
Lipopolysaccharide (LPS)-induced inflammatory responses are an ideal model closely resembling ALI/ARDS [14,15,16]. Raw264.7 macrophage, a critical immunological cell, plays a hub role in the pathogenesis of ALI/ARDS [17]. Here, the first proteomic study is presented to assess the effects of CFZ on Raw264.7 macrophage cells and validate the mechanisms involved in the anti-inflammatory effect in LPS-induced ALI/ARDS model. In this study, a rich resource for data mining and guidance for clinical validation is provided. Data analysis and experimental research workflow is detailed in Fig. 1.
Materials and methods
Cell and reagents
Raw264.7 macrophage cell lines were acquired from the Beijing Proteome Research Center, National Center for Protein Sciences (Beijing, China). Fetal bovine serum (FBS), Dulbecco’s modified Eagle’s medium (DMEM), penicillin and streptomycin were purchased from Thermo Fisher Scientific. Lipopolysaccharide (LPS) from Escherichia coli 0111: B4 (in vitro) and O55: B5 (in vivo) were purchased from Sigma-Aldrich. Clofazimine (CFZ, Fig. 2a) was purchased from the Cayman Chemical. Cell Counting Kit-8 (CCK-8) was obtained from Gene-Protein Link. Mouse TNF-a, and IL-1β enzyme-linked immunosorbent assay (ELISA) kits were purchased from Sigma-Aldrich. COX-2, Aim 2, TLR4, NF-κBp65, p-NF-κBp65 and HIF-1α were purchased from Cell Signaling Technology. All other chemicals were of reagent grade.
Variation in cell viability and overview of the LPS and CFZ proteomics data in Raw264.7 macrophage cells. a 3d structure of CFZ. b CFZ reduced cell viability in a concentration-dependent manner. c Cell viability of Raw264.7 cells were exposed to different concentrations of LPS for 24 h. d CCK-8 assay was applied to detect the effects of CFZ on Raw264.7 cells. e Total number of proteins identified in each timepoint (2 h, 8 h, and 24 h) in Raw264.7 macrophage cells. All values are expressed as mean ± SEM of at least three separate experiments. Variation among treatments was determined by one-way ANOVA. *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001; “ns” indicates no significant difference
Cell culture
Raw264.7 cells were grown in culture media (DMEM), supplemented with 10% FBS and 1% penicillin/streptomycin. Cells were cultured at 37 °C in a 5% CO2 humidified incubator, and grown to 80% confluence before use.
CCK-8 assay
A CCK-8 test kit was used according to the manufacturer's instructions. Briefly, Raw264.7 cells were inoculated into 96-well plates at a density of 6 × 103 per well for 24 h. Then, the cells were treated with CFZ and (or) LPS for 24 h. 10% of CCK-8 solution was added into the well and incubated for 1 h at 37 °C away from light. The absorbance was measured at 450 nm by a microplate reader (Bio-Rad) to assess the half-maximal inhibitory concentration (IC50) value and cell proliferation.
Sample preparation for proteomics analysis
First, cells were plated in triplicate wells in six-well plates. After adherence, CFZ (4 μM) or LPS (100 ng/ml) were added to cells and incubated at 37 °C for 0 h, 2 h, 8 h, and 24 h. Then, samples were lysed in lysis buffer supplemented with phosphatase inhibitors and incubated at 95 ℃ for 15 min. After sonication, the lysates were centrifuged at 16,000×g at 4 ℃ for 10 min and the supernatants were collected as whole cell extract. Next, protein concentration was determined by NanoDrop (Thermo Fisher Scientific). Subsequently, samples (100 μg per sample) were digested with trypsin (2 μg per sample) overnight at 37 ℃. Then, peptides were eluted, separated and desalted into fractions. Finally, the samples were vacuum-centrifuged to dryness (Thermo Fisher Scientific), and used for liquid chromatography tandem mass spectrometry (LC–MS/MS) analysis.
Subsequently, peptides were analyzed on an Orbitrap Q-Exactive HF Mass Spectrometer (Thermo-Fisher Scientific) operating in the data-dependent acquisition (DDA) mode. The Proteome Discoverer (Thermo Scientific) was used to analyze the raw MS files against the mouse refseq protein database. Enzyme specificity was set to trypsin. The search set none as a fixed modification, while acetylation (Protein N-term), oxidation (M), and carbamidomethylation (C) were considered variable modifications. Charges of precursor ion were limited to + 2, + 3, and + 4. The protein quantification was represented in intensity-based absolute quantification (iBAQ).
Bioinformatics analysis
Proteins with a probability p value < 0.05, fold change > 2 or < 0.5 and false discovery rate (FDR) < 0.01 were chosen as the mostly affected proteins. Then, the expression pattern of six clusters of dynamic proteins were analyzed by Mfuzz (Kumar and Futschik, 2007). Hierarchical cluster and heatmap analysis of differentially expressed proteins were performed using the ward.D’s hierarchical clustering and heatmap function in the R package. The volcano plots were visualized in Hiplot analysis with the website (https://hiplot.com.cn/advance). Enrichment analysis using Gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG), and the Database for Annotation, Visualization and Integrated Discovery (DAVID, https://david.ncifcrf.gov/home.jsp) were available. Correlation analysis was carried out using Pearson’s correlation. Enrichment p values in GO, KEGG, and correlation analysis less than 0.05 were considered significantly enriched.
ELISA assay
RAW 264.7 macrophage cells were cultured in six-well plates, and four groups were set including control group, LPS group, CFZ group, LPS + CFZ group. The cells were then treated with freshly prepared Escherichia coli LPS (100 ng/ml) and CFZ (4 μM) for 24 h. Control group was treated with equivalent endotoxin-free PBS. According to manufacturer’s protocol, cell free supernatant was obtained after 24 h incubation, and enzyme-linked immunosorbent assay (ELISA) was employed to determine the levels of TNF-α and IL-1β.
LPS‑induced ALI/ARDS model in mice
Six-week-old female C57BL/6 J mice (weighing 20–22 g) were purchased from Beijing Vital River Laboratory Animal Technology Co., Ltd. They were housed in pathogen-free cages with free access to food and water with a 12-h light/dark schedule at 22 ± 2 °C. All study protocols were accredited by the Institutional Animal Care and Use Committee of National Center for Protein Sciences (Beijing, China) (IACUC-20210702-26MT).
Mice were randomly assigned to the following groups (n = 5 per group):
-
Control group: PBS was administered intraperitoneally (i.p.) without LPS stimulation.
-
LPS group: Mice were stimulated with LPS (20 mg/kg).
-
CFZ group: Mice were administered intragastrically with CFZ (25 mg/kg). CFZ was suspended in a 0.05% (w/v) agarose solution.
-
LPS + CFZ group. Mice in this group were stimulated with LPS and then treated with CFZ.
All mice were euthanized (i.p.; 80 mg/kg sodium pentobarbital) 8 h after the last challenge. The lung tissues were quickly removed for further analyses.
Histological analysis
Lung tissues were collected, fixed and embedded in paraffin. Subsequently, the tissues were serially sectioned into 4 μm thickness, stained with hematoxylin and eosin (H&E) and then mounted in neutral gum. Lung injury scores were based on the infiltration of neutrophils (0–4), alveolar oedema (0–4), hyperemia or congestion (0–4) and intra-alveolar hemorrhage, necrosis (0–4). The severity of lung injury was scored as: 0 = normal; 1 = minimal (< 5%); 2 = mild (< 10%); 3 = moderate (15 ‑ 20%); and 4 = severe (> 25%).
Pulmonary wet-to-dry ratio
The lung tissue from each animal was harvested and placed onto pre-weighed tin foil to measure the wet weight (W). The tissue was then placed in an incubator at 37 ℃ for 48 h and obtained dry weight (D). The W/D ratio of the lung tissue was calculated.
Survival rates and weight of mice
Survival rates were studied in another set of experiments: 30 mice were randomly divided into 3 groups [Control group; LPS (60 mg/kg) group; LPS + CFZ (25 mg/kg) group] as described above (n = 10 per group). Mice were weighed every 24 h.
Western blot analysis
Lung tissues were washed three times with ice-cold PBS and lysed in RIPA lysis buffer. Protein concentration was determined using the BCA assay (Thermo Fisher Scientific). Samples were run using 10% sodium dodecyl sulfate (SDS)—polyacrylamide gels (PAGE) and electro-transferred onto nitrocellulose membranes (Bio-Rad). After blocking in 5% non-fat dry milk in TBST at room temperature for 1 h, the membranes were washed and immunoblotted with primary antibodies COX-2 (1:1000, #12,282), TLR4 (1:1000, #14,358), NF-κBp65 (1:1000, #8242), p-NF-κBp65 (1:1000, #3033), AIM 2 (1:1000, #63,660), HIF-1α (1:1000, #36,169) and GAPDH (1:5000, Wuhan Sanying, 60,004-1-Ig) overnight at 4 ℃. Then incubated with a secondary antibody conjugated with horseradish peroxidase (HRP) (1:2000; Beyotime, A0208). Enhanced chemiluminescence reagent (ECL) detection system (Thermo Fisher Scientific) was used to detect immunoreactive bands. All acquired images were calculated by Image J (v1.53e).
Statistical analysis
Data were evaluated using GraphPad Prism 8.3.0 (GraphPad Software, La Jolla, CA). Differences between the groups were performed with Student’s t test or one-way ANOVA method. A p value < 0.05 was set at statistically significant.
Results
In vitro study
Effect of CFZ on CCK-8 assay
First, after 24 h-incubation, the cytotoxic effect of CFZ on Raw264.7 cell lines by CCK-8 assay after treatment was estimated with different doses. The results showed that IC50 of CFZ in the Raw264.7 cells was 4.1 μM (Fig. 2b). The non-toxic concentrations (4 μM) were used for the following experiment. Then, after different concentrations (0.05, 0.1, 0.25, 0.5, 1, 10 and 100 μg/mL) of LPS were applied, the viability of cells remained the same after treatment with 100 μg/mL of LPS compared to the control group (Fig. 2c). Therefore, 100 ng/mL was selected as the optimal concentration of LPS for activating the macrophages. Additionally, the results of CCK-8 assay also revealed that LPS promoted the viability of Raw264.7 cells, while CFZ could significantly reduce the proliferation of Raw264.7 cells (Fig. 2d).
A dynamic protein expression landscape of CFZ and LPS
To study the damage of LPS and CFZ on the Raw264.7 cells, shotgun proteomics was utilized to discover the alteration of protein expression patterns. 4746 and 4766 proteins were identified with quantitative information in this study (Fig. 2e). Then, the correlation between LPS and CFZ proteomic data in each timepoint (0 h, 2 h, 8 h, and 24 h) was analyzed by Mfuzz. In total, it is found that six distinct clusters of temporal patterns representing proteins are regulated differently, indicating different expression kinetics. Among the 4746 proteins of LPS, clusters 1 represents up-regulated proteins, clusters 2 and 3 represent down-regulated proteins, whereas clusters 4, 5, and 6 represent proteins displaying a bimodal expression pattern (Fig. 3a, b; Supplementary Table S1). Among the 4766 proteins of CFZ, clusters 1 and 3 represent proteins are down-regulated, clusters 4 and 5 represent proteins are up-regulated, whereas clusters 2 and 6 represent proteins displaying a bimodal expression pattern (Fig. 3c, d; Supplementary Table S2). By overlaying Mouse Genome Informatics (MGI, http://www.informatics.jax.org/) containing primarily inflammatory response proteins (IRPs), we found, intriguingly, that IRPs mainly showed a low expression level in clusters 1 and 3 of CFZ and a high expression level in clusters 1 of LPS (Fig. 3e). These proteins play critical roles in regulating inflammation, such as SYK, GSDMD, CELF1, NFκB1, MYD88, AIM 2, PTPN2, STAT5A, IL16, NCF1, PIK3AP1, DDX3X, RIPK1, FCGR1, COX2, etc. (Supplementary Table S3). The enrichment of the KEGG pathway included the leishmaniasis, NF-kappa B (NF-κB) signaling pathway, hepatitis B, osteoclast differentiation, tuberculosis, B cell receptor signaling pathway, toll-like receptor signaling pathway, measles, epstein–barr virus infection, etc. (Fig. 3f).
Temporal profiles of protein expression in each timepoint (0 h, 2 h, 8 h, and 24 h) in Raw264.7 macrophage cells. a Mfuzz clustering identified six distinct temporal patterns of proteins expression of LPS. b The left pie chart shows the number of proteins of LPS in six dynamic expression clusters in (b), the right pie chart shows the number of inflammatory proteins of LPS in six dynamic expression clusters in (b). c Mfuzz clustering identified six distinct temporal patterns of proteins expression of CFZ. d The left pie chart shows the number of proteins of CFZ in six dynamic expression clusters in (d), the right pie chart shows the number of inflammatory proteins of CFZ in six dynamic expression clusters in (d). e Venn diagram shows the opposite expression patterns of 30 inflammatory proteins identified by LPS (green) and CFZ (purple) comparisons. f KEGG pathway analysis of opposite protein clusters
Hierarchical cluster analysis of the protein identification and functions
Principal component analysis (PCA) results showed that each sample within treatment groups shared a similar expression pattern, whereas it displayed a distinguished pattern between each time point (i.e., three samples per treatment, LPS and CFZ enriched for four time points, 0, 2, 8, and 24 h, p < 0.05; Fig. 4a). To gain better insight into the difference of protein expression profiles of LPS and CFZ (0 h, 2 h, 8 h, and 24 h), we performed the degree of differential proteins in each sample by hierarchical clustering and heatmap. To reveal the molecular mechanism of proteins in each cluster and understand their biological significance, KEGG pathway analysis were performed on differentially expressed proteins in the LPS and CFZ dataset (ANOVA, FDR < 0.01, p < 0.05). The LPS dataset in cluster 1 mainly involves ribosome, spliceosome, RNA transport, biosynthesis of antibiotics, and phagosome. Cluster 2 proteins participate in systemic lupus erythematosus, spliceosome, alcoholism, viral carcinogenesis, and proteasome. Cluster 3 is enriched in herpes simplex infection, viral carcinogenesis, ubiquitin-mediated proteolysis, alcoholism, spliceosome. Intriguingly, the KEGG analysis of CFZ suggest that cluster 1 is mainly enriched in pathways including biosynthesis of antibiotics, spliceosome, RNA transport, carbon metabolism, and metabolic pathways. Cluster 2 is involved in systemic lupus erythematosus, alcoholism, carbon metabolism, biosynthesis of antibiotics, and viral carcinogenesis. Cluster 3 is mainly associated with spliceosome, biosynthesis of antibiotics, metabolic pathways, carbon metabolism, and citrate cycle (TCA cycle). Overall, the CFZ clustering analysis revealed that KEGG pathway has similar biological features to those obtained from the LPS dataset. (Fig. 4b; Supplementary Table S4 and Table S5).
Essential pathways of CFZ in proteome
The distribution of proteins is identified at each time point (0, 2, 8, and 24 h), then they are represented in Venn diagrams (FDR < 0.01, p < 0.05, fold change > 2 or fold change < 0.5; Fig. 5a). Notably, by comparing the expression levels of total proteins, the differentially expressed proteins (DEPs) at each time point (0, 2, 8, and 24 h) were screened (FDR < 0.01, p < 0.05, fold change > 2 or fold change < 0.5; Fig. 5b). Volcano plots is a scatter graph used to highlight the up-regulated, down-regulated proteins and spatial occurrence in large datasets. As shown in Fig. 5c, the great alterations in the Raw264.7 proteome were induced by LPS and CFZ at the 24 h time point, where 1203 (Supplementary Table S6) and 1332 (Supplementary Table S7) proteins showed significantly changed expression compared to controls, respectively.
To extract molecular mechanisms of proteome from the 1332 DEPs of CFZ, gene ontology (GO) and KEGG enrichment analysis (Fig. 6) were used. The top enriched categories of the biological process (BP) were related to generation of precursor metabolites and energy, aerobic respiration, cellular respiration, energy derivation by oxidation of organic compounds, ATP metabolic process, oxidative phosphorylation, carboxylic acid catabolic process, organic acid catabolic process, nucleotide metabolic process, and purine-containing compound metabolic process. The cellular component (CC) terms of the DEPs were mitochondrial matrix, mitochondrial protein-containing complex, mitochondrial inner membrane, organelle inner membrane, oxidoreductase complex, myelin sheath, organellar ribosome, mitochondrial ribosome, inner mitochondrial membrane protein complex, and respiratory chain complex. The molecular function (MF) terms of the DEPs were NAD binding, structural constituent of ribosome, electron transfer activity, isomerase activity, ribonucleoprotein complex binding, iron-sulfur cluster binding, metal cluster binding, ATP hydrolysis activity, primary active transmembrane transporter activity, and ligase activity. The enrichment of the KEGG pathway included carbon metabolism, oxidative phosphorylation, fatty acid metabolism, HIF-1 signaling pathway, and lysosome.
CFZ inhibits TLR4/NF-kB/HIF-1α pathway and IL-1β and TNF-α release
NF-κB pathway has been defined as a key participant in LPS infections [18, 19]. Meanwhile, hypoxia-inducible factor-1α (HIF-1α) is considered one of endogenous ROS's core cellular targets, which further regulates the hypoxia signaling in the inflammatory response [20]. To further correlate CFZ with inflammation on the levels of transduction pathways, key proteins involved in the pathways implicated in TLR4/NF-κB/HIF-1α signaling pathway were listed (Fig. 7a). Among these protein expressions, there were better correlation in TLR4/NF-κB/HIF-1α transduction pathway (Fig. 7b).
Next, TNF-α and IL-1β expression was measured in the culture supernatants of Raw264.7 cells. It can be seen from the results that compared to the control group, the levels of TNF-α and IL-1β were significantly increased with LPS exposure alone. However, when treated with CFZ, the levels of TNF-α and IL-1β in LPS + CFZ group were markedly reduced compared with the LPS group (Fig. 8a).
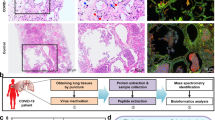
CFZ decreased the pro-inflammatory cytokines and ameliorated ALI following LPS. a The TNF-α and IL-1β level was detected by ELISA kits. All values were expressed as mean ± SEM of at least three separate experiments. b Histopathological changes of lung tissue (200 x) and lung injury score (n = 5 mice/group). c Pulmonary edema was examined by W/D ratio (n = 5 mice/group). d Survival rate of mice (n = 10 mice/group). e Body weight loss (n = 10 mice/group). Data are mean ± SEM. Variation among treatments was determined by one-way ANOVA. *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001
In vivo study
Effect of CFZ on LPS-induced lung parenchyma
As CFZ treatment significantly improved the pro-inflammatory cytokines, we examined whether this treatment reduced lung injury in mice. Intact structure and clear pulmonary alveoli in both control group and CFZ group of mice were observed. Following LPS-induced injury, the histological examination showed an increase in the areas of inflammatory infiltration, collapse of air alveoli, and pulmonary congestion. However, CFZ markedly attenuated the LPS-induced pathological changes (p < 0.05) (Fig. 8b).
Effect of CFZ on the W/D ratio
Lung edema was a predictive marker of the exudative phase of ALI/ARDS [21, 22]. We evaluated it by W/D lung weight ratio. After 8 h of LPS administration, a marked increase was observed in the average wet/dry ratio of lung tissue compared with the control group; however, the LPS + CFZ group, in particular, exhibited a significantly lower level of W/D ratio than the LPS group (p < 0.05) (Fig. 8c).
Effect of CFZ on survival rate and body weight loss of mice challenged with a lethal dose of LPS
The therapeutic effect of CFZ was evaluated on mice challenged with the lethal dose of the LPS (Fig. 8d, e). The survival rate of the mice in LPS group was markedly lower than that in the control group, while the survival rate in the LPS + CFZ group was distinctly higher than that in the LPS group during 72 h observation (p < 0.05). These results showed that CFZ could improve the survival rate of LPS-challenged mice (Fig. 8d). The effects of CFZ were further validated on LPS-induced ALI in mice. Our results suggested that on day 3, mice in the normal group increased a considerable weight, compared to day 0. Mice began to lose weight on day 1 and all weight loss on day 3 following LPS injury. Treatment of CFZ effectively improved the loss and presented a slight weight increase on day 3, compared to LPS group (p < 0.05).
Effect of CFZ on LPS-induced COX-2 and AIM 2 expression
The activation of COX-2 and AIM 2 were detected by western blot analysis to investigate the mechanism underlying the anti-inflammatory effect of CFZ. As expected, LPS application dramatically increased COX-2 and AIM 2 expression compared with the control group. However, compared with the LPS group, CFZ markedly decreased the protein expression (Fig. 9a).
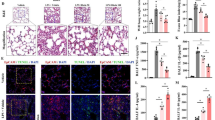
Effect of CFZ on TLR4/NF-κB/HIF-1α pathway in LPS-stimulated ALI. a Western blot corresponding to COX-2 and AIM 2 in lung lysates from mice treated with LPS then CFZ 8 h. b Western blot corresponding to TLR4, NF-κBp65, and p-NF-κBp65 in lung lysates from mice treated with LPS then CFZ 8 h. c Western blot corresponding to HIF-1α in lung lysates from mice treated with LPS then CFZ 8 h. GAPDH was used for loading control. Data are mean ± SEM., n = 5 mice/group. Variation among treatments was determined by one-way ANOVA. *p < 0.05; **p < 0.01; ***p < 0.001
Effect of CFZ on LPS-induced TLR4/NF-κB/HIF-1α expression
The protein expression levels of CFZ on TLR4/NF-κB/HIF-1α pathway were assessed for further investigating the anti-inflammatory mechanisms of CFZ. As a result (Fig. 9b), the protein levels of three essential proteins, including TLR4, NF-κBp65 and HIF-1α were significantly enhanced in LPS group. Conversely, CFZ decreased the expression of these three essential proteins in LPS + CFZ group (p < 0.05).
Discussion
Today, while most COVID-19 cases are asymptomatic or cause only mild symptoms after SARS-CoV-2 infection, 10–20% of the patients develop severe pneumonia and ARDS [23]. ALI is increasingly recognized as one of the most severe acute hyperimmune responses of COVID-19, which can be manifested as dyspnea and gradually develop into ARDS [24]. Numerous studies have also been reported that several conditions, including infection, trauma, or inflammatory pathways are associated with ALI/ARDS [25, 26]. Hence, obtaining an additional therapeutic strategy of ALI/ARDS remains a significant challenge. Many CFZ-related articles focus on its antibiotic function, like treatment for tuberculosis, leprosy, and Salmonella enterica, but anti-inflammatory effect has not yet been elucidated. With this in mind, this study aims to fill this gap in the literature. Here, our results have demonstrated the anti-inflammatory roles of CFZ in both Raw264.7 cells and the mice model.
Proteomics has become the mainstay in systematically studying the expression, function, and interaction of all proteins in cells [27]. This technology can reveal the entire protein change spectrum under specific biological processes and the mechanisms of drugs on the changes of the whole proteome [28]. The researchers performed a comprehensive proteomic analysis to study the damage of LPS and CFZ on the Raw264.7 cells, and they used bioinformatic interpretation to identify the potential alteration of proteins of LPS and CFZ. As a result, 4746 (LPS) and 4766 (CFZ) proteins were identified and contained quantitative information in this study. In addition, mainly 29 proteins showed a low expression level in clusters 1 and 3 of CFZ and a high expression level in clusters 1 of LPS, overlaying MGI containing primarily inflammatory response proteins. As a result, the expression levels of IRPs including SYK, GSDMD, CELF1, NFκB1, MYD88, AIM 2, PTPN2, STAT5A, IL16, NCF1, PIK3AP1, DDX3X, RIPK1, FCGR1, COX-2, etc. Studies have revealed the potential roles of SYK, PTPN2, STAT5A, IL‑16, NCF1, PIK3AP1, and FCGR1 in different inflammatory diseases and autoimmune pathologies [29,30,31,32,33,34,35]. Additionally, GSDMD, AIM 2, and COX-2 have recently been identified as a key role in the pathogenesis of sepsis and COVID-19 [36,37,38]. Celf1 is known as an RNA-binding protein [39]. Hu et al. found that Celf1-deficiency could improve oxidative stress and apoptosis in cardiomyocytes [40]. NFκB1 (p50) is one of the five members of the NF-κB family, which is IκB kinase-like protein [41]. MYD88 is implicated in inflammatory signaling downstream of IL-1R and mammalian TLRs [42]. Recently, a plethora of studies have implicated that massive release of TNF-a, IFN-g, IL-1b, IL-8, MCP-1, and IP-10 seen in ALI/ARDS may probably be linked to NLRP3 inflammasome activation [43]. The stress granule protein DDX3X interacts with NLRP3 to drive inflammasome activation [44]. Ripk1 is a key pharmacological target of regulated cell death as well as in the TNF signaling pathways [45]. Then, KEGG analysis suggested that the IRPs were also enriched in pathways that are strongly associated with NF-kappa B (NF-κB) signaling pathway, B cell receptor signaling pathway, and toll-like receptor (TLR) signaling pathway. NF-kappa B signaling pathway, B cell receptor signaling pathway, and toll-like receptor signaling pathway profoundly affect the activation of Raw264.7 macrophage cells, and thereby modulate the release of inflammatory cytokines and the presentation of immune responses. These results indicate that CFZ could serve as potential therapeutics on Raw264.7 macrophage cells activation.
The above PCA analysis showed that each sample within treatment groups shared a similar expression pattern, whereas displayed a distinguished pattern between each time point. Based on hierarchical clustering and heatmap, the CFZ clustering analysis revealed KEGG pathway with similar biological features to those obtained from the LPS clustering. Our results also showed that CFZ and LPS play a very similar inflammatory and immune function in Raw264.7 cell model.
To ensure the accuracy and reliability of our results, proteins, which had FDR < 0.01, p < 0.05, and fold change > 2 or fold change < 0.5, in each time point (2, 8, and 24 h) of LPS and CFZ compared with 0 h, was considered as significantly expressed proteins. Strikingly, using this cutoff, after 24 h of CFZ exposure in Raw264.7 macrophages, a total of 1332 proteins were significantly changed, including 405 up-regulated and 927 down-regulated. After 24 h of LPS exposure in Raw264.7 macrophages, a total of 1203 proteins were significantly changed, of which 537 were up-regulated and 666 down-regulated. As most of the inflammatory pathological features of Raw264.7 macrophages develop within this time, we focused on the phase (24 h) of the 1332 proteins of CFZ. KEGG pathway analysis showed that DEPs of CFZ were mainly involved in the pathways related to the carbon metabolism, oxidative phosphorylation, TCA cycle, fatty acid metabolism, HIF-1 signaling pathway, and lysosome. The importance of the carbon metabolism, oxidative phosphorylation, TCA cycle, fatty acid metabolism, HIF-1 signaling pathway, and lysosome in physiology and pathophysiology has now been confirmed by many experimental studies in LPS models. For instance, when Raw 264.7 cells were exposed to LPS, several inflammatory reactions could be inhibited by the low doses of carbon monoxide (CO) or endogenous CO [46]. HIF-1α is induced in LPS-activated macrophages, where it is pivotal involved in TCA cycle and glycolytic metabolites as well as the induction of inflammatory response, notably IL-1β. As the mechanisms for CFZ-mediated macrophage protein expression are poorly understood, then key proteins involved in the signaling pathways implicated in TLR4/NF-κB/HIF-1α signaling pathway were listed in this study according to KEGG pathway analysis. Among these protein expressions, there was better correlation in this signaling pathway. These observations suggest CFZ exerted anti-inflammatory activity via TLR4/NF-κB/HIF-1α pathway in macrophages.
Raw264.7 macrophage, an important immunological cell, plays a major role in regulating pro-inflammatory chemokines, cytokines, and mediators in the pathogenesis of ALI/ARDS [47]. Our experiments demonstrated that LPS promoted the viability of Raw264.7 cells, while CFZ could significantly decrease the proliferation of Raw264.7 macrophages. We further evaluated the anti-inflammatory effect of CFZ in Raw264.7 cells models. It was reported that targeting pro-inflammatory cytokines (such as TNF-α and IL-1β) was one of the most promising treatments against inflammatory response during inflammatory diseases [48]. Our study demonstrated that CFZ inhibited TNF-α and IL-1β production in the Raw264.7 cells caused by LPS. Similar to Yoon et al., they also reported that CFZ decreased TNF-α production and boosted IL-1RA secretion after treatment with LPS in macrophages [49].
Animal models can provide highlight ways in the complex pathophysiology of human disease. Therefore, in the present study, LPS model was also applied to examine the effect of CFZ on lung and lethality in mice. In our work, it is demonstrated that CFZ treatment decreased the severity of LPS-induced lung injury by improving the inflammatory infiltration into the lung. Previous studies have shown that pulmonary edema is the hallmark of ALI/ARDS in LPS model [25, 26]. Alveolar epithelial barrier dysfunction in response to inflammatory reaction results in lung edema in ALI/ARDS [50]. Our results also suggest that CFZ significantly attenuated the lung edema. LPS challenge contributed to a 40% survival rate within 72 h, while administration of CFZ improved the survival rate to 80%. Besides, a significant deterioration of weight loss for the first 2 days was observed, followed by weight recovery in mice treatment with CFZ compared to mice receiving LPS. The significance of TLR4/NF-κB/HIF-1α in LPS-induced ALI/ARDS model has been previously reported. Subsequently, we further investigated the anti-inflammatory effect of CFZ on TLR4/NF-κB/HIF-1α pathway in LPS-induced lung injury mice. COX-2, an evolutionary enzyme, plays a central role in controlling host defense response to viral and bacterial infections [51]. Several lines of evidence demonstrated that COX-2 inhibitor is one of the major targets in treating ALI model induced by LPS in mice [52]. AIM 2, which can be expressed in the cytosol, is a member of innate immune sensors. During the inflammatory response, expression levels of COX-2 and AIM 2 are related to the NF-κB pathway [53]. Recent studies have alluded to the fact that the NF‐κB and HIF crosstalk. For instance, macrophages or mice infected with LPS signaled through NF-κB and promoted the inflammatory cytokine to modulate HIF-1α expression. We found that LPS activates the expression of key proteins involved in TLR4/NF-κB/HIF-1α signaling, including the TLR4, NF-κB, and HIF-1α, as well as inflammatory enzyme (COX-2) and the inflammasome (AIM 2), and its stimulation was reversed through CFZ. Taken together, these results demonstrated the strong anti-inflammatory effects of CFZ on LPS-induce lung injury.
Conclusion
In summary, this project revealed that CFZ markedly inhibits LPS-challenged inflammatory response in vitro and in vivo. Through performing a systematically proteomic analysis on Raw264.7 macrophage, the progressive alteration of protein expression of LPS and CFZ dataset has been explored. Our LPS model demonstrated that CFZ decreases the level of pro-inflammatory cytokine, attenuates lung injury and pulmonary edema, and reduces the lethality of septic shock. In particular, our results show that TLR4/NF-κB/HIF-1α is a critical signaling of CFZ-mediated anti-inflammatory responses. Thus, the study of quantitative proteomic data reveals valuable information to understand the anti-inflammatory effects of CFZ and provide potential therapeutic strategies for infectious diseases.
Data sharing statement
The original contributions presented in the study are included in the article/Supplementary Material, further inquiries can be directed to the corresponding author/s. The raw data used to support the findings of this study are available online at: https://www.jianguoyun.com/p/DUJ_XZ8QsIisChjmt7EE.
References
Li J, Liu HH, Yin XD, Li CC, Wang J. COVID-19 illness and autoimmune diseases: recent insights. Inflamm Res. 2021;70(4):407–28. https://doi.org/10.1007/s00011-021-01446-1.
Kim JS, Lee JY, Yang JW, Lee KH, Effenberger M, Szpirt W, et al. Immunopathogenesis and treatment of cytokine storm in COVID-19. Theranostics. 2021;11(1):316–29. https://doi.org/10.7150/thno.49713.
Asselah T, Durantel D, Pasmant E, Lau G, Schinazi RF. COVID-19: Discovery, diagnostics and drug development. J Hepatol. 2021;74(1):168–84. https://doi.org/10.1016/j.jhep.2020.09.031.
Kassirian S, Taneja R, Mehta S. Diagnosis and management of acute respiratory distress syndrome in a time of COVID-19. Diagnostics (Basel). 2020;10(12):1053. https://doi.org/10.3390/diagnostics10121053.
Yuan S, Yin X, Meng X, Chan JF, Ye ZW, Riva L, et al. Clofazimine broadly inhibits coronaviruses including SARS-CoV-2. Nature. 2021;593(7859):418–23. https://doi.org/10.1038/s41586-021-03431-4.
Fischer M. Leprosy—an overview of clinical features, diagnosis, and treatment. J Dtsch Dermatol Ges. 2017;15(8):801–27. https://doi.org/10.1111/ddg.13301.
Lange C, Dheda K, Chesov D, Mandalakas AM, Udwadia Z, Horsburgh CR Jr. Management of drug-resistant tuberculosis. Lancet. 2019;394(10202):953–66. https://doi.org/10.1016/S0140-6736(19)31882-3.
Yoon GS, Keswani RK, Sud S, Rzeczycki PM, Murashov MD, Koehn TA, et al. Clofazimine biocrystal accumulation in macrophages upregulates interleukin 1 receptor antagonist production to induce a systemic anti-inflammatory state. Antimicrob Agents Chemother. 2016;60(6):3470–9. https://doi.org/10.1128/AAC.00265-16.
Kumar H, Chattopadhyay S, Das N, Shree S, Patel D, Mohapatra J, et al. Leprosy drug clofazimine activates peroxisome proliferator-activated receptor-γ and synergizes with imatinib to inhibit chronic myeloid leukemia cells. Haematologica. 2020;105(4):971–86. https://doi.org/10.3324/haematol.2018.194910.
Selby W, Pavli P, Crotty B, Florin T, Radford-Smith G, Gibson P, et al. Antibiotics in Crohn’s Disease Study Group. Two-year combination antibiotic therapy with clarithromycin, rifabutin, and clofazimine for Crohn’s disease. Gastroenterology. 2007;132(7):2313–9. https://doi.org/10.1053/j.gastro.2007.03.031.
Goreti Catorze M, Pereira F, Fonseca F, Morbey A, Assis PF. Pyoderma gangrenosum associated with sclerosing cholangitis, type 1 diabetes mellitus and ulcerative colitis. J Eur Acad Dermatol Venereol. 2001;15(3):257–9. https://doi.org/10.1046/j.1468-3083.2001.00260.x.
Coulson GB, Johnson BK, Zheng H, Colvin CJ, Fillinger RJ, Haiderer ER, et al. Targeting Mycobacterium tuberculosis sensitivity to thiol stress at acidic ph kills the bacterium and potentiates antibiotics. Cell Chem Biol. 2017;24(8):993-1004.e4. https://doi.org/10.1016/j.chembiol.2017.06.018.
Lu P, Heineke MH, Koul A, Andries K, Cook GM, Lill H, et al. The cytochrome bd-type quinol oxidase is important for survival of Mycobacterium smegmatis under peroxide and antibiotic-induced stress. Sci Rep. 2015;27(5):10333. https://doi.org/10.1038/srep10333.
Chen H, Bai C, Wang X. The value of the lipopolysaccharide-induced acute lung injury model in respiratory medicine. Expert Rev Respir Med. 2010;4(6):773–83. https://doi.org/10.1586/ers.10.71.
Dutra Silva J, Su Y, Calfee CS, Delucchi KL, Weiss D, McAuley DF, et al. Mesenchymal stromal cell extracellular vesicles rescue mitochondrial dysfunction and improve barrier integrity in clinically relevant models of ARDS. Eur Respir J. 2021;58(1):2002978. https://doi.org/10.1183/13993003.02978-2020.
Yang B, Wang R, Ji LL, Li XP, Li XH, Zhou HG, et al. Exploration of the function of ginsenoside RD attenuates lipopolysaccharide-induced lung injury: a study of network pharmacology and experimental validation. Shock. 2022;57(2):212–20. https://doi.org/10.1097/SHK.0000000000001824.
Park EJ, Kim YM, Kim HJ, Chang KC. Luteolin activates ERK1/2- and Ca2+-dependent HO-1 induction that reduces LPS-induced HMGB1, iNOS/NO, and COX-2 expression in RAW264.7 cells and mitigates acute lung injury of endotoxin mice. Inflamm Res. 2018;67(5):445–53. https://doi.org/10.1007/s00011-018-1137-8.
Facchin BM, Dos Reis GO, Vieira GN, Mohr ETB, da Rosa JS, Kretzer IF, Demarchi IG, Dalmarco EM. Inflammatory biomarkers on an LPS-induced RAW 264.7 cell model: a systematic review and meta-analysis. Inflamm Res. 2022;71(7–8):741–58.
Yang B, Ma L, Wei Y, Cui Y, Li X, Wei Y, et al. Isorhamnetin alleviates lipopolysaccharide-induced acute lung injury by inhibiting mTOR signaling pathway. Immunopharmacol Immunotoxicol. 2022;21:1–13. https://doi.org/10.1080/08923973.2022.2052892.
Tian M, Liu W, Li X, Zhao P, Shereen MA, Zhu C, et al. HIF-1α promotes SARS-CoV-2 infection and aggravates inflammatory responses to COVID-19. Signal Transduct Target Ther. 2021;6(1):308. https://doi.org/10.1038/s41392-021-00726-w.
Mokra D, Kosutova P. Biomarkers in acute lung injury. Respir Physiol Neurobiol. 2015;209:52–8. https://doi.org/10.1016/j.resp.2014.10.006.
Shah D, Das P, Acharya S, Agarwal B, Christensen DJ, Robertson SM, et al. Small immunomodulatory molecules as potential therapeutics in experimental murine models of acute lung injury (ALI)/acute respiratory distress syndrome (ARDS). Int J Mol Sci. 2021;22(5):2573. https://doi.org/10.3390/ijms22052573.
Tang YW, Schmitz JE, Persing DH, Stratton CW. Laboratory diagnosis of COVID-19: current issues and challenges. J Clin Microbiol. 2020;58(6):e00512-e520. https://doi.org/10.1128/JCM.00512-20.
Kyriakopoulos C, Ntritsos G, Gogali A, Milionis H, Evangelou E, Kostikas K. Tocilizumab administration for the treatment of hospitalized patients with COVID-19: a systematic review and meta-analysis. Respirology. 2021;26(11):1027–40. https://doi.org/10.1111/resp.14152.
Huppert LA, Matthay MA, Ware LB. Pathogenesis of acute respiratory distress syndrome. Semin Respir Crit Care Med. 2019;40(1):31–9. https://doi.org/10.1055/s-0039-1683996.
Yang B, Ni YF, Wang WC, Du HY, Zhang H, Zhang L, et al. Melatonin attenuates intestinal ischemia–reperfusion-induced lung injury in rats by upregulating N-myc downstream-regulated gene 2. J Surg Res. 2015;194(1):273–80. https://doi.org/10.1016/j.jss.2014.11.018.
Wilhelm M, Schlegl J, Hahne H, Gholami AM, Lieberenz M, Savitski MM, et al. Mass-spectrometry-based draft of the human proteome. Nature. 2014;509(7502):582–7. https://doi.org/10.1038/nature13319.
Lundberg E, Borner GHH. Spatial proteomics: a powerful discovery tool for cell biology. Nat Rev Mol Cell Biol. 2019;20(5):285–302. https://doi.org/10.1038/s41580-018-0094-y.
Wang L, Aschenbrenner D, Zeng Z, Cao X, Mayr D, Mehta M, et al. Gain-of-function variants in SYK cause immune dysregulation and systemic inflammation in humans and mice. Nat Genet. 2021;53(4):500–10. https://doi.org/10.1038/s41588-021-00803-4.
Spalinger MR, Sayoc-Becerra A, Santos AN, Shawki A, Canale V, Krishnan M, et al. PTPN2 regulates interactions between macrophages and intestinal epithelial cells to promote intestinal barrier function. Gastroenterology. 2020;159(5):1763-1777.e14. https://doi.org/10.1053/j.gastro.2020.07.004.
Hedl M, Sun R, Huang C, Abraham C. STAT3 and STAT5 signaling thresholds determine distinct regulation for Innate receptor-induced inflammatory cytokines, and STAT3/STAT5 disease variants modulate these outcomes. J Immunol. 2019;203(12):3325–38. https://doi.org/10.4049/jimmunol.1900031.
Normandin E, Holroyd KB, Collens SI, Shaw BM, Siddle KJ, Adams G, et al. Intrathecal inflammatory responses in the absence of SARS-CoV-2 nucleic acid in the CSF of COVID-19 hospitalized patients. J Neurol Sci. 2021;15(430): 120023. https://doi.org/10.1016/j.jns.2021.120023.
Holmdahl R, Sareila O, Olsson LM, Bäckdahl L, Wing K. Ncf1 polymorphism reveals oxidative regulation of autoimmune chronic inflammation. Immunol Rev. 2016;269(1):228–47. https://doi.org/10.1111/imr.12378.
Deason K, Troutman TD, Jain A, Challa DK, Mandraju R, Brewer T, et al. BCAP links IL-1R to the PI3K-mTOR pathway and regulates pathogenic Th17 cell differentiation. J Exp Med. 2018;215(9):2413–28. https://doi.org/10.1084/jem.20171810.
Negishi-Koga T, Gober HJ, Sumiya E, Komatsu N, Okamoto K, Sawa S, et al. Immune complexes regulate bone metabolism through FcRγ signalling. Nat Commun. 2015;31(6):6637. https://doi.org/10.1038/ncomms7637.
Ma J, Zhu F, Zhao M, Shao F, Yu D, Ma J, et al. SARS-CoV-2 nucleocapsid suppresses host pyroptosis by blocking Gasdermin D cleavage. EMBO J. 2021;40(18): e108249. https://doi.org/10.15252/embj.2021108249.
Junqueira C, Crespo Â, Ranjbar S, Ingber J, Parry B, Ravid S, et al. SARS-CoV-2 infects blood monocytes to activate NLRP3 and AIM2 inflammasomes, pyroptosis and cytokine release. medRxiv [Preprint]. 2021. https://doi.org/10.1101/2021.03.06.21252796.
Baghaki S, Yalcin CE, Baghaki HS, Aydin SY, Daghan B, Yavuz E. COX2 inhibition in the treatment of COVID-19: Review of literature to propose repositioning of celecoxib for randomized controlled studies. Int J Infect Dis. 2020;101:29–32. https://doi.org/10.1016/j.ijid.2020.09.1466.
Hu X, Wu P, Liu B, et al. RNA-binding protein CELF1 promotes cardiac hypertrophy via interaction with PEBP1 in cardiomyocytes. Cell Tissue Res. 2022;387(1):111–21. https://doi.org/10.1007/s00441-021-03541-5.
Chang KT, Wang LH, Lin YM, Cheng CF, Wang GS. CELF1 promotes vascular endothelial growth factor degradation resulting in impaired microvasculature in heart failure. FASEB J. 2021;35(5): e21512. https://doi.org/10.1096/fj.202002553R.
Vallabhapurapu S, Karin M. Regulation and function of NF-kappaB transcription factors in the immune system. Annu Rev Immunol. 2009;27:693–733. https://doi.org/10.1146/annurev.immunol.021908.132641.
Di Padova F, Quesniaux VFJ, Ryffel B. MyD88 as a therapeutic target for inflammatory lung diseases. Expert Opin Ther Targets. 2018;22(5):401–8. https://doi.org/10.1080/14728222.2018.1464139.
Root-Bernstein R. Innate receptor activation patterns involving TLR and NLR synergisms in COVID-19, ALI/ARDS and sepsis cytokine storms: a review and model making novel predictions and therapeutic suggestions. Int J Mol Sci. 2021;22(4):2108. https://doi.org/10.3390/ijms22042108.
Samir P, Kesavardhana S, Patmore DM, Gingras S, Malireddi RKS, Karki R, et al. DDX3X acts as a live-or-die checkpoint in stressed cells by regulating NLRP3 inflammasome. Nature. 2019;573(7775):590–4. https://doi.org/10.1038/s41586-019-1551-2.
Degterev A, Ofengeim D, Yuan J. Targeting RIPK1 for the treatment of human diseases. Proc Natl Acad Sci U S A. 2019;116(20):9714–22. https://doi.org/10.1073/pnas.1901179116.
Dias-Pedroso D, Ramalho JS, Sardão VA, Jones JG, Romão CC, Oliveira PJ, et al. Carbon monoxide-neuroglobin axis targeting metabolism against inflammation in BV-2 microglial cells. Mol Neurobiol. 2022;59(2):916–31. https://doi.org/10.1007/s12035-021-02630-4.
Chen X, Tang J, Shuai W, Meng J, Feng J, Han Z. Macrophage polarization and its role in the pathogenesis of acute lung injury/acute respiratory distress syndrome. Inflamm Res. 2020;69(9):883–95. https://doi.org/10.1007/s00011-020-01378-2.
Fara A, Mitrev Z, Rosalia RA, Assas BM. Cytokine storm and COVID-19: a chronicle of pro-inflammatory cytokines. Open Biol. 2020;10(9): 200160. https://doi.org/10.1098/rsob.200160.
Yoon GS, Sud S, Keswani RK, Baik J, Standiford TJ, Stringer KA, et al. Phagocytosed clofazimine biocrystals can modulate innate immune signaling by inhibiting TNFα and boosting IL-1RA secretion. Mol Pharm. 2015;12(7):2517–27. https://doi.org/10.1021/acs.molpharmaceut.5b00035.
Berthiaume Y, Matthay MA. Alveolar edema fluid clearance and acute lung injury. Respir Physiol Neurobiol. 2007;159(3):350–9. https://doi.org/10.1016/j.resp.2007.05.010.
Rumzhum NN, Ammit AJ. Cyclooxygenase 2: its regulation, role and impact in airway inflammation. Clin Exp Allergy. 2016;46(3):397–410. https://doi.org/10.1111/cea.12697.
Izadparast F, Riahi-Zajani B, Yarmohammadi F, Hayes AW, Karimi G. Protective effect of berberine against LPS-induced injury in the intestine: a review. Cell Cycle. 2022;19:1–14. https://doi.org/10.1080/15384101.2022.2100682.
Li Y, Huang B, Ye T, Wang Y, Xia D, Qian J. Physiological concentrations of bilirubin control inflammatory response by inhibiting NF-κB and inflammasome activation. Int Immunopharmacol. 2020;84: 106520. https://doi.org/10.1016/j.intimp.2020.106520.
Acknowledgements
The work is supported by grants from the Natural Science Foundation of Hebei Province (no. A2021202006), the Hebei Provincial Postdoctoral Science Foundation (no. B2021005005), the Open Projects of State Key Laboratory for Strength and Vibration of Mechanical Structures (no. SV2021-KF-15), and the Natural Science Foundation of Tianjin (no. 21JCQNJC01090). We thank the State Key Laboratory of Proteomics, Beijing Proteome Research Center for their kind technical help.
Funding
The authors declare no support from any organizations for the submitted work. The design of the study, theanalyses and the writing of the manuscript were solely the responsibility of the authors.
Author information
Authors and Affiliations
Contributions
BY: conceptualization, project administration, methodology, writing—original draft. ZG: methodology, project administration. Q-SL: methodology, project administration. X-YZ: methodology, proteomics analysis. LS: proteomics analysis, software. Y-NW: project administration, resources. X-YW: methodology, software. L-LJ: methodology, H-LX: methodology. HX: methodology. F-KF: methodology. X-PL: resources. WL: resources. RW: supervision, project administration, writing—original draft. G-SW: conceptualization, writing—review and editing.
Corresponding authors
Ethics declarations
Conflict of interest
None of the authors have any conflicts of interest to declare.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yang, B., Gao, Z., Li, QS. et al. Proteomic analysis and identification reveal the anti-inflammatory mechanism of clofazimine on lipopolysaccharide-induced acute lung injury in mice. Inflamm. Res. 71, 1327–1345 (2022). https://doi.org/10.1007/s00011-022-01623-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00011-022-01623-w