Abstract
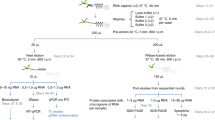
Characterizing protein–protein and protein–RNA interaction networks is a fundamental step to understanding the function of an RNA-binding protein. In many cases, these interactions are transient and highly dynamic. Therefore, capturing stable as well as transient interactions in living cells for the identification of protein-binding partners and the mapping of RNA-binding sequences is key to a successful establishment of the molecular interaction network. In this chapter, we will describe a method for capturing the molecular interactions in living cells using formaldehyde as a crosslinker and enriching a specific RNA–protein complex from cell extracts followed by mass spectrometry and Next-Gen sequencing analyses.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Glisovic T, Bachorik JL, Yong J et al (2008) RNA-binding proteins and post-transcriptional gene regulation. FEBS Lett 582(14):1977–1986. doi:10.1016/j.febslet.2008.03.004
Lunde BM, Moore C, Varani G (2007) RNA-binding proteins: modular design for efficient function. Nat Rev Mol Cell Biol 8(6):479–490
Mili S, Steitz JA (2004) Evidence for reassociation of RNA-binding proteins after cell lysis: implications for the interpretation of immunoprecipitation analyses. RNA 10(11):1692–1694. doi:10.1261/rna.7151404
Dreyfuss G, Choi YD, Adam SA (1984) Characterization of heterogeneous nuclear RNA–protein complexes in vivo with monoclonal antibodies. Mol Cell Biol 4(6):1104–1114
Huang Y, Steitz JA (2001) Splicing factors SRp20 and 9G8 promote the nucleocytoplasmic export of mRNA. Mol Cell 7(4):899–905. doi:10.1016/S1097-2765(01)00233-7
Mayrand S, Setyono B, Greenberg JR et al (1981) Structure of nuclear ribonucleoprotein: identification of proteins in contact with poly(A) + heterogeneous nuclear RNA in living HeLa cells. J Cell Biol 90(2):380–384. doi:10.1083/jcb.90.2.380
Singh G, Ricci EP, Moore MJ (2014) RIPiT-seq: a high-throughput approach for footprinting RNA:protein complexes. Methods 65(3):320–332. doi:10.1016/j.ymeth.2013.09.013
Hafner M, Landthaler M, Burger L et al (2010) Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell 141(1):129–141. doi:10.1016/j.cell.2010.03.009
Jackson V (1978) Studies on histone organization in the nucleosome using formaldehyde as a reversible cross-linking agent. Cell 15(3):945–954. doi:10.1016/0092-8674(78)90278-7
Orlando V, Strutt H, Paro R (1997) Analysis of chromatin structure by in vivo formaldehyde cross-linking. Methods 11(2):205–214. doi:10.1006/meth.1996.0407
Niranjanakumari S, Lasda E, Brazas R et al (2002) Reversible cross-linking combined with immunoprecipitation to study RNA–protein interactions in vivo. Methods 26(2):182–190. doi:10.1016/S1046-2023(02)00021-X
Fraenkel-Conrat H, Olcott HS (1948) Reaction of formaldehyde with proteins: VI. Cross-linking of amino groups with phenol, imidazole, or indole groups. J Biol Chem 174(3):827–843
Metz B, Kersten GFA, Hoogerhout P et al (2004) Identification of formaldehyde-induced modifications in proteins: reactions with model peptides. J Biol Chem 279(8):6235–6243. doi:10.1074/jbc.M310752200
Yong J, Kasim M, Bachorik JL et al (2010) Gemin5 delivers snRNA precursors to the SMN complex for snRNP biogenesis. Mol Cell 38(4):551–562. doi:10.1016/j.molcel.2010.03.014
Morozova O, Marra MA (2008) Applications of next-generation sequencing technologies in functional genomics. Genomics 92(5):255–264. doi:10.1016/j.ygeno.2008.07.001
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer Science+Business Media New York
About this protocol
Cite this protocol
Yeh, HS., Chang, JW., Yong, J. (2016). Ribo-Proteomics Approach to Profile RNA–Protein and Protein–Protein Interaction Networks. In: Lin, RJ. (eds) RNA-Protein Complexes and Interactions. Methods in Molecular Biology, vol 1421. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-3591-8_14
Download citation
DOI: https://doi.org/10.1007/978-1-4939-3591-8_14
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-3589-5
Online ISBN: 978-1-4939-3591-8
eBook Packages: Springer Protocols