Abstract
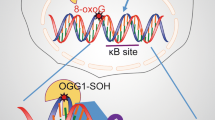
Oxidative DNA modifications are a major challenge to genomic stability. One of the most abundant oxidative DNA base damage is the 8-oxoG, a mutagenic lesion that needs to be efficiently removed to avoid G/C to T/A transversions. It has recently been postulated that 8-oxoG could also act as an epigenetic mark. There is indeed increasing evidence for the key roles played by 8-oxoG and OGG1, the DNA glycosylase responsible for initiating its removal through the base excision repair (BER) pathway, in the regulation of transcription. While it seems clear that there should be a tight coordination between both processes to avoid collision between DNA repair and transcription machineries, there is a real need in the field to better conciliate both aspects that are essential for the maintenance and the expression of our genome. In this review, we will discuss the present knowledge concerning the role of OGG1 in both DNA repair and transcriptional regulation in order to shed light on the coordination between these two major cellular processes.
Similar content being viewed by others
References
Abdella R, Talyzina A, Chen S, Inouye CJ, Tjian R, He Y (2021) Structure of the human Mediator-bound transcription preinitiation complex. Science 372:52–56. https://doi.org/10.1126/science.abg3074
Adamowicz M, Hailstone R, Demin AA, Komulainen E, Hanzlikova H, Brazina J, Gautam A, Wells SE, Caldecott KW (2021) XRCC1 protects transcription from toxic PARP1 activity during DNA base excision repair. Nat Cell Biol 23:1287–1298. https://doi.org/10.1038/s41556-021-00792-w
Aguilera-Aguirre L, Hosoki K, Bacsi A, Riadak Z, Sur S, Hedge ML, Tian B, Saavedra-Molina A, Brasier AR, Ba X, Boldogh I (2015) Whole transcriptome analysis reveals a role for OGG1-initiated DNA repair signaling in airway remodeling. Free Radic Biol Med 89:20–33. https://doi.org/10.1016/j.freeradbiomed.2015.07.007.Whole
Alberti S, Hyman AA (2021) Biomolecular condensates at the nexus of cellular stress, protein aggregation disease and ageing. Nat Rev Mol Cell Biol 22:196–213. https://doi.org/10.1038/s41580-020-00326-6
Allgayer J, Kitsera N, Bartelt S, Epe B, Khobta A (2016) Widespread transcriptional gene inactivation initiated by a repair intermediate of 8-oxoguanine. Nucleic Acids Res. https://doi.org/10.1093/nar/gkw473
Al-Mehdi AB, Pastukh VM, Swiger BM, Reed DJ, Patel MR, Bardwell GC, Pastukh VV, Alexeyev MF, Gillespie MN (2013) Perinuclear mitochondrial clustering creates an oxidant-rich nuclear domain required for hypoxia-induced transcription. Sci Signal 5:1–20
Amente S, Bertoni A, Morano A, Lania L, Avvedimento EV, Majello B (2010) LSD1-mediated demethylation of histone H3 lysine 4 triggers Myc-induced transcription. Oncogene 29:3691–3702. https://doi.org/10.1038/onc.2010.120
Amouroux R, Campalans A, Epe B, Radicella JP (2010) Oxidative stress triggers the preferential assembly of base excision repair complexes on open chromatin regions. Nucleic Acids Res 38:2878–2890. https://doi.org/10.1093/nar/gkp1247
An J, Yin M, Yin J, Wu S, Selby CP, Yang Y, Sancar A, Xu G-L, Qian M, Hu J (2021) Genome-wide analysis of 8-oxo-7,8-dihydro-2’-deoxyguanosine at single-nucleotide resolution unveils reduced occurrence of oxidative damage at G-quadruplex sites. Nucleic Acids Res 49:12252–12267. https://doi.org/10.1093/nar/gkab1022
André KM, Sipos EH, Soutourina J (2020) Mediator roles going beyond transcription. Trends Genet 37:224–234. https://doi.org/10.1016/j.tig.2020.08.015
Bhakat KK, Mantha AK, Mitra S (2009) Transcriptional regulatory functions of mammalian AP-endonuclease (APE1/Ref-1), an essential multifunctional protein. Antioxid Redox Signal 11:621–637. https://doi.org/10.1089/ars.2008.2198
Bhakat KK, Sengupta S, Mitra S (2020) Fine-tuning of DNA base excision/strand break repair via acetylation. DNA Repair (Amst) 93:102931. https://doi.org/10.1016/j.dnarep.2020.102931
Bohm KA, Hodges AJ, Czaja W, Selvam K, Smerdon MJ, Mao P, Wyrick JJ (2021) Distinct roles for RSC and SWI/SNF chromatin remodelers in genomic excision repair. Genome Res 31:1047–1059. https://doi.org/10.1101/gr.274373.120
Boiteux S, Coste F, Castaing B (2017) Repair of 8-oxo-7,8-dihydroguanine in prokaryotic and eukaryotic cells: properties and biological roles of the Fpg and OGG1 DNA N-glycosylases. Free Radic Biol Med 107:179–201. https://doi.org/10.1016/j.freeradbiomed.2016.11.042
Bravard A, Vacher M, Gouget B, Coutant A, de Boisferon FH, Marsin S, Chevillard S, Radicella JP (2006) Redox regulation of human OGG1 activity in response to cellular oxidative stress. Mol Cell Biol 26:7430–7436. https://doi.org/10.1128/MCB.00624-06
Campalans A, Kortulewski T, Amouroux R, Menoni H, Vermeulen W, Radicella JP (2013) Distinct spatiotemporal patterns and PARP dependence of XRCC1 recruitment to single-strand break and base excision repair. Nucleic Acids Res 41:3115–3129. https://doi.org/10.1093/nar/gkt025
Campalans A, Moritz E, Kortulewski T, Biard D, Epe B, Radicella JP (2015) Interaction with OGG1 Is required for efficient recruitment of XRCC1 to base excision repair and maintenance of genetic stability after exposure to oxidative stress. Mol Cell Biol 35:1648–1658. https://doi.org/10.1128/mcb.00134-15
Chakraborty A, Tapryal N, Islam A, Mitra S, Hazra T (2021a) Transcription coupled base excision repair in mammalian cells: so little is known and so much to uncover. DNA Repair (Amst) 107:103204. https://doi.org/10.1016/j.dnarep.2021.103204
Chakraborty U, Shen ZJ, Tyler J (2021b) Chaperoning histones at the DNA repair dance. DNA Repair (Amst) 108:103240. https://doi.org/10.1016/j.dnarep.2021.103240
Charles Richard JL, Shukla MS, Menoni H, Ouararhni K, Lone IN, Roulland Y, Papin C, Ben Simon E, Kundu T, Hamiche A et al (2016) FACT assists base excision repair by boosting the remodeling activity of RSC. PLoS Genet 12:1–21. https://doi.org/10.1371/journal.pgen.1006221
Chen M, Liang J, Ji H, Yang Z, Altilia S, Hu B, Schronce A, McDermott MSJ, Schools GP, Lim CU et al (2017) CDK8/19 Mediator kinases potentiate induction of transcription by NFκB. Proc Natl Acad Sci U S A 114:10208–10213. https://doi.org/10.1073/pnas.1710467114
Chiolo I, Minoda A, Colmenares SU, Polyzos A, Costes SV, Karpen GH (2011) Double-strand breaks in heterochromatin move outside of a dynamic HP1a domain to complete recombinational repair. Cell 144:732–744. https://doi.org/10.1016/j.cell.2011.02.012
Cho I, Tsai PF, Lake RJ, Basheer A, Fan HY (2013) ATP-dependent chromatin remodeling by Cockayne syndrome protein B and NAP1-like histone chaperones is required for efficient transcription-coupled DNA repair. PLoS Genet 9. https://doi.org/10.1371/journal.pgen.1003407
Clark DW, Phang T, Edwards MG, Geraci MW, Gillespie MN (2012) Promoter G-quadruplex sequences are targets for base oxidation and strand cleavage during hypoxia-induced transcription. Free Radic Biol Med 53:51–59. https://doi.org/10.1016/j.freeradbiomed.2012.04.024
Cogoi S, Ferino A, Miglietta G, Pedersen EB, Xodo LE (2018) The regulatory G4 motif of the Kirsten ras (KRAS) gene is sensitive to guanine oxidation: Implications on transcription. Nucleic Acids Res 46:661–676. https://doi.org/10.1093/nar/gkx1142
Collins AR, Cadet J, Moller L, Poulsen HE, Viña J (2004) Are we sure we know how to measure 8-oxo-7,8-dihydroguanine in DNA from human cells? Arch Biochem Biophys 423:57–65. https://doi.org/10.1016/j.abb.2003.12.022
Czaja W, Mao P, Smerdon MJ (2014) Chromatin remodeling complex RSC promotes base excision repair in chromatin of Saccharomyces cerevisiae. DNA Repair (Amst) 16:35–43. https://doi.org/10.1016/j.dnarep.2014.01.002
D’Augustin O, Huet S, Campalans A, Radicella JP (2020) Lost in the crowd: how does human 8-oxoguanine DNA glycosylase 1 (OGG1) find 8-oxoguanine in the genome? Int J Mol Sci 21:1–18. https://doi.org/10.3390/ijms21218360
D’Errico M, Parlanti E, Teson M, De Jesus BMB, Degan P, Calcagnile A, Jaruga P, Bjørås M, Crescenzi M, Pedrini AM et al (2006) New functions of XPC in the protection of human skin cells from oxidative damage. EMBO J 25:4305–4315. https://doi.org/10.1038/sj.emboj.7601277
D’Errico M, Parlanti E, Teson M, Degan P, Lemma T, Calcagnile A, Iavarone I, Jaruga P, Ropolo M, Pedrini AM et al (2007) The role of CSA in the response to oxidative DNA damage in human cells. Oncogene 26:4336–4343. https://doi.org/10.1038/sj.onc.1210232
Dianov G, Bischoff C, Sunesen M, Bohr VA (1999) Repair of 8-oxoguanine in DNA is deficient in Cockayne syndrome group B cells. Nucleic Acids Res 27:1365–1368. https://doi.org/10.1038/sj.onc.1205994
Ding Y, Fleming AM, Burrows CJ (2017) Sequencing the mouse genome for the oxidatively modified base 8-Oxo-7,8-dihydroguanine by OG-Seq. J Am Chem Soc 139:2569–2572. https://doi.org/10.1021/jacs.6b12604
El Khattabi L, Zhao H, Kalchschmidt J, Young N, Jung S, Van Blerkom P, Kieffer-Kwon P, Kieffer-Kwon KR, Park S, Wang X et al (2019) A pliable mediator acts as a functional rather than an architectural bridge between promoters and enhancers. Cell 178:1145–1158.e20. https://doi.org/10.1016/j.cell.2019.07.011
Fang Y, Zou P (2020) Genome-wide mapping of oxidative DNA damage via engineering of 8-oxoguanine DNA glycosylase. Biochemistry 59:85–89. https://doi.org/10.1021/acs.biochem.9b00782
Ferrand J, Plessier A, Polo SE (2021) Control of the chromatin response to DNA damage: histone proteins pull the strings. Semin Cell Dev Biol 113:75–87. https://doi.org/10.1016/j.semcdb.2020.07.002
Fleming AM, Burrows CJ (2021) Oxidative stress-mediated epigenetic regulation by G-quadruplexes. 3:1–16
Fortuny A, Chansard A, Caron P, Chevallier O, Leroy O, Renaud O, Polo SE (2021) Imaging the response to DNA damage in heterochromatin domains reveals core principles of heterochromatin maintenance. Nat Commun 12:1–16. https://doi.org/10.1038/s41467-021-22575-5
Friedson B, Cooper KF (2021) Cdk8 kinase module: A mediator of life and death decisions in times of stress. Microorganisms 9. https://doi.org/10.3390/microorganisms9102152
Gorini F, Scala G, Di Palo G, Dellino GI, Cocozza S, Pelicci PG, Lania L, Majello B, Amente S (2020) The genomic landscape of 8-oxodG reveals enrichment at specific inherently fragile promoters. Nucleic Acids Res 48:4309–4324. https://doi.org/10.1093/NAR/GKAA175
Guha M, Srinivasan S, Ruthel G, Kashina AK, Carstens RP, Mendoza A, Khanna C, Winkle V, Avadhani NG (2014) Mitochondrial retrograde signaling induces epithelial-mesenchymal transition and generates breast cancer stem cells. Oncogene 33:5238–5250. https://doi.org/10.1038/onc.2013.467
Guo YE, Manteiga JC, Henninger JE, Sabari BR, Agnese AD, Shrinivas K, Abraham BJ, Hannett NM, Spille J, Afeyan LK et al (2019) Pol II phosphorylation regulates a switch between transcriptional and splicing condensates. Nature. https://doi.org/10.1038/s41586-019-1464-0
Hanawalt PC (2015) Historical perspective on the DNA damage response. DNA Repair (Amst) 36:2–7. https://doi.org/10.1016/j.dnarep.2015.10.001
Hao W, Wang J, Zhang Y, Wang C, Xia L, Zhang W, Zafar M, Kang JY, Wang R, Ali Bohio A et al (2020) Enzymatically inactive OGG1 binds to DNA and steers base excision repair toward gene transcription. FASEB J 34:7427–7441. https://doi.org/10.1096/fj.201902243R
Jang S, Kumar N, Beckwitt EC, Kong M, Fouquerel E, Rapić-Otrin V, Prasad R, Watkins SC, Khuu C, Majumdar C et al (2019) Damage sensor role of UV-DDB during base excision repair. Nat Struct Mol Biol 26:695–703. https://doi.org/10.1038/s41594-019-0261-7
Jeronimo C, Robert F (2017) The mediator complex: at the Nexus of RNA polymerase II transcription. Trends Cell Biol 27:765–783. https://doi.org/10.1016/j.tcb.2017.07.001
Ježek, J.; Cooper, K.F.; Strich, R. The impact of mitochondrial fission-stimulated ROS production on pro-apoptotic chemotherapy. 2021.
Kagey MH, Newman JJ, Bilodeau S, Zhan Y, Orlando DA, Van Berkum NL, Ebmeier CC, Goossens J, Rahl PB, Levine SS et al (2010) Mediator and cohesin connect gene expression and chromatin architecture. Nature 467:430–435. https://doi.org/10.1038/nature09380
Kim D, Kim KI, Baek SH (2021) Roles of lysine-specific demethylase 1 (LSD1) in homeostasis and diseases. J Biomed Sci 28:1–14. https://doi.org/10.1186/s12929-021-00737-3
Kumar N, Theil AF, Roginskaya V, Ali Y, Calderon M, Watkins SC, Barnes RP, Opresko PL, Pines A, Lans H et al (2022) Global and transcription-coupled repair of 8-oxoG is initiated by nucleotide excision repair proteins. Nat Commun 13:1–16. https://doi.org/10.1038/s41467-022-28642-9
Lebraud E, Pinna G, Siberchicot C, Depagne J, Busso D, Fantini D, Irbah L, Robeska E, Kratassiouk G, Ravanat J-L et al (2020) Chromatin recruitment of OGG1 requires cohesin and mediator and is essential for efficient 8-oxoG removal. Nucleic Acids Res. https://doi.org/10.1093/nar/gkaa611
Lia D, Reyes A, de Melo Campos JTA, Piolot T, Baijer J, Radicella JP, Campalans A (2018) Mitochondrial maintenance under oxidative stress depends on mitochondrially localised α-OGG1. J Cell Sci 131:jcs213538. https://doi.org/10.1242/jcs.213538
Mao P, Brown AJ, Malc EP, Mieczkowski PA, Smerdon MJ, Roberts SA, Wyrick JJ (2017) Genome-wide maps of alkylation damage, repair, and mutagenesis in yeast reveal mechanisms of mutational heterogeneity. Genome Res 27:1674–1684. https://doi.org/10.1101/gr.225771.117
Menoni H, Hoeijmakers JHJ, Vermeulen W (2012) Nucleotide excision repair-initiating proteins bind to oxidative DNA lesions in vivo. J Cell Biol 199:1037–1046. https://doi.org/10.1083/jcb.201205149
Menoni H, Di Mascio P, Cadet J, Dimitrov S, Angelov D (2017) Chromatin associated mechanisms in base excision repair – nucleosome remodeling and DNA transcription, two key players. Free Radic Biol Med 107:159–169. https://doi.org/10.1016/j.freeradbiomed.2016.12.026
Menoni H, Wienholz F, Theil AF, Janssens RC, Lans H, Campalans A, Radicella JP, Marteijn JA, Vermeulen W (2018) The transcription-coupled DNA repair-initiating protein CSB promotes XRCC1 recruitment to oxidative DNA damage. Nucleic Acids Res 46:7747–7756. https://doi.org/10.1093/nar/gky579
Nilsen H, Lindahl T, Verreault A (2002) DNA base excision repair of uracil residues in reconstituted nucleosome core particles. EMBO J 21:5943–5952. https://doi.org/10.1093/emboj/cdf581
Ohno M, Miura T, Furuichi M, Tominaga Y, Tsuchimoto D, Sakumi K, Nakabeppu Y (2006) A genome-wide distribution of 8-oxoguanine correlates with the preferred regions for recombination and single nucleotide polymorphism in the human genome. Genome Res 16:567–575. https://doi.org/10.1101/gr.4769606
Olmon ED, Delaney S (2017) Differential ability of five DNA glycosylases to recognize and repair damage on nucleosomal DNA. ACS Chem Biol 12:692–701. https://doi.org/10.1021/acschembio.6b00921
Osman S, Mohammad E, Lidschreiber M, Stuetzer A, Bazsó FL, Maier KC, Urlaub H, Cramer P (2021) The Cdk8 kinase module regulates interaction of the mediator complex with RNA polymerase II. J Biol Chem 296:100734. https://doi.org/10.1016/j.jbc.2021.100734
Pastukh V, Roberts JT, Clark DW, Bardwell GC, Patel M, Al-Mehdi AB, Borchert GM, Gillespie MN (2015) An oxidative DNA “damage” and repair mechanism localized in the VEGF promoter is important for hypoxia-induced VEGF mRNA expression. Am J Physiol Lung Cell Mol Physiol 309:L1367–L1375. https://doi.org/10.1152/ajplung.00236.2015
Pelish HE, Liau BB, Nitulescu II, Tangpeerachaikul A, Poss ZC, Da Silva DH, Caruso BT, Arefolov A, Fadeyi O, Christie AL, Du K et al (2015) Mediator kinase inhibition further activates super-enhancer associated genes in AML. Nature 526:273–276. https://doi.org/10.1038/nature14904.Mediator
Perillo B, Ombra MN, Bertoni A, Cuozzo C, Sacchetti S, Sasso A, Chiariotti L, Malorni A, Abbondanza C, Avvedimento EV (2008) DNA oxidation as triggered by H3K9me2 demethylation drives estrogen-induced gene expression. Science 202–207
Pezone A, Zuchegna C, Tramontano A, Romano A, Russo G, de Rosa M, Vinciguerra M, Porcellini A, Gottesman ME, Avvedimento EV (2019) RNA stabilizes transcription-dependent chromatin loops induced by nuclear hormones. Sci Rep 9:1–12. https://doi.org/10.1038/s41598-019-40123-6
Pezone A, Taddei ML, Tramontano A, Dolcini J, Boffo FL, De Rosa M, Parri M, Stinziani S, Comito G, Porcellini A et al (2020) Targeted DNA oxidation by LSD1-SMAD2/3 primes TGF-β1/EMT genes for activation or repression. Nucleic Acids Res 48:8943–8958. https://doi.org/10.1093/nar/gkaa599
Poetsch AR (2020) The genomics of oxidative DNA damage, repair, and resulting mutagenesis. Comput Struct Biotechnol J 18:207–219. https://doi.org/10.1016/j.csbj.2019.12.013
Rong Z, Tu P, Xu P, Sun Y, Yu F, Tu N, Guo L, Yang Y (2021) The mitochondrial response to DNA damage. Front Cell Dev Biol 9:1–10
Roychoudhury S, Pramanik S, Harris HL, Tarpley M, Sarkar A, Spagnol G, Sorgen PL, Chowdhury D, Band V, Klinkebiel D et al (2020) Endogenous oxidized DNA bases and APE1 regulate the formation of G-quadruplex structures in the genome. Proc Natl Acad Sci U S A 117. https://doi.org/10.1073/pnas.1912355117
Sabari BR, Dall’Agnese A, Boija A, Klein IA, Coffey EL, Shrinivas K, Abraham BJ, Hannett NM, Zamudio AV, Manteiga JC et al (2018) Coactivator condensation at super-enhancers links phase separation and gene control. Science 361:eaar3958. https://doi.org/10.1126/science.aar3958
Schuster-Böckler B, Lehner B (2012) Chromatin organization is a major influence on regional mutation rates in human cancer cells. Nature 488:504–507. https://doi.org/10.1038/nature11273
Seifermann M, Epe B (2017) Oxidatively generated base modifications in DNA: not only carcinogenic risk factor but also regulatory mark? Free Radic Biol Med. https://doi.org/10.1016/j.freeradbiomed.2016.11.018
Seifermann M, Ulges A, Bopp T, Melcea S, Schäfer A, Oka S, Nakabeppu Y, Klungland A, Niehrs C, Epe B (2017) Role of the DNA repair glycosylase OGG1 in the activation of murine splenocytes. DNA Repair (Amst) 58:13–20. https://doi.org/10.1016/j.dnarep.2017.08.005
Shigdel UK, Ovchinnikov V, Lee SJ, Shih JA, Karplus M, Nam K, Verdine GL (2020) The trajectory of intrahelical lesion recognition and extrusion by the human 8-oxoguanine DNA glycosylase. Nat Commun 11:1–8. https://doi.org/10.1038/s41467-020-18290-2
Soutourina J (2018) Transcription regulation by the Mediator complex. Nat Rev Mol Cell Biol 19:262–274. https://doi.org/10.1038/nrm.2017.115
Spegg V, Altmeyer M (2021) Biomolecular condensates at sites of DNA damage: more than just a phase. DNA Repair (Amst) 106:103179. https://doi.org/10.1016/j.dnarep.2021.103179
Trapp C, Reite K, Klungland A, Epe B (2007) Deficiency of the Cockayne syndrome B (CSB) gene aggravates the genomic instability caused by endogenous oxidative DNA base damage in mice. Oncogene 26:4044–4048. https://doi.org/10.1038/sj.onc.1210167
Tsai KL, Sato S, Tomomori-Sato C, Conaway RC, Conaway JW, Asturias FJ (2013) A conserved Mediator-CDK8 kinase module association regulates Mediator-RNA polymerase II interaction. Nat Struct Mol Biol 20:611–619. https://doi.org/10.1038/nsmb.2549
Visnes T, Cázares-Körner A, Hao W, Wallner O, Masuyer G, Loseva O, Mortusewicz O, Wiita E, Sarno A, Manoilov A et al (2018) Small-molecule inhibitor of OGG1 suppresses proinflammatory gene expression and inflammation. Science 362:834–839. https://doi.org/10.1126/science.aar8048
Wang R, Hao W, Pan L, Boldogh I, Ba X (2018) The roles of base excision repair enzyme OGG1 in gene expression. Cell Mol Life Sci 75:3741–3750
Wang K, Maayah M, Sweasy JB, Alnajjar KS (2021) The role of cysteines in the structure and function of OGG1. J Biol Chem 296:100093. https://doi.org/10.1074/jbc.RA120.016126
Whyte WA, Orlando DA, Hnisz D, Abraham BJ, Lin CY, Kagey MH, Rahl PB, Lee TI, Young RA (2013) Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153:307–319. https://doi.org/10.1016/j.cell.2013.03.035
Wu J, McKeague M, Sturla SJ (2018) Nucleotide-resolution genome-wide mapping of oxidative DNA damage by Click-Code-Seq. J Am Chem Soc 140:9783–9787. https://doi.org/10.1021/jacs.8b03715
Wu W, Hill SE, Nathan WJ, Paiano J, Callen E, Wang D, Shinoda K, van Wietmarschen N, Colón-Mercado JM, Zong D et al (2021) Neuronal enhancers are hotspots for DNA single-strand break repair. Nature. https://doi.org/10.1038/s41586-021-03468-5
Yoshihara M, Jiang L, Akatsuka S, Suyama M, Toyokuni S (2014) Genome-wide profiling of 8-oxoguanine reveals its association with spatial positioning in nucleus. DNA Res 21:603–612. https://doi.org/10.1093/dnares/dsu023
Zuchegna C, Aceto F, Bertoni A, Romano A, Perillo B, Laccetti P, Gottesman ME, Avvedimento EV, Porcellini A (2014) Mechanism of retinoic acid-induced transcription: histone code, DNA oxidation and formation of chromatin loops. Nucleic Acids Res 42:11040–11055. https://doi.org/10.1093/nar/gku823
Acknowledgments
We thank Bernd Epe and Juan Pablo Radicella for the critical reading of the manuscript and for fruitful discussions. We apologize to the authors of many excellent manuscripts that could not be cited during this review because of strong restrictions in the number of references. A.C. lab is funded by Agence National de la Recherche (project ANR TG-TOX), CEA Radiobiology program, Foncer contre le cancer, and Electricité de France.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Section Editor information
Rights and permissions
Copyright information
© 2023 Springer Nature Singapore Pte Ltd.
About this entry
Cite this entry
Di Guilmi, AM., Fonknechten, N., Campalans, A. (2023). OGG1 at the Crossroads Between Repair and Transcriptional Regulation. In: Sugimoto, N. (eds) Handbook of Chemical Biology of Nucleic Acids. Springer, Singapore. https://doi.org/10.1007/978-981-16-1313-5_50-1
Download citation
DOI: https://doi.org/10.1007/978-981-16-1313-5_50-1
Received:
Accepted:
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-16-1313-5
Online ISBN: 978-981-16-1313-5
eBook Packages: Springer Reference Chemistry and Mat. ScienceReference Module Physical and Materials ScienceReference Module Chemistry, Materials and Physics