Abstract
In this chapter, we give a not-so-long and self-contained introduction to computational molecular evolution. In particular, we present the emergence of the use of likelihood-based methods, review the standard DNA substitution models, and introduce how model choice operates. We also present recent developments in inferring absolute divergence times and rates on a phylogeny, before showing how state-of-the-art models take inspiration from diffusion theory to link population genetics, which traditionally focuses at a taxonomic level below that of the species, and molecular evolution. Although this is not a cookbook chapter, we try and point to popular programs and implementations along the way.
You have full access to this open access chapter, Download protocol PDF
Similar content being viewed by others
Key words
1 Introduction
Many books [1,2,3,4,5,6,7] and review papers [8,9,10] have been published in recent years on the topic of computational molecular evolution, so that updating our previous primer on the very same topic [11] may seem redundant. However, the field is continuously undergoing changes, as both models and algorithms become even more sophisticated, efficient, robust, and accurate. This increase in refinement has not been motivated by a desire to complicate existing models, but rather to make an old wish come true: that of having integrated methods that can take unaligned sequences as an input, and simultaneously output the alignment, the tree, and other estimates of interest, in a sound statistical framework justified by sound principles: those of population genetics.
The aim of this chapter is still to provide readers with the essentials of computational molecular evolution, offering a brief overview of recent progress, both in terms of modeling and algorithm development. Some of the details will be left out as they are dealt with by others in this volume. Likewise, the analysis of genomic-scale data is briefly touched upon, but the details are left to other chapters.
2 Parsimony and Likelihood
2.1 A Brief Overview of Parsimony
The simplest phylogenetic question pertains to the reconstruction of a rooted tree with three sequences (Fig. 1). The sequences can be made of DNA, RNA, amino acids, or codons, but for the sake of simplicity we focus on DNA throughout this chapter. In the basic example below, based on [12], DNA sequences are assumed to have been sampled from three different species that diverged a “long time ago.” In this context, we assume that the data or gene sequences have been aligned (see Subheading 6), and that the DNA alignment is:
s 1 | ATGACCCCAATACGCAAAACTAACCCCCTAATAAAATTAATTAACCACTCCTTC |
s 2 | ATGACCCCAATACGGAAAACTAACCCCCAAATAAAATTAATTAACCACTCATTC |
s 3 | ATGACGCCAATACGCAAAACTAACCGCCTAATAAAATTAATTTACCACTCATTC |
The objective is to estimate which of the three fully resolved topologies in Fig. 1 is supported by the data. In order to go further, we recode the data in terms of site patterns, which correspond to the patterns observed in each column of our alignment. This recoding implies that columns, or sites, in our alignment evolve according to an identically and independently distributed (iid) process. With this in mind, our alignment can be recoded in the following manner. When all the characters (nucleotides) in a column are identical, the same letter is assigned to each character, for example, x, irrespective of the actual character state. When a substitution occurs in one of the three sequences, we have three corresponding site patterns: xxy, xyx, and yxx, where the order within each site pattern respects the order of the sequences in the alignment, s 1 s 2 s 3.
s 1 | xxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxxyxxx |
s 2 | xxxxxxxxxxxxxxyxxxxxxxxxxxxxyxxxxxxxxxxxxxxxxxxxxxxxxx |
s 3 | xxxxxyxxxxxxxxxxxxxxxxxxxyxxxxxxxxxxxxxxxxyxxxxxxxxxxx |
The first informative site pattern, xxy, implies that at this particular site, sequences s 1 and s 2 are more similar than any of these to s 3, so that this site pattern supports topology T 1, which groups sequences s 1 and s 2 together (Fig. 1). The most intuitive idea, called the winning-site strategy, is that the topology supported by the data corresponds to the fully resolved topology that has the largest number of site patterns in its favor. In the example shown above, topology T 1 is supported by three columns (with site pattern xxy), topology T 2 by two columns (xyx), and T 3 by one column (yxx; see Table 1). This is the intuition behind parsimony, which minimizes the amount of change along a topology. Strictly speaking, unordered parsimony cannot distinguish these three trees as they all require at least one single change. Yet, it can be argued that if tree T 1 is the true tree, site pattern xxy is more likely than any other patterns as xxy requires at least one change along a long branch (the one leading to sequence s 3) while both xyx and yxx require a change along a short branch (see p. 28 sqq. in [13]; [12]).
A number of methodological variations exist. A very condensed overview can be found in the books by Durbin [14] or, with more details, Felsenstein [15]. Most computer programs that implement substitution models where sites are iid condense the alignment as an array of site patterns; some, like PAML [16], even output these site patterns.
Note that in obtaining this topology estimate, most of the site columns were discarded from our alignment (all the xxx site patterns, representing 89% of the site in our example above). Most of our data were phylogenetically uninformative (for parsimony). We also failed to take evolutionary time into account, or any process of basic molecular biology, such as the observation that transitions (substitution of a purine [A or G] by a purine, or a pyrimidine by a pyrimidine) are more frequent than transversions (substitution between a purine and a pyrimidine).
2.2 Assessing the Reliability of an Estimate: The Bootstrap
As with any statistical exercise estimating a quantity of interest, we would like to have a confidence interval, taken at a particular level, so that we can gauge the reliability of our estimate. A standard approach to derive confidence intervals is the bootstrap [17], a computational technique that resamples data points with replacement to simulate the distribution of any test statistic under the null hypothesis that is tested. The bootstrap, particularly useful in complicated nonparametric problems where no asymptotic results can be obtained [18], was adapted by Felsenstein to the nonstandard phylogenetic problem [19]. Indeed, the problem is nonstandard in that the object for which we wish to assess accuracy is not a real-valued parameter, but a graph.
The basic idea, clearly explained in [20], consists in resampling columns of the alignment, with replacement, to construct a “synthetic” alignment of the same size as the original alignment. This synthetic or bootstrap replicate is then subjected to the same tree-reconstruction algorithm used on the original data (Fig. 2). This exercise is repeated a large number of times (e.g., × 106), and the proportion of each original bipartition (internal node) in the set of bootstrapped trees is recorded. In Fig. 2, for instance, the bipartition s 1 s 2|s 3 is found in two bootstrap trees out of three, so the bootstrap support for this node is 66.7%. In this simple case with three sequences, the bootstrap support for topology T 1 is also 66.7%. This bootstrap proportion for topologies (or for trees when branch lengths are taken into account, in a maximum likelihood context, for instance—see below) can be computed very quickly by bootstrapping the sitewise log-likelihood values, instead of the columns of the alignment; this bootstrap is called RELL, for “resampling estimated log-likelihood” [21].
However, this approach is no longer used or cited extensively since 2008 (source: ISI Thompson). One alternative that has gained momentum is the one based on the approximated likelihood ratio test (aLRT) [22], implemented, for instance, in phyml [23, 24]. Instead of resampling any quantity (sites or sitewise log-likelihood values), the aLRT tests the null hypothesis that an interior branch length is zero. In spite of being slightly conservative in simulations, the approach is extremely fast and hence highly practical [22].
The meaning of the bootstrap has been a matter of debate for years. As noted before [8] (see also [22]), the bootstrap proportion P can be seen as assessing the correctness of an internal node, and failing to do so [25], or 1 − P can be interpreted as a conservative probability of falsely supporting monophyly [26]. Since bootstrap proportions are either too liberal or too conservative depending on the actual interpretation of P [27], it is difficult to adjust the threshold below which monophyly can be confidently ruled out [28]. Alternatively, an intuitive geometric argument was proposed to explain the conservativeness of bootstrap probabilities [18] and was further developed into the approximately unbiased or AU test, implemented in CONSEL [29]. In spite of these difficulties, the bootstrap is still widely used—and mandatory in all publications featuring a phylogeny—to assess the confidence one can have in the tree estimated from the data under a particular scheme or model (see Subheading 2.9.3 below). Lastly, note that bootstrap support has often been abused [30], as a high value does not necessarily indicate high phylogenetic signal, and can be the result of systematic biases [31] due to the use of the wrong model of evolution, for instance, as detailed below.
2.3 Parsimony and LBA
Now that we have a means of evaluating the support for the different topologies, we can test some of the conditions under which parsimony estimates the correct tree topology. Ideally, a good method should return the correct answer with a probability of one when the number of sites increases to infinity. This desirable statistical property is called consistency. One serious criticism of parsimony is its sensitivity to long branch attraction, or LBA, even in the presence of an infinite amount of data (infinite alignment length) [31]. In other words, parsimony is not statistically consistent.
Different types of model misspecification can lead to LBA, and new ones are continually identified. The topology originally used to demonstrate the artifact is represented in Fig. 3, where two long branches are separated by a shorter one. Felsenstein demonstrated that, under a simple evolutionary process, the artifact or LBA tree is reconstructed. Note that parsimony is not the only phylogenetic method affected by LBA, but because it posits a very simple model of evolution [32,33,34], parsimony is particularly sensitive to the artifact. In spite of this, one particular journal chose to enforce the use of parsimony, stating that authors should estimate their phylogenies by parsimony but also that, if estimated by some other method, they would need to defend their position “on philosophical grounds” [35]; there is of course no valid scientific justification for taking such a step—derided in the “Twittersphere” as “#parsimonygate.”
The long branch attraction artifact. The true tree topology has two long branches separated by a short one. The tree reconstructed under a simple model of evolution (a) is the artifact or LBA tree on the left. The tree reconstructed under the correct model of evolution (b) is the correct tree, on the right
The LBA artifact has been shown to plague the analysis of numerous data sets, and a number of empirical approaches have been used to detect the artifact [36, 37]. Most recent papers based on multigene analyses (e.g., [38, 39]) now examine carefully the effect of across-site and across-lineage rate variation (in addition to the use of heterogeneous models). For both sites and lineages, the procedure is the same and consists in successively removing either the sites that evolve the fastest or the taxa that show the longest root-to-tip branch lengths.
2.4 Origin of the Problem
By definition, parsimony minimizes the number of changes along each branch of the tree. When there is only a small number of changes per branch, the method is expected to be accurate. However, when sequences are quite divergent, the parsimony assumption leads to underestimating the actual number of changes (Fig. 4; see also [40]).
Consequently, we would like a tree-reconstruction method that accounts for multiple substitutions. We would also like a method that (1) takes into account less parsimonious as well as most parsimonious state reconstructions (intervals, tests), (2) weights changes differently if they occur on branches of different length (evolutionary time), and (3) weights different kinds of events (transitions, transversions) differently (biological realism). Likelihood methods include such considerations explicitly, as they require modeling the substitution process itself.
2.5 Modeling Molecular Evolution
The basic model of DNA substitution (Fig. 5) is defined on the DNA state space, made of the four nucleotides thymine (T), cytosine (C), adenine (A), and guanine (G). Note that T and C are pyrimidines (biochemically, six-membered rings), while A and G are purines (fused five- and six-membered heterocyclic compounds). Depending on these two biochemical categories, two different types of substitutions can happen: transitions within a category, and transversions between categories. Their respective rates are denoted α and β in Fig. 5.
The process we want to model should describe the substitution process of the different nucleotides of a DNA sequence. Again, we will make the simplifying assumption that sites evolve under a time-homogeneous Markov process and are iid, as above. We can therefore concentrate on one single site for now (e.g., [41]).
At a particular site, we want to describe the change in nucleotide frequency after a short amount of time dt, during which the nucleotide frequency of A, for instance, after dt will change from f A(t) to f A(t + dt). According to Fig. 5, f A(t + dt) will be equal to what we had at time t, f A(t), minus the quantity of A that “disappeared” by mutation during dt, plus the quantity of A that “appeared” by mutation during dt. Denoting the mutation rate as μ, the quantity of A that “disappeared” by mutation during dt is simply f A(t)μ A dt. These mutations away from A generated quantities of T, C, and G, in which we are not interested at the moment since we only want to know what happens to A. There are three different ways to generate A: from either T, C, or G (Fig. 5). Coming from T, mutation will generate f T(t)μ T→A dt of A during dt. Similar expressions exist for C and for G, so that in total, over the three non-A nucleotides, mutation will generate ∑i≠A f i(t)μ i→A dt. Mathematically, we can express these ideas as:
Equation 1 describes the change of frequency of A during a short time interval dt. Similar equations can be written for T, C, and G, so that we actually have a system of four equations describing the change in nucleotide frequencies over a short time interval dt:
which, in matrix notation, can simply be rewritten as:
with an obvious notation for F, while the instantaneous rate matrix Q is
In all the following matrices, we will use the same order for nucleotide: T, C, A, and G, which follows the order in which codon tables are usually written. Recall that μ ij is the mutation rate from nucleotide i to nucleotide j. Note also that the sum of each row is 0.
Let us rearrange the matrix notation from Eq. 3 as:
and take the variation limit when dt → 0:
which is a first-order differential equation that can be integrated as:
Very often, this last equation 7 is written as F(t) = P(t)F(0), where F(0) is conveniently taken to be the identity matrix and P(t) = {P i,j(t)} = e Qt is the matrix of probabilities of going from state i to j during a finite time duration t. Note that the right-hand side of this equation is a matrix exponentiation, which is not the same as the exponential of all the elements (row and columns) of that matrix. The computation of the term e Qt demands that a spectral decomposition of the matrix Q be realized. This means finding a diagonal matrix D of eigenvalues and a matrix M of (right) eigenvectors so that:
The exponential of the diagonal matrix D is simply the exponential of the diagonal terms.
Except in the simplest models of evolution, finding analytical solutions for the eigenvalues and associated eigenvectors can be tedious. As a result, numerical procedures are employed to solve Eq. 8. Alternatively, a Taylor expansion can be used to approximate P(t).
If all entries in Q are positive, any state or nucleotide can be reached from any other in a finite number of steps (all states “communicate”) and the base frequencies have a stationary distribution π = (π T, π C, π A, π G). This is the steady state reached after an “infinite” amount of time, or long enough for the Markov process to forget its initial state, starting from “random” base frequencies.
2.6 Computation on a Tree
Now that we know how to determine the rate of change of nucleotide frequencies during a time interval dt, we can compute the probability of a particular nucleotide change on a tree. The simplest case, though somewhat artificial with only two sequences, is depicted in Fig. 6.
We are looking at a particular nucleotide position, denoted j, for two aligned sequences. The observed nucleotides at this position are T in sequence 1, and C in sequence 2. The branch separating T from C has a total length of t 0 + t 1. For the sake of convenience, we set an arbitrary root along this path. The likelihood at site j is then given by the probability of going from the fictive root i to T in t 0, and from i to C in t 1. Any of the four nucleotides can be present at the fictive root. As we do not know which one was there, we sum these probabilities over all possible state, weighted by their prior probabilities, the equilibrium frequencies π i. In all, we have the likelihood ℓ j at site j:
which is equivalent to the Chapman–Kolmogorov equation [42]. As all the sites are assumed to be iid, the likelihood of an alignment is the product of the site likelihoods in Eq. 9. Because all these sitewise probabilities can be small, and that the product of small numbers can become smaller than what a computer can represent in memory (underflow), all computations are done on a logarithmic scale and may include some form of rescaling [43].
Note that this example is somewhat artificial: with only two sequences, we can compute the likelihood directly with π T P T,C(t 0 + t 1) = π C P C,T(t 0 + t 1); the full summation over unknown states as in Eq. 9 is required with three sequences or more. When analyzing a multiple-sequence alignment of S sequences, there will be many nodes in the tree for which the character state is unknown, which means that the summation required will involve many terms. Specifically, the sum will be over 4S−3 terms. Fortunately, terms can be factored out of the summation, and a dynamic programing algorithm with a complexity of the order of \( \mathcal{O}\left({4}^2S\right) \), called the pruning algorithm [44], can be used (see [15] for details).
2.7 Substitution Models and Instantaneous Rate Matrices Q
Now that we have almost all the elements to compute the likelihood of a set of parameters, including the tree (i.e., the set of branch lengths and the tree topology; see Subheading 2.10), the only missing element required to compute the likelihood at each site, as in Eq. 9, for instance, is the specification of the instantaneous rate matrix Q as in Eq. 4. Remember that the μ i,j represent mutation rates from state (nucleotide) i to j. This matrix is generally rewritten as:
so that each entry r ij is a rate of change from nucleotide i to nucleotide j. The diagonal entries are left out, indicated by a “−,” and are in fact calculated as the negative sum of the off-diagonal entries (as rows sum to 0).
The simplest specification of Q would be that all rates of change are identical, so that Q becomes (leaving out the mutation rate μ and indexing the matrix to indicate the difference):
which is the model proposed by Jukes and Cantor [45] and often noted “JC” or “JC69”. Under the specification of Eq. 11, this model has no free parameter. The process is generally scaled such that the unit of branch lengths can be interpreted as an expected number of substitutions per site.
Of course, this model is extremely simplistic and neglects a fair amount of basic molecular biology. In particular, it overlooks two observations. First, base frequencies are not all equal in actual DNA sequences, but are rather skewed, and second, transitions are more frequent than transversions (see Subheading 2.5).
The way to account for this first “biological realism” is as follows. If DNA sequences were made exclusively of As, for instance, that would mean that all mutations are towards the observed base, in this case A, whose equilibrium or stationary frequency is π A. The same reasoning can be used for arbitrary equilibrium frequencies π, so that all relative rates of change in Q become proportional to the vector of equilibrium frequency π of the target nucleotide. In other words, the instantaneous rate matrix Q becomes:
again with the requirement that rows sum to 0. This matrix represents the Felsenstein or F81 model [44]. This model has four parameters (the four base frequencies), but since base frequencies sum to 1, we only have three free parameters.
The second “biological realism,” accounting for the different rates of transversions and transitions, can be described by saying that transitions occur κ times faster than transversions. From Fig. 5, recall that transitions are mutations from T to C (and vice versa) and from A to G (and vice versa). This translates into:
This model is called the Kimura two-parameter model or K80 (or K2P) [46]. The model is alternatively described with the two rates α and β (see Fig. 5). In the “κ version” of the model as in Eq. 13, there is only one free parameter.
Of course it is possible to account for both kinds of “biological realism,” unequal equilibrium base frequencies and transition bias, all in the same model, whose generator Q becomes:
which corresponds to the Hasegawa–Kishino–Yano or HKY (or HKY85) model [47]. This model has four free parameters: κ and three base frequencies.
The level of “sophistication” goes “up to” the general time-reversible model [48], denoted GTR or REV, which has for generator:
The number of free parameters is now eight (three base frequencies plus five nucleotide propensities). The name is derived from the time-reversibility constraint, which implies that the likelihood is independent of the actual orientation of time.
In fact, there exist only a few “named” additional substitution models [15], most of which are time-reversible models, while a total of 203 models can be derived from GTR [49]. We have focused solely on DNA models in this chapter, but the problem is similar with amino acid or codon models, except that the number of parameters increases quickly. We have also limited ourselves to time-reversible time-homogeneous models, but irreversible non-homogeneous models were developed some time ago [50] and are used, for instance, to root phylogenies [51] or to help alleviate the effects of LBA [39].
2.8 Some Computational Aspects
2.8.1 Optimization of the Likelihood Function
For a given substitution model, how should parameters be estimated, given the (potentially) high dimensionality of the model? Analytical solutions consist in determining when the first derivative of the likelihood function is equal to zero (with a change of sign in the second derivative). However, finding the root of the likelihood function analytically is only possible in the simple case of three sequences of binary characters under the assumption of the molecular clock (see Subheading 3.1) [12]. As a result, numerical solutions must be found to maximize the likelihood function.
A number of ideas have been combined to search efficiently for the parameter values that maximize the likelihood function. Most programs will start from a random starting point, for example, \( \left({\theta}_1^{(0)},{\theta}_2^{(0)}\right) \), denoted by an x in Fig. 7, where we limit ourselves to a two-parameter example. The optimization procedure can follow one of the two strategies. In the first one, parameters are optimized one at a time. In Fig. 7a, parameter θ 1 is first optimized to maximize the likelihood function with a line search, which defines a direction along which the other parameter (θ 2) or parameters in the multidimensional case are kept constant. Once \( {\theta}_1^{(1)} \) is found, a new direction is defined to optimize θ 2, and so on so forth until convergence to the maximum of the likelihood function. As shown in Fig. 7a, many iterations can be required, in particular when the parameters θ 1 and θ 2 are correlated. The alternative to optimizing one parameter at a time is to optimize all parameters simultaneously. In this case (Fig. 7b), an initial direction is defined at \( \left({\theta}_1^{(0)},{\theta}_2^{(0)}\right) \) such that the slope at this point is maximized. The process is repeated until convergence. More technical details can be found in [52]. The simultaneous optimization procedure generally requires fewer steps than optimizing parameters one at a time, but not always. Since the computation of the likelihood function is the most expensive computation of these algorithms, the simultaneous optimization is much more efficient, at least in our toy example.
Two optimization strategies. The likelihood surface of a function with two parameters θ 1 and θ 2 (e.g., two branch lengths) is depicted as a contour plot, whose highest peak is at the + sign. (a) Optimization one parameter at a time. (b) Optimization of all parameters simultaneously. See text for details
How general is this result? Simultaneously optimizing parameters of the substitution model, while optimizing branch lengths one at a time, was shown to be more effective on large data sets [43], potentially because of the correlation that exists between some of the parameters entering the Q matrix (see Subheading 2.7).
2.8.2 Convergence
Convergence is usually reached either when the increment in the log-likelihood score becomes smaller than an ε value, usually set to a small number such as 10−6 (but yet a number larger than the machine ε: the smallest number that a given computer can represent), or when the log-likelihood score has not changed after a predetermined number of iterations. However, none of these stopping rules guarantees that the global maximum of the likelihood function has been found. Therefore, it is generally recommended to run the optimization procedure at least twice, starting from different initial values of the model parameters, and to check that the likelihood score after optimization is the same across the different runs (Fig. 8). If this is not the case, additional runs may be required, and the one with the largest likelihood is chosen for inference (e.g., [53]).
Likelihood surfaces behaving badly. Schematic of the probability surface of the function p(X|θ) is plotted as a function of θ. Most line search strategies will converge (CV) to the MLE when the initial value is in the “CV” interval, and fail when it is in the “no CV” interval. Adapted with permission from [54]
In many instances though, different substitution models will give different tree topologies, and therefore different biological conclusions. One difficulty is therefore to know which model should be used to analyze a particular data set.
2.9 Selection of the Appropriate Substitution Model
One important issue in model selection is about the trade-off between bias and variance [55]: a simple model will fail to capture all the sophistication of the actual substitution process, and will therefore be highly biased even if all the parameters can be estimated with tight precision (little variance). Alternatively, a highly parameterized model will “spread” the information available from the data over a large number of parameters, hereby making their estimation difficult (flat likelihood surface; see Subheading 2.8.1), with a large variance, in spite of perhaps being a more realistic model with less bias. The objective of most model selection procedure is therefore to find not the best model in terms of likelihood score, but the most appropriate model, the one that strikes the right balance between bias and variance in terms of number of parameters. However, we argue that optimizing for this bias–variance trade-off works only for statistical procedures, be they, for instance, frequentist (LRT, likelihood ratio test) or Bayesian (BF, Bayes factor). On the other hand, information-theoretic criteria (e.g., AIC, Akaike information criterion) aim at selecting the model that is approximately closest to the “true” biological process.
The bias–variance trade-off mainly concerns the comparison of models that are based on the same underlying rationale, for instance, choosing among the 203 models that can be derived from GTR. We may also be interested in comparing models that are based on very different rationales. The likelihood ratio test is suited for assessing the bias–variance trade-off, while Bayesian and information-theoretic approaches, as well as cross-validation (CV), can be used for more general model comparisons. Here we review four approaches to model selection: LRT, BF, AIC, and CV.
2.9.1 The Likelihood Ratio Test
The substitution models presented above have one key property: it is possible to reduce the most sophisticated time-reversible named model (GTR+Γ+I) to any simpler model by imposing some constraints on parameters. As a result, the models are said to be nested, and statistical theory (the Neyman–Pearson lemma) tells us that there is an optimal (most powerful) way of comparing two nested models (a simple null vs. a simple alternative hypothesis) based on the likelihood ratio test or LRT.
The test statistic of the LRT is twice the log-likelihood difference between the most sophisticated model (which by definition is always the one with the highest likelihood—if this is not the case, there is a convergence issue; see Subheading 2.8.1) and the simpler model. This test statistic follows asymptotically a χ 2 distribution (under certain regularity conditions), and the degree of freedom of the test is equal to the difference in the number of free parameters between the two models.
The null hypothesis is that the two competing models explain the data equally well. The alternative is that the most sophisticated model explains the data better than the simpler model. If the null hypothesis cannot be rejected at a certain level (type-I error rate), then, based on the argument developed above, the simpler model should be used to analyze the data. Otherwise, if the null hypothesis can be rejected, the more sophisticated model should be used to analyze the data. Note that a test never leads to accepting a null hypothesis; the only outcomes of a test are either reject or fail to reject a null hypothesis.
Intuitively, we can see the null hypothesis H 0 as stating that a certain parameter θ is equal to θ 0. The maximum likelihood estimate (MLE) is at \( \widehat{\theta} \), which is our alternative hypothesis H 1, left unspecified. We note the log-likelihood as \( \ln p\left(X|\theta \right)=\ell \left(\theta \right) \), where X represents the data. Under H 0, we have θ = θ 0, while under H 1 we have \( \theta =\widehat{\theta} \). The log-likelihood ratio is therefore \( \ln LR=\ell \left(\widehat{\theta}\right)-\ell \left({\theta}_0\right) \). Under the null H 0, \( \ell \left(\widehat{\theta}\right)=0 \) (by definition). The log-likelihood ratio then reduces to \( \ln LR=-\ell \left({\theta}_0\right) \). We can then take the Taylor expansion of the log-likelihood function ℓ around \( \widehat{\theta} \), which gives us \( \ell \approx \frac{1}{2}{\left(\widehat{\theta}-{\theta}_0\right)}^2\frac{d^2\ell }{d{\theta}^2} \) (recall that \( \ell \left(\widehat{\theta}\right)=0 \), so that the first terms of the series “disappear”). Therefore, log-likelihood ratio can be approximated by \( -\frac{1}{2}{\left(\widehat{\theta}-{\theta}_0\right)}^2\frac{d^2\ell }{d{\theta}^2} \). Recall that Fisher’s information is negative reciprocal of the second derivative of the likelihood function, so that:
which follows asymptotically half a χ 2 distribution. Hence the usual approximation:
with k being the difference in the number of free parameters between the two models 0 and 1. The important points in this intuitive outline of the proof are that (1) the two hypotheses need to be nested and (2) taking the Taylor expansion around \( \widehat{\theta} \) requires that the likelihood function be continuous at that point, which implies that ℓ is differentiable left and right of \( \widehat{\theta} \). Therefore, testing points at the boundary of the parameter space cannot be done by approximating the distribution of the test statistic of the LRT by a regular χ 2 distribution, as noted many times in molecular evolution [56,57,58,59,60,61,62,63,64]. A solution still involves the LRT, but the asymptotic distribution becomes a mixture of χ 2 distributions [65].
An approach that has become popular under the widespread adoption of computer programs such as ModelTest [66] and jModelTest [67] is the hierarchical LRT (hLRT). This hierarchy goes from the simplest model (JC) to the set of most complex models (+Γ+I), traversing a tree of models. The issue is that there is more than one way to traverse this tree of models, and that depending on which way is adopted, the procedure may end up selecting different models [68, 69].
2.9.2 Information-Theoretic Approaches
Information theory provides us with a number of solutions to circumvent the three limitations of the LRT (nestedness, continuity, and dependency on the order in which models are compared).
The core of the information-based approach is the Kullback–Leibler (KL) distance, or information [70], which measures the distance between an approximating model g and a “true” model f [55]. This distance is computed as:
where θ is a vector of parameters entering the approximating model g and x represents the data. Note that this distance is not symmetric, as typically d KL(f, g) ≠ d KL(g, f), and that the “true” model f is unknown. The idea is to rewrite d KL(f, g) in a slightly different form, to make it clear that Eq. 18 is actually a difference between two expectations, both taken with respect to the unknown “truth” f:
Equation 19 therefore measures the loss of information incurred by fitting g when the data x actually come from f. As f is unknown, d KL(f, g) cannot be computed as such.
Two points are key to deriving the criterion proposed by Akaike (see [55]). First, we usually want to compare at least two approximating models, g 0 and g 1. We can then measure which one is closest to the “true” process f by taking the difference between their respective Kullback–Leibler distances. In the process, the direct reference to the “true” process cancels out. As a result, the “best” model among g 0 and g 1 is the one that is closest to the “true” process f: it is the model that minimizes the distance to f. By setting model parameters to their MLEs, we now deal with estimated distances, but these are still with respect to the unknown f.
Second, in the context of a frequentist approach, we would repeat the experiment of sampling data an infinite number of times. We would then compute the expected estimated KL distance, so that model selection can be done on the sole estimated log-likelihood value. Akaike, however, showed that this latter approximation is biased, and must be adjusted by a term that is approximately equal to the number of parameters K entering model g (see [55]). For “historical reasons” (similarity with asymptotic theory with the normal distribution), the selection criterion is multiplied by 2 to give the well-known definition of the Akaike information criterion or AIC:
Unlike the case of the hLRT, where we were selecting the “most appropriate model” (with respect to the bias–variance trade-off), in the case of AIC we can select the best model. This best model is the one that is closest to the “true” unknown model (f), with the smallest relative estimated expected KL distance. The best AIC model therefore minimizes the criterion in Eq. 20.
A small-sample second-order version of AIC exists, where the penalty for extra parameters (2K in Eq. 20) is slightly modified to account for the trade-off between information content in the data and K (see [55]). In our experience, we find it advisable to use this small-sample correction irrespective of the actual size of the data, since this correction vanishes in large and informative samples, but corrects for proper model ranking when K becomes very large compared to the amount of information (e.g., in phylogenomics where models are partitioned with respect to hundreds of genes).
The AIC has been shown to tend to favor parameter-rich models [71,72,73,74,75], which has motivated the use and development of alternative approaches in computational molecular evolution. These include, the Bayesian information criterion [76], and the decision theory or DT approach, which is based on ΔAIC weighted by squared branch length differences [71]. Most of these approaches, including the hLRT, have recently been compared in a simulation study that suggests, in agreement with empirical studies [72, 77], that both BIC and DT have the highest accuracy and precision [75].
One particular drawback of these information-theoretic approaches is that they require that every single model of evolution, or at least the most “popular” models (the few named ones), be evaluated. This step can be time-consuming, especially if a full maximum likelihood optimization is performed under each model. A first set of heuristics consists in fixing the tree topology to a tree estimated with a quick distance-based method such as BioNJ [78], and then estimating just the branch lengths and the parameters of the substitution model, as implemented in jModelTest [67]. As the optimizations are independent of each other under each substitution model, these computations are typically forked to multiple cores or processors [79]. Further heuristics exist to avoid all these independent optimizations [79], as implemented in SMS (Smart Model Selection in PhyML), which is reported to be cutting runtimes in half without forfeiting accuracy [80].
Note finally that all these approaches are not limited to selecting the most appropriate or the best model of evolution. Disregarding the hLRT, which requires that models be nested (to be able to use the χ 2 approximation; otherwise, see [65]), AIC, BIC, etc. allow us to compare non-nested models and, in particular, phylogenetic trees (branch lengths plus topology).
2.9.3 The Bayesian Approach
The Bayesian framework has permitted the development of two main approaches, which are actually two sides of the same coin: one based on finding the model that is the most probable a posteriori, and one based on ranking models and estimating a quantity called the Bayes factor.
In a nutshell, the frequentist approaches developed in the previous sections are based on the likelihood, which is the probability of the data, given the parameters: p(X|θ). However, this approach may not be the most intuitive, since most practitioners are not interested in knowing the conditional probability of their data, as the data were collected to learn more about the processes that generated them. It can therefore be argued that the Bayesian approach, which considers the probability of the parameters given the data or p(θ|X), is more intuitive than the frequentist approach. Unlike likelihood, which relies on the function p(X|θ) and permits point estimation, Bayesian inference is based on the posterior distribution p(θ|X). This distribution is often summarized by a centrality measure such as its mode, mean, or median. Measures of uncertainty are based on credibility intervals, the Bayesian equivalent of confidence intervals. Typically, credibility intervals are taken at the 95% cutoff and are called highest posterior densities (HPDs).
The connection between posterior probability and likelihood is made with Bayes’ inversion formula, also called Bayes’ theorem, by means of a quantity called the prior distribution p(θ):
The prior represents what we think about the process that generated the data, before analyzing the data, and is at the origin of all controversies surrounding Bayesian inference. In practice, priors are more typically chosen based on statistical convenience, and often have nothing to do with our genuine state of knowledge about parameters before observing the available data. We will see in Subheading 3.1 that priors can be used to distinguish between parameters that are confounded in a maximum likelihood analysis (model), so that we argue that the frequentist vs. Bayesian controversy is sterile, and we advocate a more pragmatic approach, that often results in the mixing of both approaches (in their concepts and techniques) [81, 82].
All models have parameters. Subheading 2.7 treats substitution models, which can have eight free parameters in the case of GTR + Γ. Most people are not really interested in these parameters θ or in their estimates \( \widehat{\theta} \), but have to use them in order to estimate a phylogenetic tree τ. These parameters θ are called nuisance parameters because they enter the model but are not the focus of inference. The likelihood solution consists in setting these parameters to their MLE, ignoring the uncertainty with which they can be estimated, while the Bayesian approach will integrate them out, directly accounting for their uncertainty:
One difficulty in Bayesian inference is about the denominator in Eq. 21, as this denominator often has no analytical solution. In spite of being a normalizing constant, p(X) requires integrating out nuisance parameters by means of prior distributions as in Eq. 22. Thus, it is easy to see from Eq. 21 that the posterior distribution of the variable of interest (e.g., τ) can quickly become complicated:
where τ and θ are assumed to be independent and the discrete sum is taken over the set T of all possible topologies (see Subheading 2.10.1). However, the ratio of posteriors evaluated at two different points will simplify: as the denominator in Eq. 23 is a constant, it will cancel out from the ratio. This simple observation is at the origin of an integration technique for approximating the posterior distribution in Eq. 23: Markov chain Monte Carlo (MCMC) samplers. A very clear introduction can be found in [83].
Building on this, two approaches can be formulated to compare models in a Bayesian framework. The first is to treat the model as a “random variable,” and compute its posterior probability. The best model is then the one that has the highest posterior probability. This approach is typically implemented in a reversible-jump MCMC (or rjMCMC) sampler (e.g., see [49]).
The alternative is to use the Bayesian equivalent of the LRT, the Bayes factor. Rather than comparing two likelihoods, the Bayes factor compares the probability of the data under two models, M 0 and M 1:
More specifically, BF0,1 evaluates the weight of evidence in favor of model M 0 against model M 1, with BF0,1 > 1 considered as evidence in favor of M 0. Just as in a frequentist context, where a null hypothesis is significantly rejected at a certain threshold, 5%, 1%, or less depending on different costs or error types, Bayes factors can be evaluated on a specific scale [84]. However, because this scale is just as ad hoc as in a frequentist setting, it might be preferable to use the probability of the data under a particular model p(X|M i) as a means of ranking models M i.
The quantity p(X|M 0), which is the denominator in Eq. 23 (where we did not include the dependence on the model in the notation), is called the marginal likelihood. Note that it is also an expectation with respect to a prior probability distribution:
A number of approximations to evaluate Eq. 25 exist and are reviewed in [85] (see also [86, 87]). The simplest one is based on the harmonic mean of the likelihood sampled from the posterior distribution [88], also known as the harmonic mean estimator (HME). The way this estimator is derived demands to understand how integrals can be approximated. Briefly, to compute \( I=\int g\left(\theta \right)\kern0.3em p\left(\theta \right)\kern0.3em d\theta \), generate a sample from a distribution p ⋆(θ) and calculate the simulation-consistent estimator \( I=\sum {w}_i\kern0.3em g\left(\theta \right)/ \sum {w}_i \) where w i is the importance function p(θ)∕p ⋆(θ). Take g = p(X|θ) and p ⋆(θ) = p(X|θ) p(θ)∕p(X), then \( \widehat{I}=\widehat{p}\left(X|{M}_0\right)={\lim}_{N\to \infty }{\left(\frac{1}{N}\sum \frac{1}{p\left(X|{\theta}_i\right)}\right)}^{-1} \) with θ ∼ p(θ|X) (see supplementary information in [89]). As a result, a very simple way to estimate the marginal likelihood and Bayes factors is to take the output of an MCMC sampler and compute the harmonic mean of the likelihood values (not the log-likelihood values) sampled from the posterior distribution.
Because of its simplicity, this estimator is now implemented in most popular programs such as MrBayes [90] or BEAST [91]. However, it might be considered as the worst estimator possible, because its results are unstable [88, 92] and biased towards the selection of parameter-rich models [86]. An alternative and reliable estimator, based on thermodynamic integration (TI; [86]—also known as path sampling; [93, 94]), is much more demanding in terms of computation. Indeed, it requires running MCMC samplers morphing one model into the other (and vice versa), which can increase computation time by up to an order of magnitude [86]. Improvements of the TI estimator are however available. The stepping-stone (SS) approach builds on importance sampling and TI to speed up the computation while maintaining the accuracy of the standard TI estimator [87, 95].
Moving away from the estimation of marginal likelihoods, an analogue of AIC that can be obtained through the output of an MCMC sampler (AICM) was proposed [96]. In essence, it relies on the asymptotic convergence of the posterior distribution of the log-likelihood on a gamma distribution [97]. As such, it becomes possible to estimate the effective number of parameters as twice the sample variance of posterior distribution of the log-likelihood, which itself can be estimated by a resampling procedure [96]. This gives a very elegant means of estimating AIC, from the posterior simulations. However, although AICM seems to be a more stable measure of model ranking than HME, both TI and SS still seem to outperform this estimator, at least in the case of the comparison of demographic and relaxed molecular clock models [96] (see Subheading 3).
2.9.4 Cross-Validation
Cross-validation is another model selection approach, which is extremely versatile in that it can be used to compare any set of models of interest. Besides, the approach is very intuitive. In its simplest form, cross-validation consists in dividing the available data into two sets, one used for “training” and the other one used for “validating.” In the training step (TS), the model of interest is fitted to the training data in order to obtain a set of MLEs. These MLEs are then used to compute the likelihood using the validation data (validation step, VS). Because the validation data were not part of the training data, the likelihood values computed during VS can be directly used to compare models, without requiring any explicit correction for model dimensionality.
The robustness of the cross-validation scores can be explored in various ways, such as repeating the above procedure with a switched labeling of training and validation data (hence the expression cross-validation). Of course, this simple 2-fold cross-validation could be extended to n-fold cross-validation, where the data are subdivided into n subsets, with n − 1 subsets serving for training, and one for validation. Ideally, the procedure is repeated n − 1 additional times.
We know of only two examples of its use in phylogenetics, one in the ML framework [98] and one with a Bayesian approach [99]. Given the increasing size of modern data sets, putting aside some of the data for validation is probably not going to dramatically affect the information content of the whole data set. As a result, model selection via cross-validation, which is statistically sound, could become a very popular approach.
2.10 Finding the Best Tree Topology
2.10.1 Counting Trees
Now that we can select a model of evolution (Subheading 2.9) and estimate model parameters (Subheading 2.8) under a particular model (Subheading 2.5), how do we find the optimal tree? The basic example in Subheading 2.1 suggested that we score all possible tree topologies and choose for inference the one that has the highest score. However, a simple counting exercise shows that an exhaustive examination of all possible topologies is not realistic.
Figure 9 shows how to count tree topologies. Starting from the simplest possible unrooted tree, with three taxa, there are three positions where a fourth branch (leading to a fourth taxon) can be added. As a result, there are three possible topologies with four taxa. For each of these, there are four places on the tree where a fifth branch can be added, which leads to a total of 3 × 5 = 15 topologies with five taxa. A recursion appears immediately, and it can be shown that the total number of unrooted topologies with n taxa is equal to 1 × 3 ×⋯ × 2n − 5 [100] (see [15] for the deeper history), which, as given in [101], is equal to:
where the Γ function for any real number x is defined as \( \varGamma (x)={\int}_0^{\infty }{t}^{x-1}\kern0.3em {e}^{-t}\kern0.3em dt \). An approximation based on Stirling number is also given in [101].
Procedure to count the number of unrooted topologies. The top line shows the current number of taxa included in the tree below. Gray arrows indicate locations where an additional branch can be grafted to add one taxon. Black arrows show the resulting number of topologies after addition of a branch (taxon). Only one such possible topology is represented at the next step. The bottom line indicates the number of possibilities. These numbers multiply to obtain the total number of trees
The same exercise can be done for rooted trees (Fig. 10), where the number of possible rooted topologies with n taxa becomes 1 × 3 ×⋯ × 2n − 3, which is
Note that \( {N}_{\mathrm{unrooted}}^{T(n)}={N}_{\mathrm{rooted}}^{T\left(n-1\right)} \), as Table 2 clearly suggests.
Procedure to count the number of rooted topologies. See Fig. 9 for legend and text for details
As a result, the number of possible topologies quickly becomes very large when the number n of sequences increases, even with a very modest n, so that heuristics become necessary to find the best-scoring tree.
2.10.2 Some Heuristics to Find the Best Tree
The simplest approach builds upon the idea presented in Figs. 9 and 10. Stepwise addition, for instance, starts with three sequences drawn at random among the n sequences to be analyzed, and adds sequences one at a time, keeping only the tree that has the highest score at each step (e.g., [52]). However, there is no guarantee that the final tree is the optimal tree [44]. The idea behind branch-and-bound [102], refined in [103], is to have a look-ahead routine that prevents entrapment in suboptimal trees. This routine sets a bound on the trees selected at each round of additions, such that only the trees that have a score at least as good as that of the trees obtained in the next round are kept in the search algorithm. Solutions found by the branch-and-bound algorithm are optimal, but computing time becomes quickly prohibitive with more than 20 sequences.
As a result, most tree-search algorithms will start with a quickly obtained tree, often reconstructed with an algorithm based on pairwise distances such as neighbor-joining [104] or a related approach [78, 105], and then alter the tree randomly until no further improvement is obtained or after a certain number of unsuccessful attempts are reached. Examples of such algorithms include nearest neighbor interchange (NNI), subtree pruning and regrafting (SPR), or tree bisection and reconnection (TBR), see, for instance, [6] for a full description. While the details are of little importance here, the critical point is the extent of topological rearrangement in each case. With, e.g., NNI, each rearrangement can give rise to two topologies. The result is that exploring the topology space is slow, especially in problems with large n. On the other hand, TBR has, among the three methods cited above, the largest number of neighbors. As a result, the topology space is explored quickly, but the optimal tree can be “missed” simply because a dramatic change is attempted, so that the computational cost increases. Alternatively, the chance of finding the optimal tree \( \widehat{\tau} \) when \( \widehat{\tau} \) is very different from the current tree is higher when the algorithm can create some dramatic rearrangements. Some programs, such as PhyML ver. 3.0, now use a combination of NNI and SPR to address this issue [24]. MCMC samplers that search the tree space implement somewhat similar tree-perturbation algorithms that are either “global,” and modify the topology dramatically, or “local” [106] (see also [107] for a correction of the original local moves). As a result, MCMC samplers are affected by the same issues as traditional likelihood methods. Much of the difficulty therefore comes from this kind of trade-off between larger rearrangements that are expected to improve accuracy and the computational burden associated with these extra computations [108].
2.10.3 Cutting Corners with ABC and AI
As some of the above computations can become quite costly (high runtimes, heavy memory footprints, poor scalability with large data sets, etc.), computational workarounds have been and are being explored. One of these resorts to approximate Bayes computing (ABC), which is essentially a likelihood-free approach. First developed in the context of population genetics [109, 110], the driving idea is to bypass the optimization procedures and replace them with simulations in the context of a rejection sampler. In population genetics, the problem could be about a gene tree, which is usually appropriately described by a coalescence tree [111, 112], for which we want to estimate some model parameters. As we are able to simulate trees from such a process, it is possible to place prior distributions on these model parameters, and simulate trees by drawing parameters until the simulated trees “look like” the observed tree. The set of parameters thus drawn approximates the posterior distribution of the corresponding variables. This forms the basis of a naïve rejection sampler, that is quite flexible as it does not even require that a probabilistic model be formulated, but one that can be inefficient, especially if the posterior distribution is far from the prior distribution—which is usually the case. As a result, a number of variations have been described, trying either to correlate sample draws as in MCMC samplers [113] or to resample sequentially from the past [114, 115]. In spite of recent reviews of the computational promises and deliveries of ABC samplers [116,117,118], the few applications in molecular evolution have been, to date, mostly limited to molecular epidemiology [119,120,121,122]. One of the major challenges to estimate a phylogenetic tree from a sequence alignment with ABC is the lack of a proper and efficient simulation strategy: it is possible to simulate trees under various processes (we saw the coalescent above), it is also possible to simulate an alignment from a given (possibly simulated tree), so that in theory one could imagine an ABC algorithm that would use this backward process to estimate phylogenetic trees by comparing a simulated alignment to an “actual” alignment. This, however, would most likely be a very inefficient sampler.
A second area that holds promises is the use of artificial intelligence (AI), and more specifically of machine learning (ML), in molecular evolution. Here again, attempts have been made to using standard ML approaches such as support vector machines [123] to guide the comparison of tree shapes, for instance, [124], which can then be used in epidemiology [121], but estimating a phylogenetic tree has proved more challenging. In one notable exception, an alignment-free distance-based tree-reconstruction method was proposed [125], but its main legacy seems to be in the development of k-mers, or unaligned sequences chopped into words of length k, to reconstruct phylogenetic trees—in particular in the context of phylogenomics (phylogenetics at a genomics scale) [126, 127]. To the best of our knowledge, nobody has ever tried, yet, to train a neural network or even a deep learning algorithm [128,129,130] on a database of phylogenetic trees with corresponding alignments such as TreeBASE [131] or PANDIT [132]. As applications of deep learning start emerging in genomics [133] and proteomics [134], it is likely that phylogenetics will come next.
3 Uncovering Processes and Times
3.1 Dating the Tree of Life: Always Deeper?
Similar to the problem of estimating the tree of life, dating the tree of life poses many challenges [135]. Since it was first proposed in 1965 [40], the idea of estimating divergence times has since undergone a dramatic change, and new approaches are regularly proposed. Population geneticists have their own approaches, which are either fully Bayesian [136] or based on approximate Bayesian computation in the coalescent framework [137]. All these approaches make it possible to infer divergence times between recently diverged species, as in the case of humans and chimpanzees, or to date demographic events such as the migrations “out of Africa” of early human populations [138].
In the context of molecular evolution, we are usually interested in estimating deeper divergence times, such as those between species, which are available online, for instance, at www.timetree.org [139], recently revamped and extended to cover close to 100k species [140]. While early “molecular dates” were systematically biased towards ages that are too old [135], we argue here that recent developments in the field have led to more accurate methods and also to a better understanding of methodological limitations.
3.1.1 The Strict Molecular Clock
One quantity that we can estimate when comparing pairs of sequences is the number of differences that exist. This number, estimated as a branch length b, can be corrected for multiple substitutions (see Subheading 2.7), but basically remains an expected number of substitutions per site. With “dating” (defined here as the activity of estimating divergence times [141]), we are interested in estimating time t, which relates to the expected numbers of substitutions b according to the following equation:
where Δt is a period of time and r the rate of evolution. In technical terms, times and rates are said to be confounded, because we cannot estimate one without making an assumption about the other.
The molecular clock hypothesis does just this by assuming that rates of evolution are constant in time [40] (see also [142], p. 65). Under this assumption, the estimated tree is ultrametric as in the basic example represented in Fig. 11, which implies that all the tips are level, or equivalently that the distance from root to tip is the same for all branches.
The strict molecular clock. The tree is ultrametric. The node marked with a star indicates the presence of a fossil, dated in this example to 10 million years ago (MYA). This is the point that we will use to calibrate the clock, that is, to estimate the global rate of evolution. The number of substitutions that accumulated from the marked node to the tips (present) is indicated on the right weights in at 0.1 substitutions/site. The node that is the most recent common ancestor of S2 and S5 is the node of interest. The number of substitutions from this node to the tips is 0.02 substitutions/site
In this example (Fig. 11), the branch length from the fossil-dated node is 0.1 substitutions/site (sub/site), and the fossil was estimated to be present 10 million years ago (MYA). Under the strict molecular clock assumption (equal rates over the whole tree), we can (1) estimate the rate of evolution (0.1∕10 = 0.01 sub/site/my) and (2) date all the other nodes on the tree. For instance, the most recent common ancestor of S2 and S5 is separated from the tips by a branch length of 0.02 sub/site. Its divergence time is therefore 0.02∕0.01 = 2 MYA.
As with any hypothesis, the strict clock can be tested. Tests based on relative rates assess whether two species evolve at the same rate as a third one, used as an outgroup. Originally formulated in a distance-based context [143], likelihood versions have been described [44, 144]. However, because of their low power [145] their use is on the wane. The most powerful test is again the LRT (see Subheading 2.9.1). The test proceeds as usual, first calculating the test statistic 2Δℓ (twice the difference of log-likelihood values). The null hypothesis (strict clock) is nested within the alternative hypothesis (clock not enforced), so that 2Δℓ follows a χ 2 distribution. The degree of freedom is calculated following Fig. 12. With an alignment of n sequences, we can estimate n − 1 divergence times under the null model (disregarding parameters of the substitution model) and we have 2n − 3 branch lengths under the alternative model. The difference in number of free parameters is therefore n − 2, which is our degree of freedom. This version of the test actually assesses whether all tips are at the same distance from the root of the tree [44]. For time-stamped data, serially sampled in time as in the case of viruses, the alternative model incorporates information on tip dates [146].
Testing the strict molecular clock. The divergence times that can be estimated under the strict clock assumption are denoted t i. The branch lengths that can be estimated without the clock are denoted b i. In the case depicted, with n = 7 sequences, we have n − 1 = 6 divergence times and 2n − 3 = 11 branch lengths
This linear regression model suggested by the molecular clock hypothesis has often been portrayed as a recipe [147], which gave rise in the late twentieth to early twenty-first century to a veritable cottage industry [148,149,150,151], culminating with a paper suggesting that the age of the tree of life might be older than the age of planet Earth [152]. This recipe was put down by two factors: (1) the publication of a piece written in a rather unusual style for a scientific paper [153], and (2) new methodological developments. The main points made in [153] are that (1) most of the early dating studies relied on one analysis [149] that used a fossil-based calibration point for the divergence of birds at 310 MYA to estimate a number of molecular dates for vertebrates, and that (2) these molecular dates were then used in subsequent studies as a proxy for calibration points, disregarding their uncertainty. As a result, estimation errors were passed on and amplified from study to study, leading to the nonsensical results in [152].
3.1.2 Local Molecular Clocks
This “debacle” has motivated further theoretical developments in the dating field. The simplest idea is that, if a global clock does not hold for the entire tree, then perhaps groups of related species share the same rate. That is, if a global clock does not hold, perhaps the tree can be subdivided into local molecular clocks. An initial idea was proposed in the context of quartets of sequences [154] and was later generalized to a tree of any size with any number of local clocks on the tree [155] (constrained by the number of branches on the tree and calibration points). Because of the arbitrariness of such local clocks, methods have been devised to place the clocks on the tree [156] and to estimate the appropriate number of clocks that should be used [157]. A Bayesian approach now estimates all these parameters and their placement in an integrated statistical framework [158].
3.1.3 Correlated Relaxed Clocks
The idea of a correlated relaxed molecular clock goes back to Sanderson [159] (see also [160]), who considered that rates of evolution can change from branch to branch on a tree. By constraining rates of evolution to vary in an autocorrelated manner on a tree, it is possible to devise a method that minimizes the amount of rate change.
The idea of an autocorrelated process governing the evolution of the rates of evolution is attributed to [161] in [159], but could all the same be attributed to Darwin. Thorne et al. developed this idea further in a Bayesian framework [162]. Building upon the basic theory covered in Subheading 2.9.3, the idea is to place prior distributions on the quantities in the right-hand side of Eq. 28. The target distribution is p(t|X). It is proportional to p(X|t) p(t) according to Bayes’ theorem, but all that we can estimate is
One of the possible ways to expand the joint distribution of rates and times is p(r, t) is p(r|t) p(t), which posits a process where rate change depends on the length of time separating two divergences. The “art” is now in choosing prior distributions, conditional on the obvious constraint that rates and times should take positive values. A number of such prior distributions for rates have been proposed and assessed [163] and one of the best-performing model for rates is, in our experience, the log-normal model [162, 164]. The prior on times is either a pure-birth (Yule) model or a birth-and-death process possibly incorporating species sampling effects [165]. If sequences are sampled at the population level, a coalescent process is more appropriate (see [112] for an introduction). In this case, the past demography of the sampled sequences can be traced back taking inspiration from spline regression techniques [166, 167] or multiple change-point models [168].
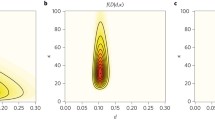
Once these priors are specified, an MCMC sampler will draw from the target distribution in Eq. 29, and marginal distributions for times and rates can easily be obtained. The rationale behind the sampler is represented in Fig. 13. As per Eq. 28, the relationship between rates and times is the branch of a hyperbolic curve, where the priors on rates and on times define a region of higher posterior probability, symbolized here by a contour plot superimposed on the hyperbolic curve. On top of this, fossil information is incorporated into the analysis as constraints on times. A very influential piece stimulated a discussion about the shape of these prior distributions [153], which was taken up [169], and further developed in [170]. Briefly, fossil information is usually imprecise, as paleontologists can only provide minimum and maximum ages (Fig. 13). Of these two ages, the minimum age is often the most reliable. Under the assumption that the placement of the fossil on the tree is correct, the idea is to place on fossil dates a prior distribution that will be highly skewed towards older (maximum) ages. A “hard bound” can be placed on the minimum age, possibly by shifting this prior distribution by an offset equal to the minimum age, while the tails of the prior distribution will act as “soft bounds,” because they do not impose on the tree a strict (or hard) constraint. Empirical studies agree, however, that both reliability and precision of fossil calibrations are critical to estimating divergence times [136, 171].
3.1.4 Uncorrelated Relaxed Clocks
Because of the autocorrelation between the rate of each branch and that of its ancestral branch (except for the root, which obviously requires a special treatment), the tree topology is fixed under the autocorrelated models described above. By relaxing this assumption about rate autocorrelation, [172] were able to implement a model that also integrates over topological uncertainty. In spite of the somewhat counter-intuitive nature of the relaxation of the autocorrelated process, as implemented in BEAST [91, 173], empirical studies have found this approach to be one of the best-performing (e.g., [157]).
When first published, it was proposed that making use of an uncorrelated relaxed molecular clock could improve phylogenetic inference [172]. The idea was that calibration points and their placement on the tree could act as additional information. However, a simulation study suggests that relaxed molecular clocks might not improve phylogenetic accuracy [174], a result that might be due to the lack of calibration constraints in this particular simulation study.
3.1.5 Some Applications of Relaxed Clock Models
Since the advent of relaxed molecular clocks, two very exciting developments have seen the light of day. The first concerns the inclusion of spatial statistics into dating models [175, 176]. Spatial statistics are not new in population genetics [177] and have been used with success in combination with analyses in computational molecular evolution (e.g., [178]). However, the originality in [176], for instance, is to combine in a single statistical framework molecular data with geographical and environmental information to infer the diffusion of sequences through both space and time. While these preliminary models seem to deal appropriately with natural barriers to gene flow such as coastlines, a more detailed set of constraints on gene flow may further enhance their current predictive power.
The second development coming from relaxed molecular clocks concerns the mapping of ancestral characters onto uncertain phylogenies. This is not a novel topic, as a Bayesian approach was first described in 2004 [179, 180]. The novelty is that we now have the tools to correlate morphological and molecular evolution in terms of their absolute rates and to allow both molecular and morphological rates of evolution to vary in time [181]. Further development will certainly integrate over topological uncertainty. While there has been a heated controversy about the existence of such a correlation in the past [182], all previous studies were using branch length as a proxy for rate of molecular evolution, which is clearly incorrect. We can therefore expect some more accurate results on this topic very soon. More details and examples can be found in recent and extensive reviews [183,184,185] that further discuss applications to biogeographic studies [186], or extensions to viral [187, 188], as well as other types of genomic [189] and morphological [190] data.
4 Molecular Population Phylogenomics
Population genetics is rich in theory regarding the relative roles of mutation, drift, and selection. Much research in population genomics is now focusing on using this theory to develop statistical procedures to infer past processes based on population-level data, such as those of the 1000-genome project [191], the UK’s 10,000 genome project [192], and always more ambitious projects [193]. One limitation of these inference procedures is that they all focus on a thin slice of evolutionary time by studying evolution at the level of populations. If we wish to study longer evolutionary time scales, for example, tens or hundreds of millions of years, we must resort to interspecific data. In such a context, which is becoming intrinsically phylogenetic, the most important event is a substitution, that is, a mutation that has been fixed. Yet substitution rates can be defined from several features. In particular, from a population genetics perspective, it is of interest to model both mutational features and selective effects, combining them multiplicatively to specify substitution rates. We review briefly how substitution models that invoke codons as the state space lend themselves naturally to these objectives in a first section below (Subheading 4.1), before explaining the origin (and a shortcoming) of all the approaches developed so far (Subheading 4.2).
4.1 Bridging the Gap Between Population Genetics and Phylogenetics
Assuming a point-mutation process, such that events only change one nucleotide of a codon during a small time interval, Muse and Gaut proposed a codon substitution model with rates specified from the Q GTR nucleotide-level matrix (see Subheading 2.7), along with one parameter that modulates synonymous events and another one that modulates nonsynonymous events [194]. In most subsequent formulations, the parameter associated with synonymous events is assumed to be fixed, such that the model only modulates nonsynonymous rates by means of a parameter denoted ω. This parameter has traditionally been interpreted as the nonsynonymous to synonymous rate ratio, and is generally associated with a different formulation of the codon model proposed by Goldman and Yang [195]. More details on codon models can be found in Chapter 4.1 [196]. There continues to be a debate regarding the interpretation of the ω parameter [197, 198]. Regardless of how this issue is settled, it is clear that ω is aimed at capturing the net overall effects of selection, irrespective of the exact nature of these effects.
With the intention to model selective effects themselves, Halpern and Bruno proposed a codon substitution model that combines a nucleotide-level layer, as described above, for controlling mutational features, along with a fixation factor that is proportional to the fixation probability of the mutational event [199]. The fixation factor is in turn specified from an account of amino acid or codon preferences. One objective of the model, then, consists in teasing apart mutation and selection. While [199] proposed their model with site-specific fixation factors, later work has explored simpler specifications, where all sites have the same fixation factor [200]. Other models that aimed at capturing across-site heterogeneities in fixation factors were proposed using nonparametric devices and empirical mixtures [201]. Another core idea behind these approaches is to construct a more appropriate null model against which to test for features of the evolutionary process. This idea has been put into practice for the detection of adaptive evolution in protein-coding genes [202, 203]. Recent developments include sequence-wide fixation factors [9, 197, 204, 205], and we predict that these models will play a role in bridging the gap between molecular evolution at the population and at the species levels.
4.2 Origin of Mutation–Selection Models: The Genic Selection Model
In order to understand a shortcoming of these models, we need to go back to the development of fixation probabilities that took place in the second half of the twentieth century. The basic unit or quantum of evolution is a change in allele frequency p. Allele frequencies can be affected by four processes: migration, mutation, selection, and drift. Because of the symmetry between migration and mutation [206], which only differ in their magnitude, these two processes can be treated as one. We are left with three forces: mutation, selection, and drift. The question is then, what is the fate of an allele under the combined action of these processes? Our development here follows [207] (but see [208] for a very clear account).
4.2.1 Fixation Probabilities
Of the three processes affecting allele frequencies, mutation and selection can be seen as directional forces in that their action will shift the distribution of allele frequencies towards a particular point, be it an internal equilibrium, or fixation/loss of an allele. On the other hand, drift is a non-directional process that will increase the variance in allele frequencies across populations, and will therefore spread out the distribution of allele frequencies. This distribution is denoted Ψ(p, t). We also must assume that the magnitude of all three processes, mutation, selection, and drift, is small and of the order of \( \frac{1}{2{N}_e} \), where N e is the effective population size. To derive the fate of an allele after a certain number of generations, we also need to define g(p, ε;dt), the probability that allele frequency changes from p to p + ε during a time interval dt.
In phylogenetics (and population genetics) we are generally interested in predicting the past. The tool making this possible is called the Kolmogorov backward equation, which predicts the frequency of an allele at some time t, given its frequency p 0 at time t 0:
We can take the Taylor expansion of Eq. 30 around p 0, neglect all terms whose order is larger than two (\( o\left({p}_0^2\right) \)) and since Ψ is not a function of ε, we obtain:
This formulation leads to the definition of two terms that represent the directional processes affecting allele frequencies (M) and the non-directional process, or drift (V ):
that we can substitute into Eq. 31. At equilibrium, \( \frac{\partial \varPsi }{\partial t}=0 \) and, after a bit of calculus, we obtain:
for which we need to specify boundary conditions and a model of selection. The boundary conditions are the two absorbing states of the system: (1) once fixed, an allele remains fixed (Ψ(1, ∞; 1) = 1) and (2) once lost, an allele remains lost (Ψ(1, ∞; 0) = 0). With these two requirements, the probability that the allele frequency is 1 given that it was p 0 in the distant past is the fixation probability:
We therefore only need to compute M and V under a particular model of selection to fully specify the fixation probability of an allele in a mutation–selection-drift system. All that is required now to go further is a selection model.
4.2.2 The Case of Genic Selection
We are now ready to derive an explicit form to Ψ(1, ∞; p 0) in Eq. 34 in the case of the genic selection model (Table 3; [209]). We obtain:
which can be approximated by 1 + 2phs (the result is exact only when h = 1∕2). Therefore, \( d\overline{w}/ dp=2 hs \), and we can calculate the M and V terms to obtain the popular result:
Now, the initial frequency of a mutation in a diploid population of (census) size N is p 0 = 1∕2N (following [208]; [207] considered that p 0 = 1∕2N e; this debate is beyond the scope of this chapter), which leads to:
If N e is of the order of N, the numerator of the right-hand side of Eq. 37 becomes approximately e −2hs − 1, whose Taylor approximation around hs = 0 is simply − 2hs. We then obtain the result used in [199], and in all the papers that implemented mutation–selection (-drift) models (e.g., [197, 199,200,201, 204]):
Two critical points should be noted here. First, none of the recent codon models [197, 199,200,201,202, 204, 210, 211] ever investigated the role of dominance h, as they all consider that the allele under (positive) selection is fully dominant. Second, Table 3 shows that another class of selection models, those based on balancing selection, has never been considered so far. The impact of the selection model on the predictions made by the mutation–selection (-drift) models is currently unknown.
5 High-Performance Computing for Phylogenetics
5.1 Parallelization
Because of the dependency of the likelihood computations on the shape of a particular tree (see Subheading 2.6), most phylogenetic computations cannot be parallelized to take advantage of a multiprocessor (or multicore) environment. Nevertheless, two main directions have been explored to speed up computations: first, in computing the likelihood of substitution models that incorporate among-site rate variation and second, in distributing bootstrap replicates to several processors, as both types of computations can be done independently. A third route is explored in Chapter 7.4 [212].
In the first case, among-site rate variation is usually modeled with a Γ distribution [213] that is discretized over a finite (and small) number of categories [214]. The likelihood then takes the form of a weighted sum of likelihood functions, one for each discrete rate category, so that each of these functions can be evaluated independently. The route most commonly used is the plain “embarrassingly parallel” solution, where completely independent computations are farmed out to different processors. Such is the case for bootstrap replicates, for which a version of PhyML [24] exists, or in a Bayesian context for independent MCMC samplers [215] (see Subheading 2.9.3). The PhyloBayes-MPI package implements distributed likelihood calculations across sites over several compute-cores, allowing for a genuinely parallelized MCMC run [216, 217].
5.2 HPC and Cloud Computing
More recent work has focused on the development of heuristics that make large-scale phylogenetics amenable to high-performance computing (HPC) that are performed on computer clusters. Because of the algorithmic complexity of resolving phylogenetic trees, an approach based on “algorithmic engineering” was developed [218]. The underlying idea is akin to the training phase in supervised machine learning [123], except that here the target is not the performance of a classifier but that of search heuristics. All of these heuristics reuse parameter estimates, avoid the computation of the full likelihood function for all the bootstrap replicates, or seed the search algorithm for every n replicate on the results of previous replicates [218]. For instance, in the “lazy subtree rearrangement” [219], topologies are modified by SPR (see Subheading 2.10.2), but instead of recomputing the likelihood on the whole tree, only the branch lengths around the perturbation are re-optimized. This approximation is used to rank candidate topologies, and the actual likelihood is evaluated on the complete tree only for the best candidates. These heuristics now permit the analysis of thousands of sequences in a probabilistic framework [220], but the actual convergence of these algorithms remains difficult to evaluate, especially on very large data sets (e.g., >104 sequences).
In addition to the reduction of the memory footprint for sparse data matrices [221], an alternative direction to “tweaking likelihood algorithms” has been to take direct advantage of the computing architecture available. One particular effort aims at tapping directly into the computing power of graphics processing units or GPUs, taking advantage of their shared common memory, their highly parallelized architecture, and the comparatively negligible cost of spawning and destroying threads on them. As a result, it is possible to distribute some of the summation entering the pruning algorithm (see Subheading 2.6) to different GPUs [222]. The number of programs taking advantage of these developments is widening and includes popular options such as BEAST [91] and MrBayes [223].
All these fast algorithms can be installed on a local computer cluster, a solution adopted by many research groups since the late 1990s. However, installing a cluster can be demanding and costly because a dedicated room is required with appropriate cooling and power supply (not to mention securing the room, physically). Besides, redundancy requirements, both in terms of power supply and data storage, as well as basic software maintenance and user management, may demand hiring a system administrator. An alternative is to run analyses on a remote HPC server, in the “cloud.” Canada, for instance, has a number of such facilities, thanks to national funding bodies (CAC at cac.queensu.ca, SHARCNET at www.sharcnet.ca, or Calcul Quebec at www.calculquebec.ca, just to name a few), and commercial solutions are just a few clicks away (e.g., Amazon Elastic Compute Cloud or EC2). Researchers can obtain access to these HPC solutions according to a number of business models (free, on demand, yearly subscription, etc.) that are associated with a wide spectrum of costs [224]. But in spite of the technical support offered in the price, users usually still have to install their preferred phylogenetic software manually or put a formal request to the team of system administrators managing the HPC facility, all of which is not always convenient.
To make the algorithmic and technological developments described above more accessible, the recent past has seen the emergence of cloud computing [225] dedicated to the phylogenetics community. Examples include the CIPRES Science Gateway (www.phylo.org), or Phylogeny.fr (www.phylogeny.fr, [226]). Many include web portals that do not require that users be well versed in Unix commands, while others may include an application programming interface to cater to the most computer-savvy users. One potential limitation of these services is the bandwidth necessary to transfer large files, and storage requirements—especially in the context of next generation sequencing data. The management of relatively large files will remain a potential issue, unless phylogenetics practitioners are ready to discard these files after analysis, the end product of which is a single tree file a few kilobytes in size, in the same way that people involved in genome projects delete the original image files produced by massively parallel sequencers. Data security or privacy might not be a problem in most applications, except in projects dealing with human subjects or viruses such as HIV that expose the sexual practices of subjects. However, once these various hurdles are out of the way, users could very well imagine running their phylogenetic analyses with millions of sequences from their smartphone while commuting.
6 Conclusions
Although most of the initial applications of likelihood-based methods were motivated by the shortcomings of parsimony, they have now become well accepted as they constitute principled inference approaches that rely on probabilistic logic. Moreover, they allow biologists to evaluate more rigorously the relative importance of different aspects of evolution. The models presented in this chapter have the ability to disentangle rates from times (Subheading 3), or mutation from selection (Subheading 4), while in most cases accounting for the uncertainty about nuisance parameters. But the latest developments described above still make a number of restrictive assumptions (Subheading 4.2), and while many variations in model formulations can be envisaged, they still remain to be explored in practice.
Although some progress has been made in developing integrative approaches (e.g., [176, 181]), throughout this chapter we have assumed that a reliable alignment was available as a starting point. A number of methods exist to co-estimate an alignment and a phylogenetic tree (see Part I of this book), but the computational requirements and convergence of some of these approaches can be daunting, even on the smallest data sets by today’s standards.
This brings us, finally, to the issue of tractability of most of these models in the face of very large data sets. The field of phylogenomics is developing quickly (see Part III), at a pace that is ever increasing given the output rate of whole genome sequencing projects. Environmental questions are drawing more and more attention, and metagenomes (see Part VI) will be analyzed in the context of what will soon be called metaphylogenomics. Exploring the numerous available and foreseeable substitution models in such contexts will require continued work in computational methodologies. As such, modeling efforts will continue to go hand-in-hand with, and maybe dependent on, algorithmic developments [227]. It is also not impossible that in the near future, the use of likelihood-free approach such as ABC or machine learning algorithms in computational molecular evolution be more thoroughly explored.
References
Nei M, Kumar S (2000) Molecular evolution and phylogenetics. Oxford University Press, Oxford
Higgs PG, Attwood TK (2005) Bioinformatics and molecular evolution. Blackwell Publishing, Oxford
Balding DJ, Bishop MJ, Cannings C (2007) Handbook of statistical genetics, 3rd edn. Wiley, Chichester
Salemi M, Vandamme A-M, Lemey P (2009) The phylogenetic handbook: a practical approach to phylogenetic analysis and hypothesis testing, 2nd edn. Cambridge University Press, Cambridge
Hall BG (2011) Phylogenetic trees made easy: a how to manual. Sinauer Associates, Sunderland
Yang Z (2014) Molecular evolution: a statistical approach. Oxford University Press, Oxford
Drummond AJ, Bouckaert RR (2015) Bayesian evolutionary analysis with BEAST. Cambridge University Press, Cambridge
Aris-Brosou S, Xia X (2008) Phylogenetic analyses: a toolbox expanding towards Bayesian methods. Int J Plant Genomics 2008:683509
Rodrigue N, Philippe H (2010) Mechanistic revisions of phenomenological modeling strategies in molecular evolution. Trends Genet 26:248–252
Yang Z, Rannala B (2012) Molecular phylogenetics: principles and practice. Nat Rev Genet 13:303–314
Aris-Brosou S, Rodrigue N (2012) The essentials of computational molecular evolution. Methods Mol Biol 855:111–152
Yang Z (2000) Complexity of the simplest phylogenetic estimation problem. Proc Biol Sci 267:109–116
Sober E (1988) Reconstructing the past: parsimony, evolution, and inference. MIT Press, Cambridge
Durbin R, Eddy SR, Krogh A, Mitchison G (1998) Biological sequence analysis: probabilistic models of proteins and nucleic acids. Cambridge University Press, Cambridge
Felsenstein J (2004) Inferring phylogenies. Sinauer Associates, Sunderland
Yang Z (2007) PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591
Efron B, Tibshirani R (1993) An introduction to the bootstrap, vol 57. Chapman and Hall, Boca Raton
Efron B, Halloran E, Holmes S (1996) Bootstrap confidence levels for phylogenetic trees. Proc Natl Acad Sci USA 93:7085–7090
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Baldauf SL (2003) Phylogeny for the faint of heart: a tutorial. Trends Genet 19:345–351
Hasegawa M, Kishino H (1989) Confidence limits of the maximum-likelihood estimate of the hominoid three from mitochondrial-DNA sequences. Evolution 43:672–677
Anisimova M, Gascuel O (2006) Approximate likelihood-ratio test for branches: a fast, accurate, and powerful alternative. Syst Biol 55:539–552
Guindon S, Delsuc F, Dufayard J-F, Gascuel O (2009) Estimating maximum likelihood phylogenies with phyml. Methods Mol Biol 537:113–137
Guindon S, Dufayard J-F, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321
Hillis DM, Bull JJ (1993) An empirical test of bootstrapping as a method for assessing confidence in phylogenetic analysis. Syst Biol 42:182–192
Felsenstein J, Kishino H (1993) Is there something wrong with the bootstrap on phylogenies? A reply to Hillis and Bull. Syst Biol 42:193–200
Yang Z, Rannala B (2005) Branch-length prior influences Bayesian posterior probability of phylogeny. Syst Biol 54:455–470
Berry V, Gascuel O (1996) On the interpretation of bootstrap trees: appropriate threshold of clade selection and induced gain. Mol Biol Evol 13:999
Shimodaira H, Hasegawa M (2001) CONSEL: for assessing the confidence of phylogenetic tree selection. Bioinformatics 17:1246–1247
Salichos L, Rokas A (2013) Inferring ancient divergences requires genes with strong phylogenetic signals. Nature 497:327–331
Felsenstein J (1978) Cases in which parsimony or compatibility methods will be positively misleading. Syst Zool 27:401–410
Tuffley C, Steel M (1997) Links between maximum likelihood and maximum parsimony under a simple model of site substitution. Bull Math Biol 59:581–607
Steel M, Penny D (2000) Parsimony, likelihood, and the role of models in molecular phylogenetics. Mol Biol Evol 17:839–850
Holder MT, Lewis PO, Swofford DL (2010) The Akaike information criterion will not choose the no common mechanism model. Syst Biol 59:477–485
Editors T (2016) Editorial. Cladistics 32:1. https://doi.org/10.1111/cla.12148
Philippe H, Zhou Y, Brinkmann H, Rodrigue N, Delsuc F (2005) Heterotachy and long-branch attraction in phylogenetics. BMC Evol Biol 5:50
Brinkmann H, van der Giezen M, Zhou Y, de Raucourt GP, Philippe H (2005) An empirical assessment of long-branch attraction artefacts in deep eukaryotic phylogenomics. Syst Biol 54:743–757
Hampl V, Hug L, Leigh JW, Dacks JB, Lang BF, Simpson AG, Roger AJ (2009) Phylogenomic analyses support the monophyly of Excavata and resolve relationships among eukaryotic “supergroups”. Proc Natl Acad Sci USA 106:3859–3864
Liu H, Aris-Brosou S, Probert I, de Vargas C (2010) A timeline of the environmental genetics of the haptophytes. Mol Biol Evol 27:161–176
Zuckerkandl E, Pauling L (1965) Evolutionary divergence and convergence in proteins. In: Bryson V, Vogel HJ (eds) Evolving genes and proteins. Academic, Cambridge, pp 97–166
Galtier N, Gascuel O, Jean-Marie A (2005) Markov models in molecular evolution. In: Nielsen R (ed) Statistical methods in molecular evolution. Statistics for biology and health. Springer, New York, pp 3–24
Cox DR, Miller HD (1965) The theory of stochastic processes. Chapman and Hall/CRC, Boca Raton
Yang Z (2000) Maximum likelihood estimation on large phylogenies and analysis of adaptive evolution in human influenza virus A. J Mol Evol 51:423–432
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Jukes JC, Cantor CR (1969) Evolution of protein molecules. In: Munro HN (ed) Mammalian protein metabolism. Academic, New York, pp 21–123
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Hasegawa M, Kishino H, Yano T (1985) Dating of the human-ape splitting by a molecular clock of mitochondrial DNA. J Mol Evol 22:160–174
Tavaré S (1986) Some probabilistic and statistical problems in the analysis of DNA sequences. Lect Math Life Sci 17:57–86
Huelsenbeck JP, Larget B, Alfaro ME (2004) Bayesian phylogenetic model selection using reversible jump Markov chain Monte Carlo. Mol Biol Evol 21:1123–1133
Yang Z, Roberts D (1995) On the use of nucleic acid sequences to infer early branchings in the tree of life. Mol Biol Evol 12:451–458
Huelsenbeck JP, Bollback JP, Levine AM (2002) Inferring the root of a phylogenetic tree. Syst Biol 51:32–43
Yang Z (2006) Computational molecular evolution. Oxford University Press, Oxford
Aris-Brosou S (2005) Determinants of adaptive evolution at the molecular level: the extended complexity hypothesis. Mol Biol Evol 22:200–209
Anisimova M, Yang Z (2004) Molecular evolution of the hepatitis delta virus antigen gene: recombination or positive selection? J Mol Evol 59:815–826
Burnham KP, Anderson DR (1998) Model selection and inference: a practical information-theoretic approach. Springer, Berlin
Anisimova M, Bielawski JP, Yang Z (2001) Accuracy and power of the likelihood ratio test in detecting adaptive molecular evolution. Mol Biol Evol 18:1585–1592
Whelan S, Goldman N (2004) Estimating the frequency of events that cause multiple-nucleotide changes. Genetics 167:2027–2043
Wong WS, Yang Z, Goldman N, Nielsen R (2004) Accuracy and power of statistical methods for detecting adaptive evolution in protein coding sequences and for identifying positively selected sites. Genetics 168:1041–1051
Massingham T, Goldman N (2005) Detecting amino acid sites under positive selection and purifying selection. Genetics 169:1753–1762
Zhang J, Nielsen R, Yang Z (2005) Evaluation of an improved branch-site likelihood method for detecting positive selection at the molecular level. Mol Biol Evol 22:2472–2479
Anisimova M, Yang Z (2007) Multiple hypothesis testing to detect lineages under positive selection that affects only a few sites. Mol Biol Evol 24:1219–1228
Yang Z (2010) A likelihood ratio test of speciation with gene flow using genomic sequence data. Genome Biol Evol 2:200–211
Fletcher W, Yang Z (2010) The effect of insertions, deletions, and alignment errors on the branch-site test of positive selection. Mol Biol Evol 27:2257–2267
Yang Z, dos Reis M (2011) Statistical properties of the branch-site test of positive selection. Mol Biol Evol 28:1217–1228
Self SG, Liang K-Y (1987) Asymptotic properties of maximum likelihood estimators and likelihood ratio tests under nonstandard conditions. J Am Stat Assoc 82:605–610
Posada D, Crandall KA (1998) MODELTEST: testing the model of DNA substitution. Bioinformatics 14:817–818
Posada D (2008) jModelTest: phylogenetic model averaging. Mol Biol Evol 25:1253–1256
Cunningham CW, Zhu H, Hillis DM (1998) Best-fit maximum-likelihood models for phylogenetic inference: empirical tests with known phylogenies. Evolution 52:978–987
Pol D (2004) Empirical problems of the hierarchical likelihood ratio test for model selection. Syst Biol 53:949–962
Kullback S, Leibler RA (1951) On information and sufficiency. Ann Math Stat 22:79–86
Minin V, Abdo Z, Joyce P, Sullivan J (2003) Performance-based selection of likelihood models for phylogeny estimation. Syst Biol 52:674–683
Ripplinger J, Sullivan J (2008) Does choice in model selection affect maximum likelihood analysis? Syst Biol 57:76–85
Posada D, Crandall KA (2001) Selecting the best-fit model of nucleotide substitution. Syst Biol 50:580–601
Abdo Z, Minin VN, Joyce P, Sullivan J (2005) Accounting for uncertainty in the tree topology has little effect on the decision-theoretic approach to model selection in phylogeny estimation. Mol Biol Evol 22:691–703
Luo A, Qiao H, Zhang Y, Shi W, Ho SY, Xu W, Zhang A, Zhu C (2010) Performance of criteria for selecting evolutionary models in phylogenetics: a comprehensive study based on simulated datasets. BMC Evol Biol 10:242
Schwarz G (1978) Estimating the dimension of a model. Ann Stat 6:461–464
Evans J, Sullivan J (2011) Approximating model probabilities in Bayesian information criterion and decision-theoretic approaches to model selection in phylogenetics. Mol Biol Evol 28:343–349
Gascuel O (1997) BIONJ: an improved version of the NJ algorithm based on a simple model of sequence data. Mol Biol Evol 14:685–695
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9:772–772
Lefort V, Longueville J-E, Gascuel O (2017) SMS: smart model selection in PhyML. Mol Biol Evol 34:2422–2424
Kleinman CL, Rodrigue N, Bonnard C, Philippe H, Lartillot N (2006) A maximum likelihood framework for protein design. BMC Bioinformatics 7:326
Rodrigue N, Philippe H, Lartillot N (2007) Exploring fast computational strategies for probabilistic phylogenetic analysis. Syst Biol 56:711–726
Yang Z (2005) Bayesian inference in molecular phylogenetics. In: Gascuel O (ed) Mathematics of evolution and phylogeny. Oxford University Press, Oxford, pp 63–90
Jeffreys H (1939) Theory of probability. The International series of monographs on physics. The Clarendon Press, Oxford
Kass RE, Raftery AE (1995) Bayes factors. J Am Stat Assoc 90:773–795
Lartillot N, Philippe H (2006) Computing Bayes factors using thermodynamic integration. Syst Biol 55:195–207
Fan Y, Wu R, Chen MH, Kuo L, Lewis PO (2011) Choosing among partition models in Bayesian phylogenetics. Mol Biol Evol 28:523–32
Newton MA, Raftery AE (1994) Approximating Bayesian inference with the weighted likelihood bootstrap. J R Stat Soc B 56:3–48
Aris-Brosou S (2003) How Bayes tests of molecular phylogenies compare with frequentist approaches. Bioinformatics 19:618–624
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Drummond AJ, Rambaut A (2007) BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol 7:214
Raftery AE (1996) Hypothesis testing and model selection. In: Gilks WR, Richardson S, Spiegelhalter DJ (eds) Markov chain Monte Carlo in practice. Chapman & Hall, Boca Raton, pp 163–187
Ogata Y (1989) A Monte Carlo method for high dimensional integration. Numer Math 55:137–157
Gelman A, Meng X-L (1998) Simulating normalizing constants: from importance sampling to bridge sampling to path sampling. Stat Sci 13:163–185
Xie W, Lewis PO, Fan Y, Kuo L, Chen MH (2011) Improving marginal likelihood estimation for Bayesian phylogenetic model selection. Syst Biol 60:150–60
Baele G, Lemey P, Bedford T, Rambaut A, Suchard MA, Alekseyenko AV (2012) Improving the accuracy of demographic and molecular clock model comparison while accommodating phylogenetic uncertainty. Mol Biol Evol 29:2157–2167
Raftery AE, Newton MA, Satagopan JM, Krivitsky PN (2007) Estimating the integrated likelihood via posterior simulation using the harmonic mean identity. Bayesian Stat 8:1–45
Smyth P (2000) Model selection for probabilistic clustering using cross-validated likelihood. Stat Comput 10:63–72
Lartillot N, Brinkmann H, Philippe H (2007) Suppression of long-branch attraction artefacts in the animal phylogeny using a site-heterogeneous model. BMC Evol Biol 7(Suppl 1):S4
Cavalli-Sforza LL, Edwards AW (1967) Phylogenetic analysis. Models and estimation procedures. Am J Hum Genet 19:233–257
Aris-Brosou S (2003) Least and most powerful phylogenetic tests to elucidate the origin of the seed plants in the presence of conflicting signals under misspecified models. Syst Biol 52:781–793
Foulds LR, Penny D, Hendy MD (1979) A general approach to proving the minimality of phylogenetic trees illustrated by an example with a set of 23 vertebrates. J Mol Evol 13:151–166
Hendy MD, Penny D (1982) Branch and bound algorithms to determine minimal evolutionary trees. Math Biosci 59:277–290
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Bruno WJ, Socci ND, Halpern AL (2000) Weighted neighbor joining: a likelihood-based approach to distance-based phylogeny reconstruction. Mol Biol Evol 17:189–197
Larget B, Simon D (1999) Markov chain Monte Carlo algorithms for the Bayesian analysis of phylogenetic trees. Mol Biol Evol 16:750
Holder MT, Lewis PO, Swofford DL, Larget B (2005) Hastings ratio of the LOCAL proposal used in Bayesian phylogenetics. Syst Biol 54:961–965
Whelan S (2007) New approaches to phylogenetic tree search and their application to large numbers of protein alignments. Syst Biol 56:727–740
Pritchard JK, Seielstad MT, Perez-Lezaun A, Feldman MW (1999) Population growth of human y chromosomes: a study of Y chromosome microsatellites. Mol Biol Evol 16:1791–1798
Beaumont MA, Zhang W, Balding DJ (2002) Approximate Bayesian computation in population genetics. Genetics 162:2025–2035
Kingman JFC (1982) The coalescent. Stoch Process Appl 13:235–248
Hein J, Schierup MH, Wiuf C (2005) Gene genealogies, variation and evolution: a primer in coalescent theory. Oxford University Press, Oxford
Marjoram P, Molitor J, Plagnol V, Tavaré S (2003) Markov chain Monte Carlo without likelihoods. Proc Natl Acad Sci 100:15324–15328
Sisson SA, Fan Y, Tanaka MM (2007) Sequential Monte Carlo without likelihoods. Proc Natl Acad Sci 104:1760–1765
Toni T, Welch D, Strelkowa N, Ipsen A, Stumpf MP (2009) Approximate Bayesian computation scheme for parameter inference and model selection in dynamical systems. J R Soc Interface 6:187–202
Beaumont MA (2010) Approximate Bayesian computation in evolution and ecology. Annu Rev Ecol Evol Syst 41:379–406
Sunnåker M, Busetto AG, Numminen E, Corander J, Foll M, Dessimoz C (2013) Approximate Bayesian computation. PLoS Comput Biol 9:e1002803
Lintusaari J, Gutmann MU, Dutta R, Kaski S, Corander J (2017) Fundamentals and recent developments in approximate Bayesian computation. Syst Biol 66:e66–e82
Ratmann O, Donker G, Meijer A, Fraser C, Koelle K (2012) Phylodynamic inference and model assessment with approximate Bayesian computation: influenza as a case study. PLoS Comput Biol 8:e1002835
Zheng Y, Aris-Brosou S (2013) Approximate Bayesian computation algorithms for estimating network model parameters. In: Joint statistical meeting proceedings (2013)—biometrics section, pp 2239–2253
Poon AF (2015) Phylodynamic inference with kernel ABC and its application to HIV epidemiology. Mol Biol Evol 32:2483–2495
Ibeh N, Aris-Brosou S (2016) Estimation of sub-epidemic dynamics by means of sequential Monte Carlo approximate Bayesian computation: an application to the Swiss HIV cohort study. https://doi.org/10.1101/085993
Hastie T, Tibshirani R, Friedman JH (2009) The elements of statistical learning: data mining, inference, and prediction. Springer series in statistics, 2nd edn. Springer, New York
Poon AF, Walker LW, Murray H, McCloskey RM, Harrigan PR, Liang RH (2013) Mapping the shapes of phylogenetic trees from human and zoonotic RNA viruses. PLoS One 8:e78122
Schwarz RF, Fletcher W, Förster F, Merget B, Wolf M, Schultz J, Markowetz F (2010) Evolutionary distances in the twilight zone—a rational kernel approach. PLoS One 5:e15788
Höhl M, Ragan MA (2007) Is multiple-sequence alignment required for accurate inference of phylogeny? Syst Biol 56:206–221
Sanderson M, Nicolae M, McMahon M (2017) Homology-aware phylogenomics at gigabase scales. Syst Biol 66:590–603
Jordan MI, Mitchell TM (2015) Machine learning: trends, perspectives, and prospects. Science 349:255–260
Rusk N (2016) Deep learning. Nat Methods 13:35
Esteva A, Kuprel B, Novoa RA, Ko J, Swetter SM, Blau HM, Thrun S (2017) Dermatologist-level classification of skin cancer with deep neural networks. Nature 542:115–118
Morell V (1996) TreeBASE: the roots of phylogeny. Science 273:569
Whelan S, de Bakker PIW, Quevillon E, Rodriguez N, Goldman N (2006) PANDIT: an evolution-centric database of protein and associated nucleotide domains with inferred trees. Nucleic Acids Res 34:D327–D331
Zhou J, Troyanskaya OG (2015) Predicting effects of noncoding variants with deep learning-based sequence model. Nat Methods 12:931–934
Tran NH, Zhang X, Xin L, Shan B, Li M (2017) De novo peptide sequencing by deep learning. Proc Natl Acad Sci. https://doi.org/10.1073/pnas.1705691114
Benton MJ, Ayala FJ (2003) Dating the tree of life. Science 300:1698–700
Rannala B, Yang Z (2007) Inferring speciation times under an episodic molecular clock. Syst Biol 56:453–66
Wegmann D, Leuenberger C, Excoffier L (2009) Efficient approximate Bayesian computation coupled with Markov chain Monte Carlo without likelihood. Genetics 182:1207–1218
Reich D, Green RE, Kircher M et al (2010) Genetic history of an archaic hominin group from Denisova Cave in Siberia. Nature 468:1053–1060
Hedges SB, Dudley J, Kumar S (2006) TimeTree: a public knowledge-base of divergence times among organisms. Bioinformatics 22:2971–2972
Kumar S, Stecher G, Suleski M, Hedges SB (2017) TimeTree: a resource for timelines, timetrees, and divergence times. Mol Biol Evol 34:1812–1819
Welch JJ, Bromham L (2005) Molecular dating when rates vary. Trends Ecol Evol 20:320–327
Kimura M (1983) The neutral theory of molecular evolution. Cambridge University Press, Cambridge
Sarich VM, Wilson AC (1973) Generation time and genomic evolution in primates. Science 179:1144–1147
Muse SV, Weir BS (1992) Testing for equality of evolutionary rates. Genetics 132:269–276
Bromham L, Penny D, Rambaut A, Hendy MD (2000) The power of relative rates tests depends on the data. J Mol Evol 50:296–301
Rambaut A (2000) Estimating the rate of molecular evolution: incorporating non-contemporaneous sequences into maximum likelihood phylogenies. Bioinformatics 16:395–399
Martin AP (2001) Molecular clocks. Encyclopedia of life sciences. Wiley, Hoboken, pp 1–6
Wray GA, Levinton JS, Shapiro LH (1996) Molecular evidence for deep Precambrian divergences among Metazoan phyla. Science 274:568–573
Kumar S, Hedges SB (1998) A molecular timescale for vertebrate evolution. Nature 392:917–920
Wang DY, Kumar S, Hedges SB (1999) Divergence time estimates for the early history of animal phyla and the origin of plants, animals and fungi. Proc Biol Sci 266:163–171
Heckman DS, Geiser DM, Eidell BR, Stauffer RL, Kardos NL, Hedges SB (2001) Molecular evidence for the early colonization of land by fungi and plants. Science 293:1129–1133
Hedges SB, Chen H, Kumar S, Wang DY, Thompson AS, Watanabe H (2001) A genomic timescale for the origin of eukaryotes. BMC Evol Biol 1:4
Graur D, Martin W (2004) Reading the entrails of chickens: molecular timescales of evolution and the illusion of precision. Trends Genet 20:80–86
Rambaut A, Bromham L (1998) Estimating divergence dates from molecular sequences. Mol Biol Evol 15:442–448
Yoder AD, Yang Z (2000) Estimation of primate speciation dates using local molecular clocks. Mol Biol Evol 17:1081–1090
Yang Z (2004) A heuristic rate smoothing procedure for maximum likelihood estimation of species divergence times. Acta Zool Sin 50:645–656
Aris-Brosou S (2007) Dating phylogenies with hybrid local molecular clocks. PLoS One 2:e879
Drummond AJ, Suchard MA (2010) Bayesian random local clocks, or one rate to rule them all. BMC Biol 8:114
Sanderson M (1997) A nonparametric approach to estimating divergence times in the absence of rate constancy. Mol Biol Evol 14:1218
Sanderson MJ (2002) Estimating absolute rates of molecular evolution and divergence times: a penalized likelihood approach. Mol Biol Evol 19:101–109
Gillespie JH (1991) The causes of molecular evolution. Oxford University Press, Oxford
Thorne JL, Kishino H, Painter IS (1998) Estimating the rate of evolution of the rate of molecular evolution. Mol Biol Evol 15:1647–1657
Aris-Brosou S, Yang Z (2002) Effects of models of rate evolution on estimation of divergence dates with special reference to the metazoan 18S ribosomal RNA phylogeny. Syst Biol 51:703–714
Aris-Brosou S, Yang Z (2003) Bayesian models of episodic evolution support a late precambrian explosive diversification of the Metazoa. Mol Biol Evol 20:1947–1954
Rannala B, Yang Z (1996) Probability distribution of molecular evolutionary trees: a new method of phylogenetic inference. J Mol Evol 43:304–311
Pybus OG, Rambaut A, Harvey PH (2000) An integrated framework for the inference of viral population history from reconstructed genealogies. Genetics 155:1429–1437
Drummond AJ, Rambaut A, Shapiro B, Pybus OG (2005) Bayesian coalescent inference of past population dynamics from molecular sequences. Mol Biol Evol 22:1185–1192
Minin VN, Bloomquist EW, Suchard MA (2008) Smooth skyride through a rough skyline: Bayesian coalescent-based inference of population dynamics. Mol Biol Evol 25:1459–1471
Hedges SB, Kumar S (2004) Precision of molecular time estimates. Trends Genet 20:242–247
Yang Z, Rannala B (2006) Bayesian estimation of species divergence times under a molecular clock using multiple fossil calibrations with soft bounds. Mol Biol Evol 23:212–226
Inoue J, Donoghue PCJ, Yang Z (2010) The impact of the representation of fossil calibrations on Bayesian estimation of species divergence times. Syst Biol 59:74–89
Drummond AJ, Ho SYW, Phillips MJ, Rambaut A (2006) Relaxed phylogenetics and dating with confidence. PLoS Biol 4:e88
Bouckaert R, Heled J, Kühnert D, Vaughan T, Wu CH, Xie D, Suchard MA, Rambaut A, Drummond AJ (2014) BEAST 2: a software platform for Bayesian evolutionary analysis. PLoS Comput Biol 10:e1003537
Wertheim JO, Sanderson MJ, Worobey M, Bjork A (2010) Relaxed molecular clocks, the bias-variance trade-off, and the quality of phylogenetic inference. Syst Biol 59:1–8
Lemey P, Rambaut A, Drummond AJ, Suchard MA (2009) Bayesian phylogeography finds its roots. PLoS Comput Biol 5:e1000520
Lemey P, Rambaut A, Welch JJ, Suchard MA (2010) Phylogeography takes a relaxed random walk in continuous space and time. Mol Biol Evol 27:1877–1885
Guillot G, Santos F, Estoup A (2008) Analysing georeferenced population genetics data with Geneland: a new algorithm to deal with null alleles and a friendly graphical user interface. Bioinformatics 24:1406–1407
Nadin-Davis SA, Feng Y, Mousse D, Wandeler AI, Aris-Brosou ST (2010) Spatial and temporal dynamics of rabies virus variants in big brown bat populations across Canada: footprints of an emerging zoonosis. Mol Ecol 19:2120–2136
Pagel M, Meade A (2004) A phylogenetic mixture model for detecting pattern-heterogeneity in gene sequence or character-state data. Syst Biol 53:571–581
Pagel M, Meade A, Barker D (2004) Bayesian estimation of ancestral character states on phylogenies. Syst Biol 53:673–684
Lartillot N, Poujol R (2011) A phylogenetic model for investigating correlated evolution of substitution rates and continuous phenotypic characters. Mol Biol Evol 28:729–744
Bromham L, Woolfit M, Lee MS, Rambaut A (2002) Testing the relationship between morphological and molecular rates of change along phylogenies. Evolution 56:1921–1930
Ho SYW, Duchêne S (2014) Molecular-clock methods for estimating evolutionary rates and timescales. Mol Ecol 23:5947–5965
dos Reis M, Donoghue PCJ, Yang Z (2016) Bayesian molecular clock dating of species divergences in the genomics era. Nat Rev Genet 17:71–80
Donoghue PCJ, Yang Z (2016) The evolution of methods for establishing evolutionary timescales. Philos Trans R Soc Lond B Biol Sci. https://doi.org/10.1098/rstb.2016.0020
Ho SY, Tong KJ, Foster CS, Ritchie AM, Lo N, Crisp MD (2015) Biogeographic calibrations for the molecular clock. Biol Lett 11:20150194
Kühnert D, Wu C-H, Drummond AJ (2011) Phylogenetic and epidemic modeling of rapidly evolving infectious diseases. Infect Genet Evol 11:1825–1141
Rieux A, Balloux F (2016) Inferences from tip-calibrated phylogenies: a review and a practical guide. Mol Ecol 25:1911–1924
Ho SYW, Chen AXY, Lins LSF, Duchêne DA, Lo N (2016) The genome as an evolutionary timepiece. Genome Biol Evol 8:3006–3010
O’Reilly JE, dos Reis M, Donoghue PCJ (2015) Dating tips for divergence-time estimation. Trends Genet 31:637–50
1000 Genomes Project Consortium, Abecasis GR, Altshuler D, Auton A, Brooks LD, Durbin RM, Gibbs RA, Hurles ME, McVean GA (2010) A map of human genome variation from population-scale sequencing. Nature 467:1061–1073
UK10K Consortium, Walter K, Min JL, Huang J et al (2015) The UK10K project identifies rare variants in health and disease. Nature 526:82–90
Ledford H (2016) AstraZeneca launches project to sequence 2 million genomes. Nature 532:427
Muse SV, Gaut BS (1994) A likelihood approach for comparing synonymous and nonsynonymous nucleotide substitution rates, with application to the chloroplast genome. Mol Biol Evol 11:715–724
Goldman N, Yang Z (1994) A codon-based model of nucleotide substitution for protein-coding DNA sequences. Mol Biol Evol 11:725–736
Kosiol C, Anisimova M (2011) Methods for detecting natural selection in protein-coding genes. In: Anisimova M (ed) Evolutionary genomics: statistical and computational methods. Methods in molecular biology series. Humana-Springer, New York
Thorne JL, Choi SC, Yu J, Higgs PG, Kishino H (2007) Population genetics without intraspecific data. Mol Biol Evol 24:1667–1677
Choi SC, Hobolth A, Robinson DM, Kishino H, Thorne JL (2007) Quantifying the impact of protein tertiary structure on molecular evolution. Mol Biol Evol 24:1769–1782
Halpern AL, Bruno WJ (1998) Evolutionary distances for protein-coding sequences: modeling site-specific residue frequencies. Mol Biol Evol 15:910–917
Yang Z, Nielsen R (2008) Mutation-selection models of codon substitution and their use to estimate selective strengths on codon usage. Mol Biol Evol 25:568–579
Rodrigue N, Philippe H, Lartillot N (2010) Mutation-selection models of coding sequence evolution with site-heterogeneous amino acid fitness profiles. Proc Natl Acad Sci USA 107:4629–4634
Rodrigue N, Lartillot N (2017) Detecting adaptation in protein-coding genes using a Bayesian site-heterogeneous mutation-selection codon substitution model. Mol Biol Evol 34:204–214
Bloom JD (2017) Identification of positive selection in genes is greatly improved by using experimentally informed site-specific models. Biol Direct 12:1. https://doi.org/10.1186/s13062-016-0172-z
Choi SC, Redelings BD, Thorne JL (2008) Basing population genetic inferences and models of molecular evolution upon desired stationary distributions of DNA or protein sequences. Philos Trans R Soc Lond B Biol Sci 363:3931–3939
Rodrigue N, Kleinman CL, Philippe H, Lartillot N (2009) Computational methods for evaluating phylogenetic models of coding sequence evolution with dependence between codons. Mol Biol Evol 26:1663–1676
Hartl DL, Clark AG (2007) Principles of population genetics, 4th edn. Sinauer Associates, Sunderland
Kimura M (1962) On the probability of fixation of mutant genes in a population. Genetics 47:713–719
Rice SH (2004) Evolutionary theory: mathematical and conceptual foundations. Sinauer Associates, Sunderland
Kimura M (1978) Change of gene frequencies by natural selection under population number regulation. Proc Natl Acad Sci USA 75:1934–1937
Tamuri A, dos Reis M, Goldstein R (2012) Estimating the distribution of selection coefficients from phylogenetic data using sitewise mutation-selection models. Genetics 190:1101–1115
Rodrigue N (2013) On the statistical interpretation of site-specific variables in phylogeny-based substitution models. Genetics 193:557–564
Prins P, Belhachemi D, Möller S, Smant G (2011) Scalable computing in evolutionary genomics. In: Anisimova M (ed) Evolutionary genomics: statistical and computational methods. Methods in molecular biology series. Humana-Springer, New York
Yang Z (1993) Maximum-likelihood estimation of phylogeny from DNA sequences when substitution rates differ over sites. Mol Biol Evol 10:1396–1401
Yang Z (1994) Maximum likelihood phylogenetic estimation from DNA sequences with variable rates over sites: approximate methods. J Mol Evol 39:306–314
Altekar G, Dwarkadas S, Huelsenbeck JP, Ronquist F (2004) Parallel Metropolis coupled Markov chain Monte Carlo for Bayesian phylogenetic inference. Bioinformatics 20:407–415
Lartillot N, Rodrigue N, Stubbs D, Richer J (2013) PhyloBayes MPI: phylogenetic reconstruction with infinite mixtures of profiles in a parallel environment. Syst Biol 62:611–615
Rodrigue N, Lartillot N (2014) Site-heterogeneous mutation-selection models within the PhyloBayes-MPI package. Bioinformatics 30:1020–1021
Stamatakis A, Hoover P, Rougemont J (2008) A rapid bootstrap algorithm for the RAxML Web servers. Syst Biol 57:758–771
Stamatakis A, Ludwig T, Meier H (2005) RAxML-III: a fast program for maximum likelihood-based inference of large phylogenetic trees. Bioinformatics 21:456–463
Stamatakis A, Göker M, Grimm GW (2010) Maximum likelihood analyses of 3,490 rbcL sequences: scalability of comprehensive inference versus group-specific taxon sampling. Evol Bioinform Online 6:73–90
Stamatakis A, Alachiotis N (2010) Time and memory efficient likelihood-based tree searches on phylogenomic alignments with missing data. Bioinformatics 26:i132–i139
Suchard MA, Rambaut A (2009) Many-core algorithms for statistical phylogenetics. Bioinformatics 25:1370–1376
Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542
Muir P, Li S, Lou S et al (2016) The real cost of sequencing: scaling computation to keep pace with data generation. Genome Biol 17:53
Schatz MC, Langmead B, Salzberg SL (2010) Cloud computing and the DNA data race. Nat Biotechnol 28:691–693
Dereeper A, Guignon V, Blanc G et al (2008) Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res 36:W465–W469
de Koning AP, Gu W, Pollock DD (2010) Rapid likelihood analysis on large phylogenies using partial sampling of substitution histories. Mol Biol Evol 27:249–265
Acknowledgements
We would like to thank Michelle Brazeau, Eric Chen, Ilya Hekimi, Benoît Pagé, and Wayne Sawtell for their critical reading of a draft of the original chapter, as well as Jonathan Dench and George S. Long for their careful reading of the most recent draft. This work was supported by the Natural Sciences Research Council of Canada (SAB, NR).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2019 The Author(s)
About this protocol
Cite this protocol
Aris-Brosou, S., Rodrigue, N. (2019). A Not-So-Long Introduction to Computational Molecular Evolution. In: Anisimova, M. (eds) Evolutionary Genomics. Methods in Molecular Biology, vol 1910. Humana, New York, NY. https://doi.org/10.1007/978-1-4939-9074-0_3
Download citation
DOI: https://doi.org/10.1007/978-1-4939-9074-0_3
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-4939-9073-3
Online ISBN: 978-1-4939-9074-0
eBook Packages: Springer Protocols