Abstract
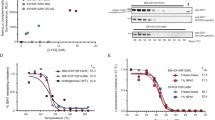
Cellular thermal shift assay (CETSA) is based on the thermal stabilization of the protein target by a compound binding. Thus, CETSA can be used to measure a compound’s cellular target engagement and permeability. HiBiT CETSA method is quantitative and has higher throughput compared to the traditional Western-based CETSA. Here, we describe the protocol for a HiBiT CETSA, which utilizes a HiBiT tag derived from the NanoLuciferase (NanoLuc) that upon complementation by LgBiT NanoLuc tag produces a bright signal enabling tracking of the effects of increasing temperature on the stability of a protein-of-interest in the presence/absence of various compounds. Exposure of a HiBiT-tagged protein to increasing temperatures induces protein denaturation and thus decreased LgBiT complementation and NanoLuc signal. As the stability of proteins at higher temperatures can be influenced by the compound binding, this method enables screening for target engagement in living or permeabilized cells.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Simon GM, Niphakis MJ, Cravatt BF (2013) Determining target engagement in living systems. Nat Chem Biol 9:200–205. https://doi.org/10.1038/NCHEMBIO.1211
Molina DM, Jafari R, Ignatushchenko M et al (2013) Monitoring drug target engagement in cells and tissues using the cellular thermal shift assay. Science 341:84–87. https://doi.org/10.1126/SCIENCE.1233606
Jafari R, Almqvist H, Axelsson H et al (2014) The cellular thermal shift assay for evaluating drug target interactions in cells. Nat Protoc 9:2100–2122. https://doi.org/10.1038/NPROT.2014.138
Hall MP, Unch J, Binkowski BF et al (2012) Engineered luciferase reporter from a deep sea shrimp utilizing a novel imidazopyrazinone substrate. ACS Chem Biol 7:1848–1857. https://doi.org/10.1021/CB3002478
Dart ML, Machleidt T, Jost E et al (2018) Homogeneous assay for target engagement utilizing bioluminescent thermal shift. ACS Med Chem Lett 9:546–551. https://doi.org/10.1021/ACSMEDCHEMLETT.8B00081/SUPPL_FILE/ML8B00081_SI_001.PDF
Sanchez TW, Owens A, Martinez NJ et al (2021) High-throughput detection of ligand-protein binding using a SplitLuc cellular thermal shift assay. Methods Mol Biol 2365:21–41. https://doi.org/10.1007/978-1-0716-1665-9_2
Martinez NJ, Asawa RR, Cyr MG et al (2018) A widely-applicable high-throughput cellular thermal shift assay (CETSA) using split Nano luciferase. Sci Reports 81(8):1–16. https://doi.org/10.1038/s41598-018-27834-y
Mortison JD, Cornella-Taracido I, Venkatchalam G et al (2021) Rapid evaluation of small molecule cellular target engagement with a luminescent thermal shift assay. ACS Med Chem Lett 12:1288–1294. https://doi.org/10.1021/ACSMEDCHEMLETT.1C00276/SUPPL_FILE/ML1C00276_SI_003.XLSX
Ramachandran S, Makukhin N, Haubrich K, et al (2022) Structure-based design of a phosphotyrosine-masked covalent ligand targeting the E3 ligase SOCS2. https://doi.org/10.26434/CHEMRXIV-2022-BVJ80
Larson HG, Zakharov AV, Sarkar S et al (2021) A genome-edited ERα-HiBiT fusion reporter cell line for the identification of ERα modulators via high-throughput screening and CETSA. Assay Drug Dev Technol 19:539–549. https://doi.org/10.1089/ADT.2021.059
Vasta JD, Corona CR, Robers MB (2021) A high-throughput method to prioritize PROTAC intracellular target engagement and cell permeability using NanoBRET. Methods Mol Biol 2365:265–282. https://doi.org/10.1007/978-1-0716-1665-9_14
Acknowledgements
This research was supported by the Natural Sciences and Engineering Research Council of Canada through a postdoctoral fellowship to V.V. and grant to D.B.L, and by the Structural Genomics Consortium, a registered charity (no: 1097737) that receives funds from Bayer AG, Boehringer Ingelheim, Bristol Myers Squibb, Genentech, Genome Canada through Ontario Genomics Institute [OGI-196], EU/EFPIA/ OICR/McGill/KTH/Diamond Innovative Medicines Initiative 2 Joint Undertaking [EUbOPEN grant 875510], Janssen, Merck KGaA (aka EMD in Canada and US), Pfizer, and Takeda. This project has also received funding from the Innovative Medicines Initiative 2 (IMI2) Joint Undertaking (JU) under grant agreement No 875510. The JU receives support from the European Union’s Horizon 2020 research and innovation program and EFPIA and Ontario Institute for Cancer Research, Royal Institution for the Advancement of Learning McGill University, Kungliga Tekniska Hoegskolan, Diamond Light Source Limited. S.R. is specifically funded by the IMI2 EUbOPEN project. S.H.B was supported by Mitacs Elevate Postdoctoral Fellowship.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Ramachandran, S., Szewczyk, M., Barghout, S.H., Ciulli, A., Barsyte-Lovejoy, D., Vu, V. (2023). HiBiT Cellular Thermal Shift Assay (HiBiT CETSA). In: Merk, D., Chaikuad, A. (eds) Chemogenomics. Methods in Molecular Biology, vol 2706. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-3397-7_11
Download citation
DOI: https://doi.org/10.1007/978-1-0716-3397-7_11
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-3396-0
Online ISBN: 978-1-0716-3397-7
eBook Packages: Springer Protocols